Abstract
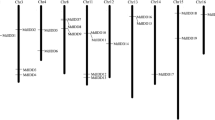
cDNA fragments representing 21 R2R3-MYB genes were isolated by RT-PCR from the Dendrobiumorchid hybrid Woo Leng. Six full-length cDNA clones were obtained from a flower cDNA library, four of which, DwMYB1, DwMYB2, DwMYB8 and DwMYB10, represent typical plant R2R3-MYB genes. The conceptual DwMYB4 protein is truncated at the C-terminal region and contains the R2 repeat and the N-terminal half of the R3 repeat (R2R3′). DwMYB4 expression is restricted to flowers. DwMYB9 contains an 8 amino acid N-terminal deletion in the R2 repeat (R2′R3) and is expressed at high levels in mature flower and inflorescence, but at very low levels in young flower buds. DwMYB8 and DwMYB10 show similar expression patterns and share very high sequence similarity in the N-terminal part of the MYB domain. Analysis of amino acid substitution indicated that the pattern and type of substitution between Arabidopsis and maize are quite different. Maize may have more conserved substitution in the MYBBRH domain than Arabidopsis.
Similar content being viewed by others

References
Arabidopsis Genome Initiative. 2000. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 791–815.
Avila, J., Nieto, C., Canas, L., Benito, M.J. and Paz-Ares, J. 1993. Petunia hybrida genes related to the maize regulatory C1 gene and to animal myb-oncogenes. Plant J. 3: 553–562.
Bender, J. and Fink, G.R. 1998. A MYB homologue, ATR1, activates tryptophan gene synthesis in Arabidopsis. Proc. Natl. Acad. Sci. USA 95: 5655–5660.
Borevitz, J.O., Xia, Y., Blount, J., Dixon, R.A. and Lamb, C. 2000. Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12: 2383–2393.
Braun, E.L. and Grotewold, E. 1999. Newly discovered plant c-Myb-like genes rewrite the evolution of the plant MYB gene family. Plant Physiol. 121: 21–24.
Cone, K.C., Coccioline, S.M., Burr, F.A. and Burr, B. 1993. Maize anthocyanin regulatory gene Pl is a duplicate of C1 that functions in plant. Plant Cell 5: 1795–1805.
Daniel, X., Lacomme, C., Morel, J.B. and Roby, D. 1999. A novel MYB oncogene homologue in Arabidopsis thaliana related to hypersensitive cell death. Plant J 20: 57–66.
Dressler, R.L. 1990. The Orchids: Natural History and Classification. Harvard University Press, Cambridge, MA.
Feldbrugge, M. 1997. PcMYB1, a novel plant protein containing a DNA-binding domain with one MYB repeat, interacts in vivo with a light-regulatory promoter unit. Plant J. 11: 1079–1093.
Franken, P., Schrell, S., Peterson, P.A., Saedler, H. and Wienand, U. 1994. Molecular analysis of protein domain function encoded by the MYB-homologous maize genes C1, Zm1 and Zm38. Plant J. 6: 21–30.
Gabrielsen, O.S., Sentenac, A. and Fromageot, P. 1991. Specific DNA binding by c-MYB: evidence for a double helix-turn-helix related motif. Science 253: 1140–1143.
Glover, B.J., Perez-Rodriguez, M. and Martin, C. 1998. Development of several epidermal cell types can be specified by the same MYB-related plant transcription factor. Development 125: 3497–3508.
Gocal, F.F.W., Poole, A.T., Gulbler, F., Watts, R.J., Blundell, C. and King, R.W. 1999. Long-day up-regulation of a GAMYB gene during Lolium temulentum inflorescence formation. Plant Physiol. 119: 1271–1278.
Grotewold, E., Frummond, B.J., Bowen, B. and Peterson, T. 1994. The MYB-homologous P gene controls phlobaphene pigmentation in maize floral organs by directly activating a flavonoid biosynthetic gene subset. Cell 76: 543–553.
Gubler, F., Kalla, R., Roberts, J.K. and Jacobsen, J.V. 1995. Gibberellin-regulated expression of a MYB gene in barley aleurone cells: Evidence for MYB transactivation of a high-pI ?-amylase gene promoter. Plant Cell 7: 1879–1891.
