Abstract
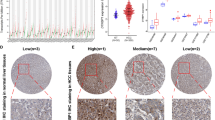
Novel and accurate biomarkers are needed for early detection and progression evaluation of hepatocellular carcinoma (HCC). Protein phosphatase 1 regulatory subunit 1A (PPP1R1A) has been studied in cancer biology; however, the expression pattern and biological function of PPP1R1A in HCC are unclear. The differentially expressed genes (DEGs) in HCC were screened by The Cancer Genome Atlas (TCGA) database. Real-time PCR and immunohistochemistry (IHC) assay were used to detect the expression of PPP1R1A in BALB/c mice, human normal tissues and corresponding tumor tissues, especially HCC. Then, Kaplan–Meier analysis of patients with HCC was performed to evaluate the relationship between PPP1R1A expression and prognosis. The transcriptional regulatory network of PPP1R1A was constructed based on the differentially expressed mRNAs, microRNAs and transcription factors (TFs). To explore the downstream regulation of PPP1R1A, the Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis and immune infiltration score were performed. A total of 4 DEGs were screened out. PPP1R1A was differentially distributed and expressed in BALB/c mice and human tissues. PPP1R1A expression was higher in normal tissues than that in tumor tissues, and patients with higher PPP1R1A expression had better clinical outcome in HCC. In addition, we constructed miR-21-3p/TAL1/PPP1R1A transcriptional network. Furthermore, PPP1R1A may modulate the activation of PI3K–Akt pathway, cell cycle, glycogen metabolism and the recruitment of M2 macrophage in HCC. This study may help to clarify the function and mechanism of PPP1R1A in HCC and provide a potential biomarker for tumor prevention and treatment.





Similar content being viewed by others
Data availability
All data is available under reasonable request.
Abbreviations
- PPP1R1A:
-
Protein phosphatase 1 regulatory subunit 1A
- PP1:
-
Protein phosphatase 1
- PPP:
-
Phosphoprotein phosphatase
- DEGs:
-
Differentially expressed genes
- TCGA:
-
The Cancer Genome Atlas
- TFs:
-
Transcription factors
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- PPI:
-
Protein–protein interaction
- TCIA:
-
The cancer immunome atlas
- OS:
-
Overall survival
- DFS:
-
Disease-free survival
- CCs:
-
Cellular components
- MFs:
-
Molecular functions
- BPs:
-
Biological processes
- PKA:
-
Protein kinase A
- IHC:
-
Immunohistochemistry
- HCC:
-
Hepatocellular carcinoma
- LIHC:
-
Liver hepatocellular carcinoma
- BLCA:
-
Bladder urothelial carcinoma
- COAD:
-
Colon adenocarcinoma
- BRCA:
-
Breast invasive carcinoma
- PAAD:
-
Pancreatic adenocarcinoma
- LGG:
-
Brain lower grade glioma
- KIRC:
-
Kidney renal clear cell carcinoma
- UVM:
-
Uveal melanoma
- ES:
-
Ewing sarcoma
- TAM:
-
Tumor-associated macrophage
References
Beste LA, Leipertz SL, Green PK, Dominitz JA, Ross D, Ioannou GN. Trends in burden of cirrhosis and hepatocellular carcinoma by underlying liver disease in US veterans, 2001–2013. Gastroenterology. 2015;149(6):1471–82 (e17-8).
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Ahn JC, Teng PC, Chen PJ, Posadas E, Tseng HR, Lu SC, et al. Detection of circulating tumor cells and their implications as a biomarker for diagnosis, prognostication, and therapeutic monitoring in hepatocellular carcinoma. Hepatology. 2021;73(1):422–36.
Jiang Y, Sun A, Zhao Y, Ying W, Sun H, Yang X, et al. Proteomics identifies new therapeutic targets of early-stage hepatocellular carcinoma. Nature. 2019;567(7747):257–61.
Wang X, Wang M, Li XY, Li J, Zhao DP. KIFC1 promotes the proliferation of hepatocellular carcinoma in vitro and in vivo. Oncol Lett. 2019;18(6):5739–46.
Lemaire S, Bollen M. Protein phosphatase-1: dual activity regulation by Inhibitor-2. Biochem Soc Trans. 2020;48(5):2229–40.
Krzyzosiak A, Sigurdardottir A, Luh L, Carrara M, Das I, Schneider K, et al. Target-based discovery of an inhibitor of the regulatory phosphatase PPP1R15B. Cell. 2018;174(5):1216–28.
Silva JV, Freitas MJ, Fardilha M. Phosphoprotein phosphatase 1 complexes in spermatogenesis. Curr Mol Pharmacol. 2014;7(2):136–46.
