Abstract
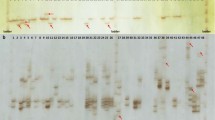
Pisum sativum specific sequence tagged microsatellite site primers were used to amplify genomic profiles from 15accessions of P. sativum L. that represented the genetic base of the Australian field pea-breeding program and five accessions of the wild related species P. fulvum. The STMS primers were used to assess genetic relationships among the Pisum accessions in two ways. Firstly, to produce RAPD-like multiple banding marker profiles using an adapted RAMS method, for intra- and interspecific diversity analysis. From the 14 flanking primer pairs assessed, 133 markers were obtained. Conservation and reproducibility of markers among individuals within accessions was demonstrated. The largest distance observed among P. sativumaccessions was 22% and among P.fulvum accessions was 40%, similar to that revealed with other PCR-based methods. The maximum distance between P.sativum and P. fulvum accessions was 46%. Phylogenetic clustering of P. sativum accessions, using the neighbour joining method and based on simple matching distances, was distinct and distant to P. fulvum. Secondly, PCR with a higher annealing temperature and fluorescent labeling identified simple and allelic loci markers useful for creating agenotype/fingerprint database for P. sativum cultivars. This is the first report to demonstrate the use of Pisum specific STMS sequences for both diversity analysis and genotype identification.
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ford, R., Le Roux, K., Itman, C. et al. Diversity analysis and genotyping in Pisum with sequence tagged microsatellite site (STMS) primers. Euphytica 124, 397–405 (2002). https://doi.org/10.1023/A:1015752907108
Issue Date:
DOI: https://doi.org/10.1023/A:1015752907108