Abstract
Pseudoachondroplasia (PSACH) and autosomal dominant multiple epiphyseal dysplasia (MED) are chondrodysplasias resulting in short-limbed dwarfism, joint pain and stiffness and early onset osteoarthritis. All PSACH, and the largest proportion of MED, result from mutations in cartilage oligomeric matrix protein (COMP). The first mutations in COMP were identified in 1995 in patients with both PSACH and MED and subsequently there has been over 30 publications describing COMP mutations in at least 250 PSACH–MED patients. However, despite these discoveries, a methodical analysis of the relationship between COMP mutations and phenotypes has not been undertaken. In particular, there has, to date, been little correlation between the type and location of a COMP mutation and the resulting phenotype of PSACH or MED. To determine if genotype to phenotype correlations could be derived for COMP, we collated 300 COMP mutations, including 25 recently identified novel mutations. The results of this analysis demonstrate that mutations in specific residues and/or regions of the type III repeats of COMP are significantly associated with either PSACH or MED. This newly derived genotype to phenotype correlation may aid in determining the prognosis of PSACH and MED, including the prediction of disease severity, and in the long term guide genetic counselling and contribute to the clinical management of patients with these diseases.
Similar content being viewed by others
Introduction
Pseudoachondroplasia (PSACH) and multiple epiphyseal dysplasia (MED) are relatively common chondrodysplasias resulting in joint pain and stiffness, short-limbed dwarfism and in many cases early onset osteoarthritis.1 PSACH results exclusively from mutations in cartilage oligomeric matrix protein (COMP),2 a large pentameric glycoprotein found in cartilage, tendon, ligament and skeletal muscle.3 In contrast, autosomal dominant MED is genetically heterogeneous and although in the European population the largest proportion results from COMP mutations, other forms of MED can be caused by mutations in the genes encoding matrilin-3 (MATN3) and type IX collagen (COL9A1, COL9A2 and COL9A3).2, 4 Both of these proteins are known to interact with COMP in the cartilage extracellular matrix and help coordinate collagen fibrillogenesis. Autosomal recessive MED appears to result exclusively from mutations in sulphate transporter solute carrier family 26.2
Despite consistent genotype to phenotype correlations across the various genes; that is, COMP (PSACH or MED), MATN3 (MED only) and type IX collagen (MED only),2 there has been no systematic investigation of the relationship between COMP mutations and phenotype (PSACH or MED). In particular, there has been no study on the type and location of a COMP mutation and the resulting phenotype.
To address this omission, we collated a comprehensive list of COMP mutations and the resulting phenotypes from 300 individual case reports that were published between 1995 and 2012 (n=260) or recently identified as part of our diagnostic service for PSACH and MED (n=40). The reported mutations were from 32 different publications, although the majority (n=183) had been described in seven articles, and our unreported cases formed the third largest cohort and included 25 novel COMP mutations.2, 4, 5, 6, 7, 8, 9 To promote the clinical utility of any genotype to phenotype correlations and provide a realistic appreciation of the clinical-diagnostic process, we recorded the phenotypes as originally reported, without any further review, which would provide a more robust model should significant correlations be identified.
Materials and methods
Mutation analysis
For the novel mutations reported in this study, mutational analysis of the COMP gene was performed as previously described.5 Briefly, bidirectional fluorescent sequence analysis was used to screen for mutations in exons 8–19 of COMP including the splice donor and acceptor sites. COMP nomenclature is according to Genebank Accession number NM_000095.2 with nucleotide 1 as the first nucleotide of translation. Mutations are accessible in the Human Mutation Database and Leiden Open Variation Database.
Statistical analysis
The Fisher exact test was used to test the following null hypotheses:-
-
1
That no association exists between the location of a mutation within the T3 repeats of COMP and the frequency of PSACH versus MED diagnosis; that is, the frequency of PSACH versus MED missense mutations reported for each T3 repeat was compared with the total frequency of PSACH versus MED mutations reported in all other COMP T3 repeats (Supplementary Table 2).
-
2
That no association exists between the location of a mutation within the N- versus C- type motifs of the T3 repeats and the frequency of PSACH versus MED diagnosis.
In all statistical analyses, cases in which mutations did not lead to a defined MED or PSACH diagnosis were excluded and the null hypothesis was rejected upon calculation of a P-value <0.05.
