Abstract
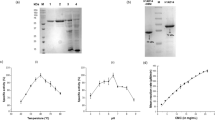
Biomass is the most abundant and short-term renewable natural resource on Earth whose recalcitrance toward enzymatic degradation represents significant challenge for a number of biotechnological applications. The not so abundant but critically necessary class of GH45 endoglucanases constitutes an essential component of tailored industrial enzyme cocktails because they randomly and internally cleave cellulose molecules. Moreover, GH45 glucanases are core constituents of major-brand detergent formulations as well as enzymatic aid components in the cotton processing industry, clipping unwanted cellulosic fibers from cotton (cellulosic)-based tissues. Here we report on a recombinant high-yield Neurospora crassa OR74A NcCel45A production system, a single-band GH45 endoglucanase purification, and a complete enzyme functional characterization. NcCel45A is a bimodular endoglucanase showing maximum activity at pH 6.0 and 60 °C, while most active against lichenan and β-glucans and lesser active toward filter paper, carboxymethylcellulose, and phosphoric acid-swollen cellulose. Gluco-oligosaccharide degradation fingerprinting experiments suggest cellopentaose as the minimal length substrate and ThermalFluor studies indicate that NcCel45A displays excellent stability at elevated temperatures up to 70 °C and pHs ranging from 5 to 9. Remarkably, we show that NcCel45A is uniquely resistant to a wide-range of organic solvents and small-angle X-ray scattering show a monkey-wrench molecular shape structure in solution, which indicates, unlike to other known cellulases, a non-fully extended conformation, thus conferring solvent protection. These NcCel45A unique enzymatic properties maybe key for specific industrial applications such as cotton fiber processing and detergent formulations.









Similar content being viewed by others
Abbreviations
- CBM:
-
Carbohydrate-binding module
- CMC:
-
Carboxymethyl cellulose
- DAM:
-
Dummy atom models
- DNS:
-
Dinitrosalycylic acid
- D max :
-
Maximum particle size
- EDTA:
-
Ethylenediaminetetraacetic acid
- GH:
-
Glycoside hydrolase
- HCCA:
-
Alpha-cyano-4-hydroxy cinnamic acid
- PASC:
-
Phosphoric acid-swollen cellulose
- PDB:
-
Protein Data Bank
- R g :
-
Radius of gyration
- pNPG:
-
p-Nitrophenyl-β-d-glucoside
- Rpm:
-
Rotations per minute
- SAXS:
-
Small-angle X-ray scattering
- T m :
-
Melting temperature
References
Jarvis, M. (2003). Chemistry—Cellulose stacks up. Nature, 426, 611–612.
Cosgrove, D. J. (2005). Growth of the plant cell wall. Nature Reviews Molecular Cell Biology, 6, 850–861.
Henrissat, B. (1994). Cellulases and their interaction with cellulose. Cellulose, 1, 169–196.
Zhang, Y. H., & Lynd, L. R. (2004). Toward an aggregated understanding of enzymatic hydrolysis of cellulose: Noncomplexed cellulase systems. Biotechnology and Bioengineering, 88, 797–824.
Horn, S. J., Vaaje-Kolstad, G., Westereng, B., & Eijsink, V. G. (2012). Novel enzymes for the degradation of cellulose. Biotechnology for Biofuels, 5, 45.
Himmel, M. E., Ding, S. Y., Johnson, D. K., Adney, W. S., Nimlos, M. R., Brady, J. W., & Foust, T. D. (2007). Biomass recalcitrance: Engineering plants and enzymes for biofuels production. Science, 315, 804–807.
Margeot, A., Hahn-Hagerdal, B., Edlund, M., Slade, R., & Monot, F. (2009). New improvements for lignocellulosic ethanol. Current Opinion in Biotechnology, 20, 372–380.
Cantarel, B. L., Coutinho, P. M., Rancurel, C., Bernard, T., Lombard, V., & Henrissat, B. (2009). The Carbohydrate-Active EnZymes database (CAZy): An expert resource for Glycogenomics. Nucleic Acids Research, 37, D233–D238.