Gubler, F., Raventos, D., Keys, M., Watts, R., Mundy, J. and Jacobsen, J.V. 1999. Target genes and regulatory domains of the GaMYB transcriptional activator in cereal aleurone. Plant J. 17: 1–9.
Hegovold, A.B. and Gabrielsen, O.S. 1996. The importane of the linker connecting the repeats of the c-MYB oncoprotein may be due to a positioning function. Nucl. Acids Res. 24: 3990–3995.
Hoeren, F.U., Dolferus, R., Wu, Y., Peacock, W.J. and Dennis, E.S. 1998. Evidence for a role of AtMYB2 in the induction of the Arabidopsis alcohol dehydrogenase gene (ADH1) by low oxygen. Genetics 149: 479–490.
Howe, K.M., Reakes, C.F.L. and Watson, R.J. 1990. Characterization of the sequence-specific interaction of mouse c-Myb protein with DNA. EMBO J. 9: 1610–1619.
International Human Genome Sequencing Consortium. 2001. Initial sequencing and analysis of the human genome. Nature 409: 860–921.
Iturriaga, G., Leyns, L., Villegas, A., Gharaibeh, R., Salamini, F. and Bartels D. 1996. A family of novel MYB-related genes from the resurrection plant Craterostigma plantagineum is specifically expressed in callus and roots in response to ABA or desiccation. Plant Mol. Biol. 32: 707–716.
Jin, H., Cominelli, E., Bailey, P., Parr, A., Mehrtens, F., Jones, J., Tonelli, C., Weisshaar, B. and Martin, C. 2000. Transcriptional repression by AtMYB4 controls production of UV-protecting sunscreens in Arabidopsis. EMBO J. 19: 6150–6161.
Kanei-Ishii, C., Sarai, A., Sawazaki, T., Nakagoshi, H., He, D.N., Ogata, K., Nishimura, Y. and Ishii, S. 1990. The tryptophan cluster: a hypothetical structure of the DNA-binding domain of the MYB proto-oncogene product. J. Biol. Chem. 265: 19990–19995.
Kirik, V., Kolle, K., Misera, S., Baumlein, H. 1998. Two novel MYB homologues with changed expression in late embryogenesisdefective Arabidopsis mutants. Plant Mol. Biol. 37: 819–827.
Kirik, V., Schnittger, A., Radchuk, V., Adler, K., Hulskamp, M. and Baumlein, H. 2001. Ectopic expression of the Arabidopsis At-MYB23 gene induces differentiation of trichome cells. Dev Biol. 235: 366–377.
Klempnauer, K.H., Gonda, T.J. and Bishop, J.M. 1982. Nucleotide sequence of the retroviral leukemia gene v-MYB and its cellular progenitor c-MYB: the architecture of a transduced oncogene. Cell 31: 453–463.
Kranz, H.D., Denekamp, M., Greco, R., Jin, H.L., Leyva, A., Meissner, R.C., Petroni, K., Urzainqiu, A., Bevan, M., Martin, C., Smeekens, S., Tonelli, C., Paz-Ares, J. and Weisshaar, B. 1998. Toward functional characterisation of the members of the R2-R3-MYB gene family from Arabidopsis thaliana. Plant J. 6: 263–276.
Kranz, H., Scholz, K. and Weisshaar, B. 2000. c-MYB oncogenelike genes encoding three MYB repeats occur in all major plant lineages. Plant J. 21: 231–235.
Lee, M.M. and Schiefelbein, J. 1999, WEREWOLF, a MYB-related protein in Arabidopsis, is a position-dependent regulator of epidermal cell patterning. Cell 99: 473–483.
Lee, M.M., Qi, M., and Yang, Y. 2001. A novel jasmonic acidinducible rice MYB gene associates with fungal infection and host cell death. Mol. Plant-Microbe Interact. 14: 527–535.
Lipsick, J.S. 1996. One billion years of MYB. Oncogene 13: 223–235.