Results
Domain-specific distribution of COMP mutations
In the first instance, we examined the domain-specific locations of the 300 mutations (Table 1 and Supplementary Table 1). Three putative mutations (1%) were identified in the type II (EGF-like) repeat domain (T2-COMP), 269 mutations (90%) in the type III repeat domain (T3-COMP) and 28 mutations (9%) in the carboxyl-terminal domain (CTD-COMP), thereby confirming that both PSACH and MED mutations are predominantly located within the type III repeat domain of COMP.
Missense mutations in the type II repeats of COMP have unresolved pathogenicity
Recently, putative missense mutations in three of the four type 2 (EGF-like) domains have been identified (c.500G>A p.(Gly167Glu)), (c.700C>T p.(Pro234Ser)) and (c.772G>C p.(Gly258Arg)) (Table 1; Supplementary Table 1 and 2); however, the scarcity of these mutations and their unresolved pathogenicity makes any correlations of limited clinical value, although they do appear to cause a range of phenotypes within the MED to mild PSACH disease spectrum, but without any distinguishing features.2
Missense mutations in the type III repeats are the major cause of PSACH & MED and show significant phenotypic correlations
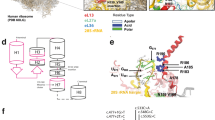
The type III repeat region of COMP comprises of amino acid residues 268–528 (n=261) arranged into eight consecutive T3 repeats typified by a consensus motif that is characteristic of a calcium-binding pocket (Figure 1). Six of the T3 repeats (T31, T32, T34, T36, T37 and T38) have both N- and C-type motifs, whereas T33 and T35 only have a single C-type motif. In addition to the repetitive nature of this domain, it is also composed of a high proportion of specific residues such aspartic acid, glycine, cysteine, asparagine and proline that have important roles in protein folding and calcium binding.
An illustration of the type III repeat region of COMP and the location of missense mutations and in-frame deletions, insertions or indels that lead to PSACH and/or MED. Each of the eight type III repeats are indicated on the left (T31, T32 etc.) along with the number of the COMP residue at the start of each repeat. The N- and C-type motifs are shown at the top and the residues that comprise each motif are boxed. The consensus sequence for the N- and C-type motifs are shown below. Residues that have missense mutations which cause MED are coloured GREEN, those that cause PSACH are coloured RED and mutations reported to cause both PSACH and MED are coloured BLUE. In-frame deletions, insertions or indels are double underlined using the same PSACH–MED colour scheme. The number of patients with a missense mutation identified at each codon is indicated above the relevant residue. Missense mutations in blocks of COMP residues that cause MED are highlighted in blue and those that cause PSACH are highlighted in green.
In total, 179 missense mutations have been identified in 73 of the 261 codons (28%) that comprise the T3 domain. Fifty three of these 73 codons (73%) encode amino acid residues that comprise the consensus motifs of the individual repeats, thereby confirming the importance of these residues for the correct folding and/or functioning of this domain. Of the remaining 20 codons, 11 encode residues such as cysteine, aspartic acid, proline and glycine, which also have potentially important roles in protein folding. Overall, this clustering suggests that the primary sequence of COMP is very highly restrained; indeed, only a single non-synonymous polymorphism (c.1156A>G p.(Asn386Asp)) has been regularly identified in the type III repeat region. We therefore hypothesised that the large number of missense mutations and high degree of amino acid similarity within the individual repeats would provide an ideal opportunity to derive, for the first time, phenotype to genotype correlations for PSACH and MED.
We firstly determined that there was no significant association between the incidence of MED and PSACH causing missense mutations and whether the mutation occurred within the N- or C-type motifs of the T3 repeats of COMP (P=0.403). We next compared the frequency of PSACH versus MED missense mutations in each of the T3 repeats, to determine whether a mutation within a given T3 repeat is more associated with PSACH or MED (Supplementary Table 2).
There was no significant association between phenotype and mutation in T31 (P=0.0524), with 14 PSACH mutations compared with five MED mutations. Likewise, mutations in the T32 (P=0.32) and T33 (P=1.00) repeats also showed no significant association with a specific phenotype.
In contrast, T34 showed a very significant association with MED compared with the other T3 repeats (P<0.0001). Indeed, it was obvious that (c.1153G>A p.(Asp385Asn)) that had been identified in 14 patients with MED was influencing this association. However, it was also clear that a contiguous stretch of MED-associated mutations were present in both the N- and C-type motifs of the T34 repeat and of the 33 mutations in nine codons, 27 were reported to have caused MED, whereas the mutation of aspartic acid at residue 376 was associated with MED in three cases and a single PSACH patient. Finally, (c.1159T>G p.(Cys387Gly)) and (c.1159T>C p.(Cys387Arg)) caused PSACH in two cases, which was comparable to the substitution of a cysteine residue at equivalent locations in the C-type motifs of the other T3 repeats.