Hirvonen, M., & Papageorgiou, A. C. (2003). Crystal structure of a family 45 endoglucanase from Melanocarpus albomyces: Mechanistic implications based on the free and cellobiose-bound forms. Journal of Molecular Biology, 329, 403–410.
Nakamura, Y., Moriya, T., Baba, Y., Yanai, K., Sumida, N., Nishimura, T., Murashima, K., Nakane, A., Yaguchi, T., Koga, J., Murakami, T. and Kono, T. (2000) Endoglucanases and cellulase preparations containing the same. WO patent 2000024879.
Jakobsen, T. S., Lindegaard, P., & Chan, M. (1998). Color-care cellulases: Fabric color shield. Inform, 9, 788–792.
Davies, G. J., Dodson, G. G., Hubbard, R. E., Tolley, S. P., Dauter, Z., Wilson, K. S., et al. (1993). Structure and function of endoglucanase V. Nature, 365, 362–364.
Couturier, M., Feliu, J., Haon, M., Navarro, D., Lesage-Meessen, L., Coutinho, P. M., & Berrin, J. G. (2011). A thermostable GH45 endoglucanase from yeast: Impact of its atypical multimodularity on activity. Microbial Cell Factories, 10, 103.
Igarashi, K., Ishida, T., Hori, C., & Samejima, M. (2008). Characterization of an endoglucanase belonging to a new subfamily of glycoside hydrolase family 45 of the basidiomycete Phanerochaete chrysosporium. Applied and Environment Microbiology, 74, 5628–5634.
Eyun, S. I., Wang, H., Pauchet, Y., Ffrench-Constant, R. H., Benson, A. K., Valencia-Jiménez, A., et al. (2014). Molecular evolution of glycoside hydrolase genes in the western corn rootworm (Diabrotica virgifera virgifera). PLoS ONE, 9, e94052.
Palomares-Rius, J. E., Hirooka, Y., Tsai, I. J., Masuya, H., Hino, A., Kanzaki, N., et al. (2014). Distribution and evolution of glycoside hydrolase family 45 cellulases in nematodes and fungi. BMC Evolutionary Biology, 14, 69.
Davies, G. J., Tolley, S. P., Henrissat, B., Hjort, C., & Schülein, M. (1995). Structures of oligosaccharide-bound forms of the endoglucanase V from Humicola insolens at 1.9 A resolution. Biochemistry, 34, 16210–16220.
Valjakka, J., & Rouvinen, J. (2003). Structure of 20 K endoglucanase from Melanocarpus albomyces at 1.8 angstrom resolution. Acta Crystallographica D, 59, 765–768.
Davis, R. H., & Perkins, D. D. (2002). Timeline: Neurospora: A model of model microbes. Nature Reviews Genetics, 3, 397–403.
Galagan, J. E., Calvo, S. E., Borkovich, K. A., Selker, E. U., Read, N. D., Jaffe, D., et al. (2003). The genome sequence of the filamentous fungus Neurospora crassa. Nature, 422, 859–868.
Romero, M. D., Aguado, J., González, L., & Ladero, M. (1999). Cellulase production by Neurospora crassa on wheat straw. Enzyme and Microbial Technology, 25, 244–250.
Eberhart, B. M., Beck, R. S., & Goolsby, K. M. (1977). Cellulase of Neurospora crassa. Journal of Bacteriology, 130, 181–186.
Yazdi, M. T., Woodward, J. R., & Radford, A. (1990). The cellulase complex of Neurospora crassa: Activity, stability and release. Journal of General Microbiology, 136, 1313–1319.
Nakamura, A., Ishida, T., Fushinobu, S., Kusaka, K., Tanaka, I., Inaka, K., et al. (2013). Phase-diagram-guided method for growth of a large crystal of glycoside hydrolase family 45 inverting cellulase suitable for neutron structural analysis. Journal of Synchrotron Radiat., 20, 859–863.
Liu, G., Wei, X., Qin, Y., & Qu, Y. (2010). Characterization of the endoglucanase and glucomannanase activities of a glycoside hydrolase family 45 protein from Penicillium decumbens 114-2. Journal of General and Applied Microbiology, 56, 223–229.