Alsina KM, Hulsurkar M, Brandenburg S, Kownatzki-Danger D, Lenz C, Urlaub H, et al. Loss of protein phosphatase 1 regulatory subunit PPP1R3A promotes atrial fibrillation. Circulation. 2019;140(8):681–93.
Verbinnen I, Ferreira M, Bollen M. Biogenesis and activity regulation of protein phosphatase 1. Biochem Soc Trans. 2017;45(1):89–99.
Tiwari A, Tashiro K, Dixit A, Soni A, Vogel K, Hall B, et al. Loss of HIF1A from pancreatic cancer cells increases expression of PPP1R1B and degradation of p53 to promote invasion and metastasis. Gastroenterology. 2020;159(5):1882–97.
Chen PC, Li C, Wang D, Luo ZW, Fu SJ, Li X, et al. PP-1alpha and PP-1gamma display antagonism and differential roles in tumorigenicity of lung cancer cells. Curr Mol Med. 2013;13(1):220–7.
Chen BY, Huang CH, Lin YH, Huang CC, Deng CX, Hsu LC. The K898E germline variant in the PP1-binding motif of BRCA1 causes defects in DNA Repair. Sci Rep. 2014;4:5812.
Schumann M, Gunzel D, Buergel N, Richter JF, Troeger H, May C, et al. Cell polarity-determining proteins Par-3 and PP-1 are involved in epithelial tight junction defects in coeliac disease. Gut. 2012;61(2):220–8.
Walsh JE, Young MR. TGF-beta regulation of focal adhesion proteins and motility of premalignant oral lesions via protein phosphatase 1. Anticancer Res. 2011;31(10):3159–64.
Van Dessel N, Boens S, Lesage B, Winkler C, Gornemann J, Van Eynde A, et al. Protein phosphatase PP1-NIPP1 activates mesenchymal genes in HeLa cells. Febs Lett. 2015;589(12):1314–21.
Paul D, Bargale AB, Rapole S, Shetty PK, Santra MK. Protein phosphatase 1 regulatory subunit SDS22 inhibits breast cancer cell tumorigenesis by functioning as a negative regulator of the AKT signaling pathway. Neoplasia. 2019;21(1):30–40.
Li H, Rokavec M, Jiang L, Horst D, Hermeking H. Antagonistic effects of p53 and HIF1A on microRNA-34a regulation of PPP1R11 and STAT3 and hypoxia-induced epithelial to mesenchymal transition in colorectal cancer cells. Gastroenterology. 2017;153(2):505–20.
Boratko A, Csortos C. TIMAP, the versatile protein phosphatase 1 regulator in endothelial cells. IUBMB Life. 2017;69(12):918–28.
Wang L, Zhao J, Ren J, Hall KH, Moorman JP, Yao ZQ, et al. Protein phosphatase 1 abrogates IRF7-mediated type I IFN response in antiviral immunity. Eur J Immunol. 2016;46(10):2409–19.
Luo W, Xu C, Ayello J, Dela CF, Rosenblum JM, Lessnick SL, et al. Protein phosphatase 1 regulatory subunit 1A in ewing sarcoma tumorigenesis and metastasis. Oncogene. 2018;37(6):798–809.
Shi SH, Jiang J, Zhang W, Sun L, Li XJ, Li C, et al. A novel lncRNA HOXC-AS3 Acts as a miR-3922-5p sponge to promote breast cancer metastasis. Cancer Invest. 2020;38(1):1–12.
Chen L, Xiang Z, Chen X, Zhu X, Peng X. A seven-gene signature model predicts overall survival in kidney renal clear cell carcinoma. Hereditas. 2020;157(1):38.
Zhou H, Chen Q, Tan W, Qiu Z, Li S, Song Y, et al. Integrated clinicopathological features and gene microarray analysis of pancreatic neuroendocrine tumors. Gene. 2017;625:72–7.
Yang M, Huang W, Sun Y, Liang H, Chen M, Wu X, et al. Prognosis and modulation mechanisms of COMMD6 in human tumours based on expression profiling and comprehensive bioinformatics analysis. Br J Cancer. 2019;121(8):699–709.
Liao L, Zhang L, Yang M, Wang X, Huang W, Wu X, et al. Expression profile of SYNE3 and bioinformatic analysis of its prognostic value and functions in tumors. J Transl Med. 2020;18(1):355.
Fan Y, Zhang L, Sun Y, Yang M, Wang X, Wu X, et al. Expression profile and bioinformatics analysis of COMMD10 in BALB/C mice and human. Cancer Gene Ther. 2020;27(3–4):216–25.
