Abstract
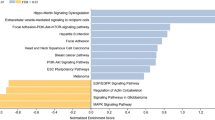
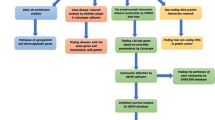
Gallbladder cancer (GBC) is one of the most fatal malignancies of the biliary tract system and is ranked sixth among the neoplasms of the gastrointestinal tract. Gallstone disease (GSD) is considered the major risk factor for GBC. However, the underlying molecular mechanism of GBC pathogenesis from different stages of GSD is not yet clearly understood. We analyzed transcriptomic datasets of GBC with reference to GSD of three different follow-up periods, i.e., GBC vs. GSD3 (1–3 years), GBC vs. GSD5 (5–10 years), and GBC vs. GSD10 (more than 10 years). We identified overlapping and specific molecular signatures in GBC compared with GSD at three different follow-up periods. Using integrative network biology approaches, such as protein–protein interaction network analysis, transcriptional regulatory network analysis, and miRNA–target gene network analysis, we have identified a few hub genes. The hub genes identified from GBC vs. GSD3, GBC vs. GSD5, and GBC vs. GSD10 were directly or indirectly associated with cancer progression and initiation from GSD. Functional enrichment analysis indicated significant correlation between GSD and GBC pathogenesis. The identified hub genes can be used for future targeted validation to develop potential diagnostic, prognostic, or therapeutic biomarkers in GBC.







Similar content being viewed by others
Dataset availability
The datasets used for this study are available at NCBI-GEO database (SRA Accession No: SRP226150).
References
Bailey TL, Boden M, Buske FA, et al. 2009 MEME Suite: Tools for motif discovery and searching. Nucleic Acids Res. 37 202–208
Barabási AL and Oltvai ZN 2004 Network biology: Understanding the cell’s functional organization. Nat. Rev. Genet. 5 101–113
Bommhardt U, Schraven B and Simeoni L 2019 Beyond TCR signaling: Emerging functions of Lck in cancer and immunotherapy. Int. J. Mol. Sci. 20 1–18
Bradley EW, Ruan MM and Oursler MJ 2008 Novel pro-survival functions of the kruppel-like transcription factor Egr2 in promotion of macrophage colony-stimulating factor-mediated osteoclast survival downstream of the MEK/ERK pathway. J. Biol. Chem. 283 8055–8064
Brenner D, Yilmaz R, Müller K, et al. 2018 Hot-spot KIF5A mutations cause familial ALS. Brain 141 688–697
Chand Y and Alam MA 2012 Network biology approach for identifying key regulatory genes by expression-based study of breast cancer. Bioinformation 8 1132–1138
Chen S, Zhou Y, Chen Y and Gu J 2018 Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34 i884–i890
Cheng D, Bao C, Zhang X, et al. 2018 LncRNA PRNCR1 interacts with HEY2 to abolish miR-448-mediated growth inhibition in non-small cell lung cancer. Biomed. Pharmacother. 107 1540–1547
Chin CH, Chen SH, Wu HH, et al. 2014 cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8 S11
Csardi G and Nepusz T 2006 The igraph software package for complex network research. InterJournal Complex Systems 1695 http://igraph.sf.net
de Sena Brandine G and Smith AD 2019 Falco: high-speed FastQC emulation for quality control of sequencing data. F1000Research 8 1874
Dennis G, Sherman BT, Hosack DA, et al. 2003 DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 4 https://doi.org/10.1186/gb-2003-4-9-r60
Forghanifard MM, Taleb S and Abbaszadegan MR 2015 Notch Signaling target genes are directly correlated to esophageal squamous cell carcinoma tumorigenesis. Pathol. Oncol. Res. 21 463–467
Franchina DG, Dostert C and Brenner D 2018 Reactive oxygen species: Involvement in T cell signaling and metabolism. Trends Immunol. 39 489–502
Fresno Vara JÁ, Casado E, de Castro J, et al. 2004 P13K/Akt signalling pathway and cancer. Cancer Treat. Rev. 30 193–204
Furlong LI 2013 Human diseases through the lens of network biology. Trends Genet. 29 150–159
Hosseinzadeh O, Hekmat Z, Nekoufar S, et al. 2020 Evaluate the gene expression of TPT1, EDN3, and ANO7 in prostate cancer tissues and their relation with age, tumor stage and family history. Meta Gene 24 100671
Hu XY, Ling ZN, Hong LL, et al. 2021 Circulating methylated THBS1 DNAs as a novel marker for predicting peritoneal dissemination in gastric cancer. J. Clin. Lab. Anal. https://doi.org/10.1002/jcla.23936