Lloyd, A.M., Walbot, V. and Davis, R.W. 1992. Arabidopsis and Nicotiana anthocyanin production activated by maize regulators R and C1. Science 258: 1773–1775.
Martin, C. and Paz-Ares, J. 1997. MYB transcription factors in plants. Trends Genet. 13: 67–73.
Meissner, R.C., Jin, H., Cominelli, E., Denekamp, M., Fuertes, A., Greco, R., Kranz, H.D., Penfield, S., Petroni, K., Urzainqui, A., et al. 1999. Function search in a large transcription factor gene family in Arabidopsis: assessing the potential of reverse genetics to identify insertional mutations in R2R3 MYB genes. Plant Cell 11: 1827–1840.
Mol, J., Grotewold, E. and Koes, R. 1998. How genes paint flowers and seeds. Trends Plant Sci. 3: 212–217.
Moyano, E., Martinez-Garcia., F. and Martinez, C. 1996. Apparent redundancy in MYB gene function provides gearing for the control of flavonoid biosynthesis in Antirrhinum flowers. Plant Cell 8: 1519–1532.
Nadeau, J.A., Zhang, X.S., Li, J. and O'Neill, S.D. 1996. Ovule development: identification of stage-specific and tissue-specific cDNAs. Plant Cell 8: 213–239.
Noda, K., Glover, B.J., Linstead, P. and Martin, C. 1994. Flower color intensity depends on specialized cell shape controlled by a MYB-related transcription factor. Nature 369: 661–664.
Ogata, K., Hojo, H., Aimoto, S., Nakai, T., Nakamura, H., Sarai, A., Ishii, S. and Nishimura, Y. 1992. Solution structure of a DNA binding unit of MYB: a helix-turn-helix-related motif with conserved tryptophans forming a hydrophobic core. Proc. Natl. Acad. Sci. USA 89: 6428–6432.
Oppenheimer, D.G., Herman, P.L., Sivakumaran, S., Esch, J. and Marks, M.D. 1991. A MYB gene required for leaf trichome differentiation in Arabidopsis is expressed in stipules. Cell 67: 483–493.
Payne, T., Clement, J., Arnold, D. and Lloyd, A. 1999. Heterologous MYB genes distinct from GL1 enhance trichome production when overexpressed in Nicotiana tabacum. Development 126: 671–682.
Payne, C.T., Zhang, F. and Lloyd, A.M. 2000. GL3 encodes a bHLH protein that regulates trichome development in Arabidopsis through interaction with GL1 and TTG1. Genetics 156: 1349–1362.
Panfield, S., Meissner, R.C., Shoue, D.A., Carpita, N.C. and Bevan, M.W. 2001. MYB61 is required for mucilage deposition and extrusion in the Arabidopsis seed coat. Plant Cell 13: 2777–2791.
Paz-Ares, J., Ghosal, D., Wienand, U., Peterson, P.A. and Saedler, H. 1987. The regulatory c1 locus of Zea mays encodes a protein with homology to MYB proto-oncogene products and with structural similarities to transcriptional activators. EMBO J. 6: 3553–3558.
Rabinowicz, P.D., Braun, E.L., Wolfe, A.D., Bowen, B. and Grotewold, E. 1999. Maize R2R3 MYB genes: sequence analysis reveals amplification in the higher plants. Genetics 153: 427–444.
Romero, I., Fuertes, A., Benito, M. J., Malpica, J. M., Leyva A. and Paz-Ares, J. 1998. More than 80 R2R3-MYB regulatory genes in the genome of Arabidopsis thaliana. Plant J. 14: 273–284.
Sablowski, R.W.M., Moyano, E., Cullianez-Macia, F.A., Schuch, W., Martin, C. and Bevan, M. 1994. A flower-specific MYB protein activates transcription of phenylpropanoid biosynthetic genes. EMBO J. 13: 128–137.
Saikumar, P., Murali, R. and Reddy, E. P. 1990. Role of tryptophan repeats and flanking amino acids in MYB-DNA interactions. Proc. Natl. Acad. Sci. 87: 8452–9456.
Saitou, N. and Nei, M. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4: 406–425.