Liang Y, Li Q, Chen K, Ni W, Zhan Z, Ye F, et al. Zinc finger protein 307 functions as a tumorsuppressor and inhibits cell proliferation by inducing apoptosis in hepatocellular carcinoma. Oncol Rep. 2017;38(4):2229–36.
Chen P, Lei L, Wang J, Zou X, Zhang D, Deng L, et al. Downregulation of Talin1 promotes hepatocellular carcinoma progression through activation of the ERK1/2 pathway. Cancer Sci. 2017;108(6):1157–68.
Gao X, Wang X, Zhang S. Bioinformatics identification of crucial genes and pathways associated with hepatocellular carcinoma. 2018. Biosci Rep. https://doi.org/10.1042/BSR20181441.
Dow M, Pyke RM, Tsui BY, Alexandrov LB, Nakagawa H, Taniguchi K, et al. Integrative genomic analysis of mouse and human hepatocellular carcinoma. Proc Natl Acad Sci U S A. 2018;115(42):E9879–88.
Lou W, Chen J, Ding B, Chen D, Zheng H, Jiang D, et al. Identification of invasion-metastasis-associated microRNAs in hepatocellular carcinoma based on bioinformatic analysis and experimental validation. J Transl Med. 2018;16(1):266.
Felgueiras J, Jeronimo C, Fardilha M. Protein phosphatase 1 in tumorigenesis: is it worth a closer look? Biochim Biophys Acta Rev Cancer. 2020;1874(2): 188433.
Luo W, Xu C, Phillips S, Gardenswartz A, Rosenblum JM, Ayello J, et al. Protein phosphatase 1 regulatory subunit 1A regulates cell cycle progression in Ewing sarcoma. Oncotarget. 2020;11(19):1691–704.
Tolsma TO, Hansen JC. Post-translational modifications and chromatin dynamics. Essays Biochem. 2019;63(1):89–96.
Gao Z, Liu H, Shi Y, Yin L, Zhu Y, Liu R. Identification of cancer stem cell molecular markers and effects of hsa-miR-21–3p on stemness in esophageal squamous cell carcinoma. Cancers (Basel). 2019;11(4):518.
Zhu Y, Tang H, Zhang L, Gong L, Wu G, Ni J, et al. Suppression of miR-21-3p enhances TRAIL-mediated apoptosis in liver cancer stem cells by suppressing the PI3K/Akt/Bad cascade via regulating PTEN. Cancer Manag Res. 2019;11:955–68.
Baez-Vega PM, Echevarria VI, Valiyeva F, Encarnacion-Rosado J, Roman A, Flores J, et al. Targeting miR-21-3p inhibits proliferation and invasion of ovarian cancer cells. Oncotarget. 2016;7(24):36321–37.
Pink RC, Samuel P, Massa D, Caley DP, Brooks SA, Carter DR. The passenger strand, miR-21-3p, plays a role in mediating cisplatin resistance in ovarian cancer cells. Gynecol Oncol. 2015;137(1):143–51.
Jiao W, Leng X, Zhou Q, Wu Y, Sun L, Tan Y, et al. Different miR-21-3p isoforms and their different features in colorectal cancer. Int J Cancer. 2017;141(10):2103–11.
Hou N, Guo Z, Zhao G, Jia G, Luo B, Shen X, et al. Inhibition of microRNA-21-3p suppresses proliferation as well as invasion and induces apoptosis by targeting RNA-binding protein with multiple splicing through Smad4/extra cellular signal-regulated protein kinase signalling pathway in human colorectal cancer HCT116 cells. Clin Exp Pharmacol Physiol. 2018;45(7):729–41.
Li Y, Liao Z, Luo H, Benyoucef A, Kang Y, Lai Q, et al. Alteration of CTCF-associated chromatin neighborhood inhibits TAL1-driven oncogenic transcription program and leukemogenesis. Nucleic Acids Res. 2020;48(6):3119–33.
Mansour MR, Sanda T, Lawton LN, Li X, Kreslavsky T, Novina CD, et al. The TAL1 complex targets the FBXW7 tumor suppressor by activating miR-223 in human T cell acute lymphoblastic leukemia. J Exp Med. 2013;210(8):1545–57.
Sanda T, Lawton LN, Barrasa MI, Fan ZP, Kohlhammer H, Gutierrez A, et al. Core transcriptional regulatory circuit controlled by the TAL1 complex in human T cell acute lymphoblastic leukemia. Cancer Cell. 2012;22(2):209–21.
Ding C, Tang W, Wu H, Fan X, Luo J, Feng J, et al. The PEAK1-PPP1R12B axis inhibits tumor growth and metastasis by regulating Grb2/PI3K/Akt signalling in colorectal cancer. Cancer Lett. 2019;442:383–95.