Hundal R and Shaffer EA 2014 Gallbladder cancer: Epidemiology and outcome. Clin. Epidemiol. 6 99–109
Hussain SP and Harris CC 2007 Inflammation and cancer: An ancient link with novel potentials. Int. J. Cancer 121 2373–2380
Janiszewska M, Primi MC and Izard T 2020 Cell adhesion in cancer: Beyond the migration of single cells. J. Biol. Chem. 295 2495–2505
Jedi M, Young GP, Pedersen SK and Symonds EL 2018 Methylation and gene expression of BCAT1 and IKZF1 in colorectal cancer tissues. Clin. Med. Insights Oncol. 12 1179554918775064
Jin Y, Jin W, Zheng Z, et al. 2017 GABRB2 plays an important role in the lymph node metastasis of papillary thyroid cancer. Biochem. Biophys. Res. Commun. 492 323–330
Kim D, Paggi JM, Park C, Bennett C and Salzberg SL 2019 Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37 907–915
Knox SS 2010 From “omics” to complex disease: A systems biology approach to gene-environment interactions in cancer. Cancer Cell Int. 10 1–13
Kumar Singh P, Kashyap A and Silakari O 2018 Exploration of the therapeutic aspects of Lck: A kinase target in inflammatory mediated pathological conditions. Biomed. Pharmacother. 108 1565–1571
Legler DF, Uetz-Von Allmen E and Hauser MA 2014 CCR7: Roles in cancer cell dissemination, migration and metastasis formation. Int. J. Biochem. Cell Biol. 54 78–82
Letelier P, Brebi P, Tapia O and Roa JC 2012 DNA promoter methylation as a diagnostic and therapeutic biomarker in gallbladder cancer. Clin. Epigenetics 4 11
Li X, Zhang Z, Yu M, et al. 2013 Involvement of miR-20a in promoting gastric cancer progression by targeting early growth response 2 (EGR2). Int. J. Mol. Sci. 14 16226–16239
Liao Y, Smyth GK and Shi W 2014 FeatureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30 923–930
Liu Z, Sanders AJ, Liang G, et al. 2017 Hey factors at the crossroad of tumorigenesis and clinical therapeutic modulation of hey for anticancer treatment. Mol. Cancer Therapeut. 16 775–786
Love MI, Anders S and Huber W 2014 Differential analysis of count data - the DESeq2 package. Genome Biol. 15 550
Marchesi F, Locatelli M, Solinas G, et al. 2010 Role of CX3CR1/CX3CL1 axis in primary and secondary involvement of the nervous system by cancer. J. Neuroimmunol. 224 39–44
Masoudi-Nejad A and Wang E 2015 Cancer modeling and network biology: Accelerating toward personalized medicine. Semin. Cancer Biol. 30 1–3
Miyamoto S, Hirata M, Yamazaki A, et al. 2004 Heparin-binding EGF-like growth factor is a promising target for ovarian cancer therapy. Cancer Res. 64 5720–5727
Miyata K, Yotsumoto F, Nam SO, Kuroki M and Miyamoto S 2012 Regulatory mechanisms of the HB-EGF autocrine loop in inflammation, homeostasis, development and cancer. Anticancer Res. 32 2347–2352
Muhammad JS, Khan MR and Ghias K 2018 DNA methylation as an epigenetic regulator of gallbladder cancer: An overview. Int. J. Surgery 53 178–183
Oliveros JC 2007 VENNY. An interactive tool for comparing lists with Venn diagrams (http://Bioinfogp.Cnb.Csic.Es/Tools/Venny/Index.Html)
Pedersen SK, Baker RT, McEvoy A, et al. 2015 A two-gene blood test for methylated DNA sensitive for colorectal cancer. PLoS One 10 1–14
Pruitt KD, Hogue CW, Groll M, et al. 2001 An automated method for finding molecular complexes in large protein interaction networks. Nucleic Acids Res. 29 137–140 https://academic.oup.com/nar/article-lookup/doi/10.1093/nar/29.1.137
Rath O and Kozielski F 2012 Kinesins and cancer. Nat. Rev. Cancer 12 527–539
Rawla P, Sunkara T, Thandra KC and Barsouk A 2019 Epidemiology of gallbladder cancer. Clin. Exp. Hepatol. 5 93–102
Roy N, Gaikwad M, Bhattacharrya DK and Barah P 2021 Identification of systems level molecular signatures from glioblastoma multiforme derived extracellular vesicles. J. Mol. Neurosci. 71 1156–1167
Sanchez-Vega F, Mina M, Armenia J, et al. 2018 Oncogenic signaling pathways in The Cancer Genome Atlas. Cell 173 321–337.e10
Sharma A, Sharma KL, Gupta A, Yadav A and Kumar A 2017 Gallbladder cancer epidemiology, pathogenesis and molecular genetics: Recent update. World J. Gastroenterol. 23 3978–3998
Shannon P, Markiel A, Ozier O, et al. 2003 Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13 2498–2504
