Abstract
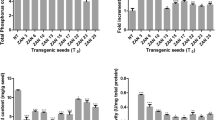
Phytate is the main storage form of phosphorus in many plant seeds, but phosphate bound in this form is not available to monogastric animals. Phytase, an enzyme that hydrolyzes phosphate from phytate, has the potential to enhance phosphorus availability in animal diets when engineered in rice seeds as a feed additive. Two genes, derived from a ruminal bacterium Selenomonas ruminantium (SrPf6) and Escherichia coli (appA), encoding highly active phytases were expressed in germinated transgenic rice seeds. Phytase expression was controlled by a germination inducible α-amylase gene (αAmy8) promoter, and extracellular phytase secretion directed by an αAmy8 signal peptide sequence. The two phytases were expressed in germinated transgenic rice seeds transiently and in a temporally controlled and tissue-specific manner. No adverse effect on plant development or seed formation was observed. Up to 0.6 and 1.4 U of phytase activity per mg of total extracted cellular proteins were obtained in germinated transgenic rice seeds expressing appA and SrPf6 phytases, respectively, which represent 46–60 times of phytase activities compared to the non-transformant. The appA and SrPf6 phytases produced in germinated transgenic rice seeds had high activity over broad pH ranges of 3.0–5.5 and 2.0–6.0, respectively. Phytase levels and inheritance of transgenes in one highly expressing plant were stable over four generations. Germinated transgenic rice seeds, which produce a highly active recombinant phytase and are rich in hydrolytic enzymes, nutrients and minerals, could potentially be an ideal feed additive for improving the phytate-phosphorus digestibility in monogastric animals.
Similar content being viewed by others
References
Austin S, Bingham ET, Koegel RG, Mathews DE, Shahan MN and Straub RJ (1994) An overview of a feasibility study for the production of industrial enzymes in transgenic alfalfa. In: Bajpai RK and Prokop A (eds), Recombinant DNA Technology II. (pp. 234–244) New York Academy of Sciences, New York.
Bae HD and Cheng KJ (2000) Production of enzyme products and raw feed materials using grain seeds. US Patent No. 6,063,431.
Bewley JD and Black M (1994) Mobilization of stored seeds reserves. In: Bewley JD and Black M (eds), Seeds Physiology of Development and Germination. Chapter 7 (pp. 293–343) Plenum Press, New York.
Brinch-Pedersen H, Olesen A, Rasmussen SK and Holm PB (2000) Generation of transgenic wheat (Tritivum aestivum L.) for constitutive accumulation of an Aspergillus phytase. Mol Breeding 6: 195–206.
Brinch-Pedersen H, Sorensen LD and Holm PB (2002) Engineering crop plants: getting a handle on phosphate. Trends Plant Sci 7: 118–125.
Chan MT, Chang HH, Ho SL, Tong WF and Yu SM (1993) Agrobacterium-mediated production of transgenic rice plants expressing a chimeric α-amylase promoter/β-glucuronidase gene.Plant Mol Biol 22: 491–506.
Chen PW, Lu CA, Yu TS, Tseng TH, Wang CS and Yu SM (2002) Rice α-amylase transcriptional enhancers direct multiple mode regulation of promoters in transgenic rice. J Biol Chem 277: 13641–13649.
Cheng KJ, Sellnger LB, Yanke LJ, Bae HD, Zhou L and Forsberg CW (1999) Phytases of ruminal microorganisms. US Patent No. 5,939,303.
Dassa J, Marck C and Boquet PL (1990) The complete nucleotide sequence of the Escherichia coli gene appA reveals significant homology between pH 2.5 acid phosphatase and glucose-1-phosphatase. J Bacteriol 172: 5497–5500.
Denbow DM, Grabau EA, Lacy GH, Kornegay ET, Russell DR and Umbeck PF (1998) Soybeans transformed with a fungal phytase gene improve phosphorus availability for broilers. Poult Sci 77: 878–881.
Golovan S, Wang G, Zhang J and Forsberg CW (2000) Characterization and overproduction of the Escherichia coli appA encoded bifunctional enzyme that exhibits both phytase and acid phosphatase activities. Can J Microbiol 46: 59–71.
Greiner R, Konietzny U and Jany K (1993) Purification and characterization of two phytases from Escherichia coli. Arch Biochem Biophys 303: 107–113.
Hajdukiewicz P, Svab Z and Maliga P (1994) The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol Biol 25: 989–994.
Hiei Y, Ohta S, Komari T and Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6: 271–282.
Hood EE, Helmer GL, Fraley RT and Chilton MD (1986) The hypervirulence of Agrobacterium tumefaciens A281 is encoded in a region of pTiBo542 outside of T-DNA. J Bacteriol 168: 1291–1301.
Howson SJ and Davis RP (1983) Production of phytatehydrolysing enzyme by some fungi. Enzyme Microb Technol 5: 377–382.
Itoh K, Yamaguchi J, Huang N, Rodriguez RL, Akazawa T and Shimamoto K (1995) Developmental and hormonal regulation of rice α-amylase (RAmy1A)–gusA fusion genes in transgenic rice seeds. Plant Physiol 107: 25–31.
