Abstract
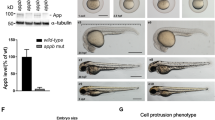
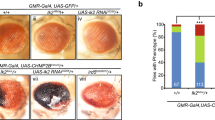
Presenilin proteins have been implicated both in developmental signalling by the cell-surface protein Notch and in the pathogenesis of Alzheimer's disease. Loss of presenilin function leads to Notch/lin-12-like mutant phenotypes in Caenorhabditis elegans1,2 and to reduced Notch1 expression in the mouse paraxial mesoderm3. In humans, presenilins that are associated with Alzheimer's disease stimulate overproduction of the neurotoxic 42-amino-acid β-amyloid derivative (Aβ42) of the amyloid-precursor protein APP4. Here we describe loss-of-function mutations in the Drosophila Presenilin gene that cause lethal Notch-like phenotypes such as maternal neurogenic effects during embryogenesis, loss of lateral inhibition within proneural cell clusters, and absence of wing margin formation. We show that presenilin is required for the normal proteolytic production of carboxy-terminal Notch fragments that are needed for receptor maturation and signalling, and that genetically it acts upstream of both the membrane-bound form and the activated nuclear form of Notch. Our findings provide evidence for the existence of distinct processing sites or modifications in the extracellular domain of Notch. They also link the role of presenilin in Notch signalling to its effect on amyloid production in Alzheimer's disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Levitan, D. & Greenwald, I. Facilitation of lin-12-mediated signalling by SEL-12, a Caenorhabditis elegans S182 Alzheimer's disease gene. Nature 377, 351–354 (1995).
Levitan, D. & Greenwald, I. Effects of SEL-12 presenilin on LIN-12 localization and function in Caenorhabditis elegans. Development 125, 3599–3603 (1998).
Wong, P. C. et al. Presenilin 1 is required for Notch1 and DII1 expression in the paraxial mesoderm. Nature 387, 288–292 (1997).
Haass, C. Presenilins: genes for life and death. Neuron 18, 687–690 (1997).
Schweisguth, F. & Posakony, J. W. Suppressor of Hairless, the Drosophila homolog of the mouse recombination signal-binding protein gene, controls sensory organ cell fates. Development 120, 1433–1441 (1994).
Fortini, M. E. & Artavanis-Tsakonas, S. The Suppressor of Hairless protein participates in Notch receptor signaling. Cell 79, 273–282 (1994).
Diaz-Benjumea, F. J. & Cohen, S. M. Serrate signals through Notch to establish a Wingless-dependent organizer at the dorsal/ventral compartment boundary of the Drosophila wing. Development 121, 4215–4225 (1995).
Kim, J., Irvine, K. D. & Carroll, S. B. Cell recognition, signal induction, and symmetrical gene activation at the dorsal–ventral boundary of the developing Drosophila wing. Cell 82, 795–802 (1995).
Couso, J. P., Knust, E. & Martinez-Arias, A. Serrate and wingless cooperate to induce vestigial gene expression and wing formation in Drosophila. Curr. Biol. 5, 1437–1448 (1995).
Rulifson, E. J. & Blair, S. S. Notch regulates wingless expression and is not required for reception of the paracrine wingless signal during wing margin neurogenesis in Drosophila. Development 121, 2813–2824 (1995).
Kim, J. et al. Integration of positional signals and regulation of wing formation and identity by Drosophila vestigial gene. Nature 382, 133–138 (1996).
Cubas, P., de Celis, J.-F., Campuzano, S. & Modelell, J. Proneural clusters of achaete-scute expression and the generation of sensory organs in the Drosophila imaginal wing disc. Genes Dev. 5, 996–1008 (1991).
Dietrich, U. & Campos-Ortega, J. A. The expression of neurogenic loci in imaginal epidermal cells of Drosophila melanogaster. J. Neurogenet. 1, 315–332 (1984).
Fehon, R. G., Johansen, K., Rebay, I. & Artavanis-Tsakonas, S. Complex cellular and subcellular regulation of Notch expression during embryonic and imaginal development of Drosophila: implications for Notch function. J. Cell. Biol. 113, 657–669 (1991).
Kooh, P. J., Fehon, R. G. & Muskavitch, M. A. T. Implications of dynamic patterns of Delta and Notch expression for cellular interactions during Drosophila development. Development 117, 493–507 (1993).