Sambrook, J., Fritsch, E.F. and Maniatis, T. 1989. Molecular Cloning: A Laboratory Manual, 2nd ed. Cold Spring Harbor Laboratory Press, Plainview, NY.
Schmitz, G., Tillmann, E., Carriero, F., Fiore, C. and Theres, K. 2002. The tomato Blind gene encodes a MYB transcription factor that controls the formation of lateral meristems. Proc. Natl. Acad. Sci. USA 99: 1064–1069.
Solano, R. 1995. Dual DNA-binding specificity of a petal epidermisspecific MYB transcription factor (MYB.PH3) from Petunia hybrida. EMBO J. 14: 1773–1784.
Stracke, R., Werber, M., and Weisshaar, B. 2001. The R2R3-MYB gene family in Arabidopsis. Curr. Opin. Plant Biol. 4: 447–456.
Swofford, D.L. 1998. PAUP*: phylogenetic analysis using parsimony (* and other methods), version 4.0b1. Sinauer Associates, Sunderland, MA.
Tamagnone, L., Merida, A., Parr, A., Mackay, S., Culianez-Macia, F.A., Roberts, K. and Martin, C. 1998. The AmMYB308 and AmMYB330 transcription factors from Antirrhinum regulate phenylpropanoid and lignin biosynthesis in transgenic tobacco. Plant Cell 10: 135–154.
Taylor, D., Badiani, P. and Weston, K.A. 1996. A dominant interfering Myb mutant cause apoptosis in T cells. Genes Dev. 10: 2732–2744.
Timmermans, M.C., Hudson, A., Becraft, P.W. and Nelson, T. 1999 ROUGH SHEATH2: a MYB protein that represses knox homeobox genes in maize lateral organ primordia. Science 284: 151–153.
Toscani, A., Mettus, R. V., Coupland, R., Simpkins, H., Litvin, J., Orth, J., Hatton, K.S. and Reddy, E.P. 1997. Arrest of spermatogenesis and defective breast development in mice lacking A-MYB. Nature 386: 713–717.
Uimari, A. and Strommer, J. 1997. MYB26: a MYB-like protein of pea flowers with affinity for promoters of phenylpropanoid genes. Plant J. 12: 1273–1284.
Urao, T., Yamaguchi, S.K., Urao, S. and Shinozaki, K. 1993. An Arabidopsis MYB homolog is induced by dehydration stress and its gene product binds to the conserved Myb recognition sequence. Plant Cell 5: 1529–1539.
van Aalten, C.M.F., Grotewold, E. and Joshua-Tor, L. 1998. Essential dynamics from NMR structures: dynamic properties of the MYB DNA-binding domain and a hinge-bending enhancing variant. Methods 14: 318–328.
Waites, R., Selvadurai, H.R.N., Oliver, I.R. and Hudson, A. 1998. The PHANTASTICA gene encodes a MYB transcription factor involved in growth and dorsoventrality of lateral organs in Antirrhinum. Cell 93: 779–789.
Williams, C.E. and Grotewold, E. 1997. Differences between plant and animal MYB domains are fundamental for DNA binding activity, and chimeric MYB domains have novel DNA binding specificities. J. Biol. Chem. 272: 563–571.
Yang, Y. and Klessig, D.F. 1996. Isolation and characterization of a tobacco mosaic virus-inducible MYB oncogene homolog from tobacco. Proc. Natl. Acad. Sci. 93: 14927–14977.
Yu, H. and Goh, C.J. 2002. Molecular genetics of reproductive biology in orchids. Plant Physiol. 127: 1390–1393.
Yu, H., Yang, S.H. and Goh, C.J. 2000. DOH1, a class 1 knox gene, is required for maintenance of the basic plant architecture and floral transition in orchid. Plant Cell 12: 2143–2159.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, XM., Lim, SH. & Yang, WC. Characterization, expression and phylogenetic study of R2R3-MYB genes in orchid. Plant Mol Biol 51, 959–972 (2003). https://doi.org/10.1023/A:1023050110077
Issue Date:
DOI: https://doi.org/10.1023/A:1023050110077