Jacobsen JV, Gubler F and Chandler PM (1995) Gibberellin action in germinated cereal grains. In: Davies PJ (ed) Plant Hormones: Physiology, Biochemistry, and Molecular Biology. (pp. 246–271) Kluwer Academic Publishers, Dordrecht.
Lee CC, Li CC and Sung FJM (1986) Physiological and genetic studies on seedling vigor in rice (Oryza sativa L.). II. Inheritances of α-amylase activity and seedling vigor in rice. J Agric Assoc China 135: 17–24.
Li J, Hegeman CE, Hanlon RW, Lacy GH, Denbow MD and Grabau EA (1997) Secretion of active recombinant phytase from soybean cell-suspension cultures. Plant Physiol 114: 1103–1111.
Lucca P, Hurrell R and Potrykus I (2001) Genetic engineering approaches to improve the bioavailability and the level of iron in rice grains. Theor Appl Genet 102: 392–397.
Maenz DD (2001) Enzymatic characterisation of phytases as they relate to their use in animal feeds. In: Bedford MR and Partridge GG (eds), Enzymes in Farm Animal Nutrition. (pp. 61–84) CAB International, Walling ford, UK.
Pen J, Verwoerd T C, van Paridin PA, Beukeder RF, van der Elzen PJM, Geerse K et al. (1993) Phytase-containing transgenic seed as a novel feed additive for improved phosphorus utilization.Bio/Technol 11: 811–814.
Ponstein AS, Bade JB, Verwoerd TC, Molendijk L, Storms J, Beudeker RF et al. (2002) Stable expression of Phytase (phyA) in canola (Brassica napus) seeds: towards a commercial product.Mol Breeding 10: 31–44.
Ravindran V, Bryden WL and Kornegay ET (1995) Phytates: occurrence, bioavailability and implications in poultry nutrition. Poult Avain Bio Rev 6: 125–143.
Sasahara T, Ikarashi H and Kambayashi M (1986) Genetic variations in embryo and endosperm weight, seedling growth parameters and α-amylase activity of the germinated grains in rice (Oryza sativa L.). Jpn J Breed 36: 248–261.
Sheu JJ, Yu TS, Tong WF and Yu SM (1996) Carbohydrate starvation stimulates differential expression of rice α-amylase genes that is modulated through complicated transcriptional and posttranscriptional processes. J Biol Chem 271: 26998–27004.
Shimizu M (1992) Purification and characterization of phytase from Bacillus subtilis (natto) N-77. Biosci Biotechnol Biochem 56: 1266–1269.
Thomas PS (1983) Hybridization of denatured RNA transferred or dotted nitrocellulose paper. Meth Enzymol 100: 255–266.
Toki S (1997) Rapid and efficient Agrobacterium-mediated transformation in rice. Plant Mol Biol Rep 15: 16–21.
Ullah AH, Sethumadhavan K, Mullaney EJ, Ziegelhoffer T and Austin-Phillips S (1999) Characterization of recombinant fungal phytase (phyA) expressed in tobacco leaves. Biochem Biophys Res Commun 264: 201–206.
Ullah AH, Sethumadhavan K, Mullaney EJ, Ziegelhoffer T and Austin-Phillips S (2002) Cloned and expressed fungal phyA gene in alfalfa produces a stable phytase. Biochem Biophys Res Commun 290: 1343–1348.
Van Gorcom RFM, van Hartingsveldt W, van Paridon PA, Veenstra AE, Luiten RGM and Selten GCM (1995) Cloning and expression of phytase from Aspergillus. US Patent No. 5,436,156.
Verwoerd TC, van Paridon PA, van Ooyen AJ, van Lent JW, Hoekema A and Pen J (1995) Stable accumulation of Aspergillus niger phytase in transgenic tobacco leaves. Plant Physiol 109: 1199–1205.
Williams JF and Peterson ML (1973) Relations between α-amylase activity and growth of rice seedlings. Crop Sci 13: 612–615.
Yanke LJ, Bae HD, Selinger LB and Cheng KJ (1998) Phytase activity of anaerobic ruminal bacteria. Microbiology 144: 1565–1573.
Yu SM, Kuo YH, Sheu G, Sheu YJ and Liu LF (1991) Metabolic derepression of α-amylase gene expression in suspensioncultured cells of rice. J Biol Chem 266: 21131–21137.
Yu SM, Lee YC, Fang SC, Chan MT, Hwa SF and Liu LF (1996) Sugars act as signal molecules and osmotica to regulate the expression of α-amylase genes and metabolic activities in germinating cereal grains. Plant Mol Biol 30: 1277–1289.
Zhang ZB, Kornegay ET, Radcliffe JS, Wilson JH and Veit HP (2000) Comparison of phytase from genetically engineered Aspergillus and canola in weanling pig diets. J Anim Sci 78: 2868–2878.
Zheng Z, Hayashomoto A, Li Z and Murai N (1991) Hygromycin resistance gene cassettes for vector construction and selection of transformed rice protoplasts. Plant Physiol 97: 832–835.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hong, CY., Cheng, KJ., Tseng, TH. et al. Production of two Highly Active Bacterial Phytases with Broad pH Optima in Germinated Transgenic Rice Seeds. Transgenic Res 13, 29–39 (2004). https://doi.org/10.1023/B:TRAG.0000017158.96765.67
Issue Date:
DOI: https://doi.org/10.1023/B:TRAG.0000017158.96765.67