Blaumueller, C. M., Qi, H., Zagouras, P. & Artavanis-Tsakonas, S. Intracellular cleavage of Notch leads to a heterodimeric receptor on the plasma membrane. Cell 90, 281–291 (1997).
Logeat, F. et al. The Notch1 receptor is cleaved constitutively by a furin-like convertase. Proc. Natl Acad. Sci. USA 95, 8108–8112 (1998).
Chan, Y.-M. & Jan, Y. N. Roles for proteolysis and trafficking in Notch maturation and signal transduction. Cell 94, 423–426 (1996).
Kopan, R., Schroeter, E. H., Weintraub, H. & Nye, J. S. Signal transduction by activated mNotch: importance of proteolytic processing and its regulation by the extracellular domain. Proc. Natl Acad. Sci. USA 93, 1683–1688 (1996).
Luo, B., Aster, J. C., Hasserjian, R. P., Kuo, F. & Sklar, J. Isolation and functional analysis of a cDNA for human Jagged2, a gene encoding a ligand for the Notch1 receptor. Mol. Cell. Biol. 17, 6057–6067 (1997).
Kidd, S., Lieber, T. & Young, M. W. Ligand-induced cleavage and regulation of nuclear entry of Notch in Drosophila melanogaster embryos. Genes Dev. 12, 3728–3740 (1998).
Pan, D. & Rubin, G. M. Kuzbanian controls proteolytic processing of Notch and mediates lateral inhibition during Drosophila and vertebrate neurogenesis. Cell 90, 271–280 (1997).
Struhl, G. & Adachi, A. Nuclear access and action of Notch in vivo. Cell 93, 649–660 (1998).
Lecourtois, M. & Schweisguth, F. Indirect evidence for Delta-dependent intracellular processing of Notch in Drosophila embryos. Curr. Biol. 8, 771–774 (1998).
Schroeter, E. H., Kisslinger, J. A. & Kopan, R. Notch-1 signalling requires ligand-induced proteolytic release of intracellular domain. Nature 393, 382–386 (1998).
Fehon, R. G. et al. Molecular interactions between the protein products of the neurogenic loci Notch and Delta, two EGF-homologous genes in Drosophila. Cell 61, 523–534 (1990).
Struhl, G., Fitzgerald, K. & Greenwald, I. Intrinsic activity of the Lin-12 and Notch intracellular domains in vivo. Cell 74, 331–345 (1993).
Rebay, I., Fehon, R. G. & Artavanis-Tsakonas, S. Specific truncations of Drosophila Notch define dominant activated and dominant negative forms of the receptor. Cell 74, 319–329 (1993).
Selkoe, D. J. Amyloid β-protein and the genetics of Alzheimer's disease. J. Biol. Chem. 271, 18295–18298 (1996).
De Strooper, B. et al. Deficiency of presenilin-1 inhibits the normal cleavage of amyloid precursor protein. Nature 391, 387–390 (1998).
Acknowledgements
We thank V. V. Roussakova for technical assistance with genetic screening and Drosophila microinjection; F. Schweisguth, G. Struhl, S. Artavanis-Tsakonas, D. Pan and R. Finkelstein for fly stocks; and S. Artavanis-Tsakonas, G. M. Rubin, Y. N. Jan and T. A. Jongens for antibodies. This work was supported by the NIH, the Life and Health Insurance Medical Research Fund, and the Alzheimer's Association.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ye, Y., Lukinova, N. & Fortini, M. Neurogenic phenotypes and altered Notch processing in Drosophila Presenilin mutants. Nature 398, 525–529 (1999). https://doi.org/10.1038/19096
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/19096
This article is cited by
-
Ubiquitylation is required for the incorporation of the Notch receptor into intraluminal vesicles to prevent prolonged and ligand-independent activation of the pathway
BMC Biology (2022)
-
Notch signaling pathway: architecture, disease, and therapeutics
Signal Transduction and Targeted Therapy (2022)
-
Structural characterization of plasmodial aminopeptidase: a combined molecular docking and QSAR-based in silico approaches
Molecular Diversity (2019)
-
Non-canonical Notch signaling represents an ancestral mechanism to regulate neural differentiation
EvoDevo (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.