Song X, Hu Y, Li Y, et al. 2020 Overview of current targeted therapy in gallbladder cancer. Signal Transduct. Target. Ther. 5 https://doi.org/10.1038/s41392-020-00324-2
Stormo GD 2000 DNA binding sites: Representation and discovery. Bioinformatics 16 16–23
Suratanee A and Plaimas K 2018 Network-based association analysis to infer new disease-gene relationships using large-scale protein interactions. PLoS One 13 1–20
Thomas-Chollier M, Sand O, Turatsinze JV, et al. 2008 RSAT: regulatory sequence analysis tools. Nucleic Acids Res. 36 https://doi.org/10.1093/nar/gkn304
Tian DW, Wu ZL, Jiang LM, et al. 2019 KIF5A promotes bladder cancer proliferation in vitro and in vivo. Dis. Markers https://doi.org/10.1155/2019/4824902
Tian S, Peng P, Li J, et al. 2020 SERPINH1 regulates EMT and gastric cancer metastasis via the Wnt/β-catenin signaling pathway. Aging 12 3574–3593
Unoki M and Nakamura Y 2003 EGR2 induces apoptosis in various cancer cell lines by direct transactivation of BNIP3L and BAK. Oncogene 22 2172–2185
Wang J, Zhu B, Zhang Y, et al. 2019 HEY2 acting as a co-repressor with smad3 and smad4 interferes with the response of TGF-beta in hepatocellular carcinoma. Am. J. Translat. Res. 11 4367–4381
Wang J, Xu C, Cheng Q, et al. 2020 RNA Sequencing revealed signals of evolution from gallbladder stone to gallbladder carcinoma. Front. Oncol. 10 https://doi.org/10.3389/fonc.2020.00823
Wangari-Talbot J, Wall BA, Goydos JS and Chen S 2012 Functional effects of GRM1 suppression in human melanoma cells. Mol. Cancer Res. 10 1440–1450
Wei LM, Cao S, Yu WD, Liu YL and Wang JT 2015 Overexpression of CX3CR1 is associated with cellular metastasis, proliferation and survival in gastric cancer. Oncol. Rep. 33 615–624
Weirauch MT, Yang A, Albu M, et al. 2014 Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158 1431–1443
Weiße J, Rosemann J, Müller L, et al. 2021 Identification of lymphocyte cell-specific protein-tyrosine kinase (LCK) as a driver for invasion and migration of oral cancer by tumor heterogeneity exploitation. Mol. Cancer 20 88
Wen Y, Li J, Koo J, et al. 2014 Activation of the glutamate receptor GRM1 enhances angiogenic signaling to drive melanoma progression. Cancer Res. 74 2499–2509
Wu W, Gao H, Li X, et al. 2019 LncRNA TPT1-AS1 promotes tumorigenesis and metastasis in epithelial ovarian cancer by inducing TPT1 expression. Cancer Sci. 110 1587–1598
Wu X, Zeng Z, Xu L, et al. 2014 Increased expression of IL17A in human gastric cancer and its potential roles in gastric carcinogenesis. Tumor Biol. 35 5347–5356
Yan L, Gong YZ, Shao MN, et al. 2020 Distinct diagnostic and prognostic values of γ-aminobutyric acid type A receptor family genes in patients with colon adenocarcinoma. Oncol. Lett. 20 275–291
Zhang L, Ye F, Zuo Z, et al. 2021 Long noncoding RNA TPT1-AS1 promotes the progression and metastasis of colorectal cancer by upregulating the TPT1-mediated FAK and JAK-STAT3 signalling pathways. Aging 13 3779–3797
Zhang X 2021 Upregulation of THBS1 is related to immunity and chemotherapy resistance in gastric cancer. J. Gen. Med. 14 4945–4957
Zhang Z, Wang Y, Zhang J, Zhong J and Yang R 2018 COL1A1 promotes metastasis in colorectal cancer by regulating the WNT/PCP pathway. Mol. Med. Rep. 17 5037–5042
Zhao X, Yu C, Zheng M and Sun J 2019 Prognostic value of the mRNA expression of gap junction α members in patients with gastric cancer. Oncol. Lett. 18 1669–1678
Acknowledgements
PB would like to acknowledge the Department of Biotechnology, India, for providing the Ramalingaswami Re-entry Fellowship grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors contributed to this manuscript declare no conflict of interests.
Additional information
Communicated by Susmita Roy.
Corresponding editor: Susmita Roy
This article is part of the Topical Collection: Emergent dynamics of biological networks.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Roy, N., Dihingia, B.R. & Barah, P. Integrative network-based approaches identified systems-level molecular signatures associated with gallbladder cancer pathogenesis from gallstone diseases. J Biosci 47, 31 (2022). https://doi.org/10.1007/s12038-022-00267-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-022-00267-6