Similarly, mutations in T35 were also very significantly associated with a greater frequency of MED compared with mutations in other T3 repeats (P=0.0009). For example, mutations in eight of the nine codons are associated with MED (n=13 cases), whereas (c.1229G>A p.(Cys410Tyr)) appears to cause the borderline phenotype of MED or mild PSACH (n=3).
Missense mutations in T36, T37 and T38 were significantly associated with a greater frequency of PSACH compared with mutations in other T3 repeats (P=0.0156, P=0.0097, P=0.0384, respectively), which therefore identified this region in COMP as ‘PSACH-specific’. This significant association did not even take account of the common PSACH (c.1417_1419delGAC p.(D469del)) mutation (present in 16% of all patients in this study), but it was clear that missense mutations at several key residues were contributing to this association. These included various mutations affecting the following residues: glycine 440 (n=9); aspartic acid 473 and asparagine 518 (n=6 each); glycine 465 (n=5 each); glycine 446, aspartic acid 509 and aspartic acid 511 (n=4 each), which all caused PSACH. Interestingly, different nucleotide transversions or transitions at codons 465, 473, 482 and 518 individually caused three or four different amino acid substitutions at each codon (Table 2).
Missense mutations in CTD show distinct clustering in functional motifs
CTD-COMP mutations were reported in 28 cases of PSACH and MED and therefore represent a minor cause of these diseases (<10%). However, these are still functionally relevant mutations and as previously reported, they cluster in distinct regions of this domain.10 In particular, the mutation of codons 583, 585 and 587 define the first region, whereas mutation of codons 718 and 719 define a second region. Furthermore, (c.2153G>C p.(Arg718Pro)) and (c.2152C>T p.(Arg718Trp)), which always causes MED, were reported in 10/28 cases (∼36%), whereas substitution of the neighbouring residue (c.2155G>A p.(Arg719Ser)) and (c.2156G>A p.(Arg719Asp)) causes PSACH (n=3; ∼11%). Interestingly, the other common CTD-COMP mutation at residue 585 (c.1754C>T p.(Thr585Met); c.1754C>G p.(Thr585Arg) and c.1754C>A p.(Thr585Lys); n=6; ∼21%) often causes a transitional phenotype of mild PSACH to severe MED9 and the association of mild myopathy with CTD-COMP mutations is well documented.3
In-frame deletions, insertions and indels are significantly associated with PSACH
Regardless of the specific COMP genomic DNA mutation, all the deletions, insertions and indels cause in-frame alterations to the COMP protein primary sequence and no frame shifts have been identified with one exception,11 which we did not include in this study as our focus was on qualitative protein defects. In total, there were 90 (30%) instances of these classes of mutations and therefore, it represented significant cause of abnormal COMP synthesis.
The majority of in-frame deletions, insertions or indels lead to PSACH (n=74; 82%), whereas a smaller proportion causes MED (n=16; 18%); however, in several instances, the same mutation was reported to cause both PSACH and MED. The high incidence of these classes of mutations is influenced by the deletion of a single aspartic acid residue (c.1417_1419delGAC p.(Asp473del)) from a contiguous stretch of five aspartic acid codons (GAC) encoding residues 469 to 473 of COMP, which has been reported in numerous studies to be a common mutation in PSACH. This observation was further corroborated by this study, which confirmed that (c.1417_1419delGAC p.(Asp473del)) accounted for 49 cases of PSACH and was the most common mutation in the entire series (16%).
Interestingly, more complex mutations are also found within this repeated region and includes the deletion of either two or three GAC codons (c.1414_1419delGACGAC p.(Asp472_Asp473del)) and (c.1411_1419delGACGACGAC p.(Asp471_Asp473del)) that cause PSACH (n=3), the duplication of single GAC codon (c.1417_1419dup p.(Asp473dup)) that consistently causes MED (n=4) and the duplication of two GAC codons (c.1414_1419dup p.(Asp472_Asp473dup)) that causes PSACH (n=2).