Sakamoto, K., & Toyohara, H. (2009). Molecular cloning of glycoside hydrolase family 45 cellulase genes from brackish water clam Corbicula japonica. Comparative Biochemistry and Physiology, 152, 390–396.
Koga, J., Baba, Y., Shimonaka, A., Nishimura, T., Hanamura, S., & Kono, T. (2008). Purification and characterization of a new family 45 endoglucanase, STCE1, from Staphylotrichum coccosporum and its overproduction in Humicola insolens. Applied and Environment Microbiology, 74, 4210–4217.
Shimonaka, A., Koga, J., Baba, Y., Nishimura, T., Murashima, K., Kubota, H., & Kono, T. (2006). Specific characteristics of family 45 endoglucanases from Mucorales in the use of textiles and laundry. Bioscience, Biotechnology, and Biochemistry, 70, 1013–1016.
Bukhtojarov, F. E., Ustinov, B. B., Salanovich, T. N., Antonov, A. I., Gusakov, A. V., Okunev, O. N., & Sinitsyn, A. P. (2004). Cellulase complex of the fungus Chrysosporium lucknowense: Isolation and characterization of endoglucanases and cellobiohydrolases. Biochemistry (Biokhimiia), 69, 542–551.
Baba, Y., Shimonaka, A., Koga, J., Kubota, H., & Kono, T. (2005). Alternative splicing produces two endoglucanases with one or two carbohydrate-binding modules in Mucor circinelloides. Journal of Bacteriology, 187, 3045–3051.
Takashima, S., Iikura, H., Nakamura, A., Hidaka, M., Masaki, H., & Uozumi, T. (1999). Comparison of gene structures and enzymatic properties between two endoglucanases from Humicola grisea. Journal of Biotechnology, 67, 85–97.
Altschul, S. F., Madden, T. L., Schäffer, A. A., Zhang, J., Zhang, Z., Miller, W., & Lipman, D. J. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402.
Notredame, C., Higgins, D. G., & Heringa, J. (2000). T-Coffee: A novel method for fast and accurate multiple sequence alignment. Journal of Molecular Biology, 302, 205–217.
Gouet, P., Robert, X., & Courcelle, E. (2003). ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Research, 31, 3320–3323.
Gasteiger, E., Hoogland, C., Gattiker, A., Duvaud, S., Wilkins, M. R., Appel, R. D. and Bairoch, A. (2005) Protein identification and analysis tools on the ExPASy server. Totowa: Humana Press.
Petersen, T. N., Brunak, S., von Heijne, G., & Nielsen, H. (2011). SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature Methods, 8, 785–786.
Steentoft, C., Vakhrushev, S. Y., Joshi, H. J., Kong, Y., Vester-Christensen, M. B., Schjoldager, K. T., et al. (2013). Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO Journal, 32, 1478–1488.
Artimo, P., Jonnalagedda, M., Arnold, K., Baratin, D., Csardi, G., de Castro, E., et al. (2012). ExPASy: SIB bioinformatics resource portal. Nucleic Acids Research, 40, W597–W603.
Wood, T. M. (1988). Preparation of crystalline, amorphous, and dyed cellulase substrates. In S. T. K. Willis & A. Wood (Eds.), Methods Enzymol (pp. 19–25). New York: Academic Press.
Segato, F., Damásio, A. R., Gonçalves, T. A., de Lucas, R. C., Squina, F. M., Decker, S. R., & Prade, R. A. (2012). High-yield secretion of multiple client proteins in Aspergillus. Enyzme and Microbial Technology, 51, 100–106.
Vogel, H. J. (1956). A convenient growth medium for Neurospora (Medium N). Microbial Genetics Bulletin, 13, 42–43.
Aslanidis, C., & de Jong, P. J. (1990). Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Research, 18, 6069–6074.
Camilo, C. M., & Polikarpov, I. (2014). High-throughput cloning, expression and purification of glycoside hydrolases using Ligation-Independent Cloning (LIC). Protein Expression and Purification, 99, 35–42.