Huang H, Tian H, Duan Z, Cao Y, Zhang XS, Sun F. microRNA-383 impairs phosphorylation of H2AX by targeting PNUTS and inducing cell cycle arrest in testicular embryonal carcinoma cells. Cell Signal. 2014;26(5):903–11.
Shahmoradgoli M, Riazalhosseini Y, Haag D, Becker N, Hovestadt V, Heck S, et al. Protein phosphatase 1, regulatory subunit 15B is a survival factor for ERalpha-positive breast cancer. Int J Cancer. 2013;132(11):2714–9.
Shen GM, Zhang FL, Liu XL, Zhang JW. Hypoxia-inducible factor 1-mediated regulation of PPP1R3C promotes glycogen accumulation in human MCF-7 cells under hypoxia. FEBS Lett. 2010;584(20):4366–72.
Ramesh A, Kumar S, Nandi D, Kulkarni A. CSF1R- and SHP2-inhibitor-loaded nanoparticles enhance cytotoxic activity and phagocytosis in tumor-associated macrophages. Adv Mater. 2019;31(51): e1904364.
Ovais M, Guo M, Chen C. Tailoring nanomaterials for targeting tumor-associated macrophages. Adv Mater. 2019;31(19): e1808303.
Yeung OW, Lo CM, Ling CC, Qi X, Geng W, Li CX, et al. Alternatively activated (M2) macrophages promote tumour growth and invasiveness in hepatocellular carcinoma. J Hepatol. 2015;62(3):607–16.
Acknowledgements
Not applicable.
Funding
This work was supported by the President Foundation of Nanfang Hospital, Southern Medical University (NO. 2020C039); Clinical Research Startup Program of Southern Medical University (NO. LC2019ZD008); Clinical Research Program of Nanfang Hospital, Southern Medical University (NO. 2018CR021 and NO. 2020CR025); National Natural Science Foundation of China (NO. 82102839); China Postdoctoral Science Foundation (NO. 2020M682815); Guangdong Basic and Applied Basic Research Foundation (NO. 2020A1515110787).
Author information
Authors and Affiliations
Contributions
JG and FY: contributed to the study design and draft revision. MY, YQW and NL: contributed to human specimen collection. XXW, LWL and LSZ: contributed to implementation and analysis of real-time PCR and IHC assay. XQW, MYM and YW: contributed to mice and tissue preparation. XXW and YW: contributed to data collection and interpretation of bioinformatics results. XXW: contributed to draft the manuscript and coordinate data collection and analysis. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interests.
Ethical approval and informed consent
This study was approved by the Ethics Committee of Nanfang Hospital of Southern Medical University. All participants offered written informed consent before operation. The study conforms to the provisions of the Declaration of Helsinki. Generated Statement: no potentially identifiable human images or data is presented in this study. All animal experiments were performed according to the institutional guidelines and approved by the Nanfang hospital animal ethic committee on January 11, 2018, application number NFYY-2018-05.
Consent for publication
All authors consent to the publication of this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13577_2022_771_MOESM1_ESM.tif
Figure S1 Biological distribution and expression analysis of PPP1R1A in human normal tissues using UCSC database. (TIF 1518 KB)
13577_2022_771_MOESM2_ESM.tif
Figure S2 Prognostic value of PPP1R1A in (A) pancreatic adenocarcinoma (PAAD), (B) brain lower grade glioma (LGG), (C) kidney renal clear cell carcinoma (KIRC) and (D) uveal melanoma (UVM). (TIF 1406 KB)
13577_2022_771_MOESM3_ESM.tif
Figure S3 Relationship between PRRX2 and PPP1R1A in HCC. A Expression of PRRX2 in HCC patients with MT or NMT based on TCGA database. B Pearson analysis of correlation between the expression of PRRX2 and PPP1R1A in HCC. (TIF 270 KB)
13577_2022_771_MOESM4_ESM.xls
Table S1 Expression of ten miRNAs with detailed transcriptional information in NMT group and MT group based on TCGA–LIHC database. NMT: without recurrence and metastasis in 3 years; MT: with recurrence and metastasis in 1 year. (XLS 61 KB)
13577_2022_771_MOESM5_ESM.xls
Table S2 Differential expression of transcription factors in NMT group compared with MT group according to TCGA–LIHC database. NMT: without recurrence and metastasis in 3 years; MT: with recurrence and metastasis in 1 year (XLS 64 KB)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, X., Wang, Y., Yang, M. et al. Exploring prognostic value and regulation network of PPP1R1A in hepatocellular carcinoma. Human Cell 35, 1856–1868 (2022). https://doi.org/10.1007/s13577-022-00771-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-022-00771-9