Other notable ‘hot-spots’ for these classes of mutations include (c.1120_1122del p.(Asp374del)) or (c.1117_1122del p.(Asp373_Asp374del)) (n=5), (c.1021_1026del p.(Glu341_Asp342del)) (n=4) and (c.1170_1181delinsTGT p.(Pro391_Asp394delinsVal)) (n=2), which most often cause PSACH, with the exception of two cases reported as MED by Kim et al 2011.4 Interestingly, (c.1371_1373del p.(Glu457del)) appears to result equally in PSACH (n=2) or MED (n=3), although a revaluation of the available clinical information would suggest that presentation is at the severe end of the MED spectrum. The remaining 15 cases of this type of mutation have only been reported in single cases and can result in a range of phenotypes within the PSACH and MED disease spectrum, although the majority cause PSACH (n=10; ∼67%).
Discussion
Even though the first COMP mutations were identified in PSACH and MED patients nearly 20 years ago, there has been no comprehensive genotype to phenotype correlation derived to date. The compilation and analysis of 300 cases in this study has allowed us to redress this oversight and to identify some important correlations that may assist in genetic counselling and disease prognosis.
The large number of missense mutations that have been identified in the type III repeats allowed us to derive the first genotype to phenotype correlation for COMP and this provides strong evidence for both PSACH- and MED-associated regions. For example, it was clear that a missense mutation in the residues encompassing T34 and T35 was more likely to result in MED, whereas mutations in T36–T38 were generally associated with PSACH. One notable correlation was the substitution of cysteine residues comprising the invariant position 14 of the C-type motif consensus sequence that causes PSACH (Cys328, n=2; Cys351, n=3; Cys387, n=2; Cys448, n=1; Cys484, n=1) in all but one position (Cys410, n=3). This latter finding highlights the potential importance of these cysteine residues to the folding of the C-type motif of COMP.
It is clear that the vast majority (>80%) of in-frame deletions, insertions or indels cause PSACH because these mutations are presumably more likely to cause a greater disruption to the folding and tertiary structure of the COMP protein. The high incidence of in-frame deletions, insertions or indels in the type III repeat region of COMP is most likely due to the repetitive nature of the DNA within the exons encoding this protein domain. To date, no in-frame deletions, insertions or indels have been identified in the carboxyl-terminal globular domain.
The deletions and insertions of Asp473 and the neighbouring residues are themselves an interesting group of mutations. p.(Asp473del) is the most common COMP mutation (n=49/300) and always causes severe PSACH, along with the deletion of either two or three GAC codons (p.(Asp472_Asp473del) and p.(Asp471_Asp473del)) (n=3/300). In contrast, the duplication of a single GAC codon p.(Asp473dup) consistently causes MED (n=4/300), whereas the duplication of two GAC codons p.(Asp472_Asp473dup) causes PSACH (n=2/300). These correlations in phenotypic presentation suggest that the deletion of a single aspartic acid residue disrupts the consensus calcium-binding pocket of the C-type motif of the T36 repeat to a greater extent than the insertion of a single aspartic acid. However, the insertion of two aspartic acid residues again has a more detrimental effect on calcium binding and protein folding, which are not mutually exclusive. It has previously been proposed that these insertions are examples of trinucleotide expansion mutations,12 but there is no evidence of genetic anticipation in PSACH–MED through an increased expansion of the GAC triplets and they are likely to be self-limiting mutations.
For 10 missense mutations and a single deletion that had been identified in more than one patient each, the phenotype had been recorded as either PSACH or MED. The discrepancies in diagnosis reflect the challenge in diagnosing these diseases and highlights the phenotypic overlap between MED and mild PSACH.13 It remains highly possible that genetic modifiers of phenotypic severity are likely to influence disease severity, which has already been reported for MED caused by MATN3 mutations.14
In conclusion, we have collated 300 COMP mutations that cause PSACH and MED and have derived the first genotype to phenotype correlations. Although there are significant associations with some residues, particularly those with >3 reported mutations, others are less clear, and extra factors (such as diagnostic interpretation and genetic modifiers) may have an influence. Nevertheless, this study provides a first framework that may help medical professionals in predicting phenotype and/or disease severity.
Accession codes
References
Briggs MD, Hoffman SM, King LM et al: Pseudoachondroplasia and multiple epiphyseal dysplasia due to mutations in the cartilage oligomeric matrix protein gene. Nat Genet 1995; 10: 330–336.