Bradford, M. M. (1976). Rapid and sensitive method for quantitation of microgram quantities of protein utilizing principle of protein-dye binding. Analytical Biochemistry, 72, 248–254.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Westermeier, R., & Naven, T. (2002). Proteomics in practice: A laboratory manual of proteome analysis. Weinheim: Weiley.
Miller, G. L. (1959). Use of dinitrosalicylic acid reagent for determination of reducing sugar. Analytical Chem, 31, 426–428.
Hammersley, A. P. (1997) FIT2D: An introduction and overview. ESRF Internal Report.
Svergun, D. I. (1992). Determination of the regularization parameter in indirect-transform methods using perceptual criteria. Journal of Applied Crystallography, 25, 495–503.
Fischer, H., Oliveira Neto, M. D., Napolitano, H. B., Polikarpov, I., & Craievich, A. F. (2009). Determination of the molecular weight of proteins in solution from a single small-angle X-ray scattering measurement on a relative scale. Journal of Applied Crystallography, 43, 101–109.
Svergun, D. I. (1999). Restoring low resolution structure of biological macromolecules from solution. Biophysical Journal, 76, 2879–2886.
Volkov, V. V., & Svergun, D. I. (2003). Uniqueness of ab initio shape determination in small-angle scattering. Journal of Applied Crystallography, 36, 860–864.
Kozin, M. B., & Svergun, D. I. (2001). Automated matching of high- and low-resolution structural models. Journal of Applied Crystallography, 34, 33–41.
Schneidman-Duhovny, D., Hammel, M., & Sali, A. (2010). FoXS: A web server for rapid computation and fitting of SAXS profiles. Nucleic Acids Research, 38, W540–W544.
Arnold, K., Bordoli, L., Kopp, J., & Schwede, T. (2006). The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics, 22, 195–201.
Bernadó, P., Mylonas, E., Petoukhov, M. V., Blackledge, M., & Svergun, D. I. (2007). Structural characterization of flexible proteins using small-angle X-ray scattering. Journal of the American Chemical Society, 129, 5656–5664.
Otagiri, M., Lopez, C. M., Kitamoto, K., Arioka, M., Kudo, T., & Moriya, S. (2013). Heterologous expression and characterization of a glycoside hydrolase family 45 endo-beta-1,4-glucanase from a symbiotic protist of the lower termite, Reticulitermes speratus. Biotechnology and Applied Biochemistry, 169, 1910–1918.
Receveur, V., Czjzek, M., Schulein, M., Panine, P., & Henrissat, B. (2002). Dimension, shape, and conformational flexibility of a two domain fungal cellulase in solution probed by small angle X-ray scattering. Journal of Biological Chemistry, 277, 40887–40892.
Guttman, M., Weinkam, P., Sali, A., & Lee, K. K. (2013). All-atom ensemble modeling to analyze small-angle X-ray scattering of glycosylated proteins. Structure, 21, 321–331.
Karlsson, J., Siika-aho, M., Tenkanen, M., & Tjerneld, F. (2002). Enzymatic properties of the low molecular mass endoglucanases Cel12A (EG III) and Cel45A (EG V) of Trichoderma reesei. Journal of Biotechnology, 99, 63–78.
Murashima, K., Shimonaka, A., Nishimura, T., Baba, Y., Koga, J., Kubota, H., & Kono, T. (2006). Exploring amino acids responsible for the temperature profile of glycoside hydrolase family 45 endoglucanase EGL3 from Humicola grisea. Bioscience, Biotechnology, and Biochemistry, 70, 2205–2212.
Vlasenko, E., Schulein, M., Cherry, J., & Xu, F. (2010). Substrate specificity of family 5, 6, 7, 9, 12, and 45 endoglucanases. Bioresource Technology, 101, 2405–2411.
Saloheimo, A., Henrissat, B., Hoffren, A. M., Teleman, O., & Penttila, M. (1994). A novel, small endoglucanase gene, egl5, from Trichoderma reesei isolated by expression in yeast. Molecular Microbiology, 13, 219–228.
Shimonaka, A., Murashima, K., Koga, J., Baba, Y., Nishimura, T., Kubota, H., & Kono, T. (2006). Amino acid regions of family 45 endoglucanases involved in cotton defibrillation and in resistance to anionic surfactants and oxidizing agents. Bioscience, Biotechnology, and Biochemistry, 70, 2460–2466.
Davies, G., & Henrissat, B. (1995). Structures and mechanisms of glycosyl hydrolases. Structure, 3, 853–859.
Goto, M., Furukawa, K., & Hayashida, S. (1992). An Avicel-affinity site in an Avicel-digesting exocellulase from a Trichoderma viride mutant. Bioscience, Biotechnology, and Biochemistry, 56, 1523–1528.
Zverlov, V. V., Velikodvorskaya, G. A., & Schwarz, W. H. (2002). A newly described cellulosomal cellobiohydrolase, CelO, from Clostridium thermocellum: Investigation of the exo-mode of hydrolysis, and binding capacity to crystalline cellulose. Microbiology, 148, 247–255.
Davies, G. J., Dodson, G., Moore, M. H., Tolley, S. P., Dauter, Z., Wilson, K. S., et al. (1996). Structure determination and refinement of the Humicola insolens endoglucanase V at 1.5 angstrom resolution. Acta Crystallographica D, 52, 7–17.
Yennamalli, R. M., Rader, A. J., Kenny, A. J., Wolt, J. D., & Sen, T. Z. (2013). Endoglucanases: insights into thermostability for biofuel applications. Biotechnology for Biofuels, 6, 136.
Vehmaanperä, J., Alapuranen, M., Puranen, T., Siika-aho, M., Kallio, J., Hooman, S., Voutilainen, S., Halonen, T. and Viikari, L. (2013) Treatment of cellulosic material and enzymes useful therein. US patent 20130224801.
Yennamalli, R. M., Rader, A. J., Wolt, J. D., & Sen, T. Z. (2011). Thermostability in endoglucanases is fold-specific. BMC Structural Biology, 11, 10.
Betz, S. F. (1993). Disulfide bonds and the stability of globular proteins. Protein Science, 2, 1551–1558.
Beeby, M., O’Connor, B. D., Ryttersgaard, C., Boutz, D. R., Perry, L. J., & Yeates, T. O. (2005). The genomics of disulfide bonding and protein stabilization in thermophiles. PLoS Biology, 3, e309.
Jorda, J., & Yeates, T. O. (2011). Widespread disulfide bonding in proteins from thermophilic archaea. Archaea. doi:10.1155/2011/409156.
Wu, Z., & Lee, Y. Y. (1997). Inhibition of the enzymatic hydrolysis of cellulose by ethanol. Biotechnology Letters, 19, 977–979.
Karnaouri, A., Topakas, E., Paschos, T., Taouki, I., & Christakopoulos, P. (2013). Cloning, expression and characterization of an ethanol tolerant GH3 beta-glucosidase from Myceliophthora thermophila. PeerJ, 1, e46.
von Ossowski, I., Eaton, J. T., Czjzek, M., Perkins, S. J., Frandsen, T. P., Schulein, M., et al. (2005). Protein disorder: conformational distribution of the flexible linker in a chimeric double cellulase. Biophysical Journal, 88, 2823–2832.
Acknowledgments
This work was supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) via grants 2008/56255-9, 2009/54035-4, 2009/11536-3, 2010/08680-2, 11/20505-4 and 09/11536-3; Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) via grants 490022/2009-0, and 301981/2011-6; and the University of São Paulo (USP) via NAP de Bioenergia & Sustentabilidade (NAPBS) e NAP de Instrumentação (CIEA_MNB). We would like to thank Livia Manzine, Alexandre Antoniazzi, Maria Auxiliadora Morim Santos and Mariana Ortiz de Godoy for technical support and the Brazilian Synchrotron Light Laboratory (LNLS, Campinas) for access to the SAXS beam line.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kadowaki, M.A.S., Camilo, C.M., Muniz, A.B. et al. Functional Characterization and Low-Resolution Structure of an Endoglucanase Cel45A from the Filamentous Fungus Neurospora crassa OR74A: Thermostable Enzyme with High Activity Toward Lichenan and β-Glucan. Mol Biotechnol 57, 574–588 (2015). https://doi.org/10.1007/s12033-015-9851-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-015-9851-8