Jackson GC, Mittaz-Crettol L, Taylor JA et al: Pseudoachondroplasia and multiple epiphyseal dysplasia: A 7-year comprehensive analysis of the known disease genes identify novel and recurrent mutations and provides an accurate assessment of their relative contribution. Hum Mutat 2012; 33: 144–157.
Pirog KA, Briggs MD : Skeletal dysplasias associated with mild myopathy-a clinical and molecular review. J Biomed Biotechnol 2010; 2010: 686457.
Kim OH, Park H, Seong MW et al: Revisit of multiple epiphyseal dysplasia: ethnic difference in genotypes and comparison of radiographic features linked to the COMP and MATN3 genes. Am J Med Genet A 2011; 155A: 2669–2680.
Kennedy J, Jackson G, Ramsden S et al: COMP mutation screening as an aid for the clinical diagnosis and counselling of patients with a suspected diagnosis of pseudoachondroplasia or multiple epiphyseal dysplasia. Eur J Hum Genet 2005; 13: 547–555.
Mabuchi A, Manabe N, Haga N et al: Novel types of COMP mutations and genotype-phenotype association in pseudoachondroplasia and multiple epiphyseal dysplasia. Hum Genet 2003; 112: 84–90.
Deere M, Sanford T, Francomano CA, Daniels K, Hecht JT : Identification of nine novel mutations in cartilage oligomeric matrix protein in patients with pseudoachondroplasia and multiple epiphyseal dysplasia. Am J Med Genet 1999; 85: 486–490.
Ikegawa S, Ohashi H, Nishimura G et al: Novel and recurrent COMP (cartilage oligomeric matrix protein) mutations in pseudoachondroplasia and multiple epiphyseal dysplasia. Hum Genet 1998; 103: 633–638.
Briggs MD, Mortier GR, Cole WG et al: Diverse mutations in the gene for cartilage oligomeric matrix protein in the pseudoachondroplasia-multiple epiphyseal dysplasia disease spectrum. Am J Hum Genet 1998; 62: 311–319.
Kennedy J, Jackson GC, Barker FS et al: Novel and recurrent mutations in the C-terminal domain of COMP cluster in two distinct regions and result in a spectrum of phenotypes within the pseudoachondroplasia — multiple epiphyseal dysplasia disease group. Hum Mutat 2005; 25: 593–594.
Mabuchi A, Haga N, Ikeda T et al: Novel mutation in exon 18 of the cartilage oligomeric matrix protein gene causes a severe pseudoachondroplasia. Am J Med Genet 2001; 104: 135–139.
Delot E, King LM, Briggs MD, Wilcox WR, Cohn DH : Trinucleotide expansion mutations in the cartilage oligomeric matrix protein (COMP) gene. Hum Mol Genet 1999; 8: 123–128.
Rimoin DL, Rasmussen IM, Briggs MD et al: A large family with features of pseudoachondroplasia and multiple epiphyseal dysplasia: exclusion of seven candidate gene loci that encode proteins of the cartilage extracellular matrix. Hum Genet 1994; 93: 236–242.
Jackson GC, Barker FS, Jakkula E et al: Missense mutations in the beta strands of the single A-domain of matrilin-3 result in multiple epiphyseal dysplasia. J Med Genet 2004; 41: 52–59.
Acknowledgements
MDB is the recipient of a Wellcome Trust Senior Research Fellowship in Basic Biomedical Science (Grant 084353/Z/07/Z). We would like to thank Kate Potter for advice on the statistical analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Briggs, M., Brock, J., Ramsden, S. et al. Genotype to phenotype correlations in cartilage oligomeric matrix protein associated chondrodysplasias. Eur J Hum Genet 22, 1278–1282 (2014). https://doi.org/10.1038/ejhg.2014.30
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2014.30
This article is cited by
-
Clinical, Biochemical, Radiological, Genetic and Therapeutic Analysis of Patients with COMP Gene Variants
Calcified Tissue International (2022)
-
A novel COMP mutation in a Chinese family with multiple epiphyseal dysplasia
BMC Medical Genetics (2020)
-
Pathogenic gene screening in 91 Chinese patients with short stature of unknown etiology with a targeted next-generation sequencing panel
BMC Medical Genetics (2018)
-
COMP and TSP-4 interact specifically with the novel GXKGHR motif only found in fibrillar collagens
Scientific Reports (2018)
-
Two novel mutations of COMP in Japanese boys with pseudoachondroplasia
Human Genome Variation (2018)