Abstract
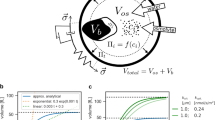
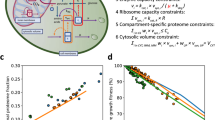
Integration of experimental studies with mathematical modeling allows insight into systems properties, prediction of perturbation effects and generation of hypotheses for further research. We present a comprehensive mathematical description of the cellular response of yeast to hyperosmotic shock. The model integrates a biochemical reaction network comprising receptor stimulation, mitogen-activated protein kinase cascade dynamics, activation of gene expression and adaptation of cellular metabolism with a thermodynamic description of volume regulation and osmotic pressure. Simulations agree well with experimental results obtained under different stress conditions or with specific mutants. The model is predictive since it suggests previously unrecognized features of the system with respect to osmolyte accumulation and feedback control, as confirmed with experiments. The mathematical description presented is a valuable tool for future studies on osmoregulation in yeast and—with appropriate modifications—other organisms. It also serves as a starting point for a comprehensive description of cellular signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Somero, G.N. & Yancey, P.H. in Handbook of Physiology (eds. Hoffmann, J.F. & Jamieson, J.D.) 441–484, (Oxford University Press, Oxford, New York, 1997).
Hohmann, S. Osmotic stress signaling and osmoadaptation in yeasts. Microbiol. Mol. Biol. Rev. 66, 300–372 (2002).
de Nadal, E., Alepuz, P.M. & Posas, F. Dealing with osmostress through MAP kinase activation. EMBO Rep. 3, 735–740 (2002).
Reiser, V., Raitt, D.C. & Saito, H. Yeast osmosensor Sln1 and plant cytokinin receptor Cre1 respond to changes in turgor pressure. J. Cell Biol. 161, 1035–1040 (2003).
Batchelor, E. & Goulian, M. Robustness and the cycle of phosphorylation and dephosphorylation in a two-component regulatory system. Proc. Natl. Acad. Sci. USA 100, 691–696 (2003).
Tyson, J.J., Chen, K. & Novak, B. Network dynamics and cell physiology. Nat. Rev. Mol. Cell Biol. 2, 908–916 (2001).
Schoeberl, B., Eichler-Jonsson, C., Gilles, E.D. & Muller, G. Computational modeling of the dynamics of the MAP kinase cascade activated by surface and internalized EGF receptors. Nat. Biotechnol. 20, 370–375 (2002).
Kitano, H. Computational systems biology. Nature 420, 206–210 (2002).
Kofahl, B. & Klipp, E. Modelling the dynamics of the yeast pheromone pathway. Yeast 21, 831–850 (2004).
Klipp, E., Herwig, R., Kowald, A., Wierling, C. & Lehrach, H. Systems Biology in Practice Concepts, Implementation and Application (Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, 2005).
Levchenko, A., Bruck, J. & Sternberg, P.W. Scaffold proteins may biphasically affect the levels of mitogen-activated protein kinase signaling and reduce its threshold properties. Proc. Natl. Acad. Sci. USA 97, 5818–5823 (2000).
Ferrell, J.E., Jr. Self-perpetuating states in signal transduction: positive feedback, double-negative feedback and bistability. Curr. Opin. Cell Biol. 14, 140–148 (2002).
Llorens, M., Nuno, J.C., Rodriguez, Y., Melendez-Hevia, E. & Montero, F. Generalization of the theory of transition times in metabolic pathways: a geometrical approach. Biophys. J. 77, 23–36 (1999).
Heinrich, R., Neel, B.G. & Rapoport, T.A. Mathematical models of protein kinase signal transduction. Mol. Cell 9, 957–970 (2002).
Huang, C.Y. & Ferrell, J.E., Jr. Ultrasensitivity in the mitogen-activated protein kinase cascade. Proc. Natl. Acad. Sci. USA 93, 10078–10083 (1996).
Bhalla, U.S. & Iyengar, R. Emergent properties of networks of biological signaling pathways. Science 283, 381–387 (1999).
Kholodenko, B.N. Negative feedback and ultrasensitivity can bring about oscillations in the mitogen-activated protein kinase cascades. Eur. J. Biochem. 267, 1583–1588 (2000).
Asthagiri, A.R. & Lauffenburger, D.A. A computational study of feedback effects on signal dynamics in a mitogen-activated protein kinase (MAPK) pathway model. Biotechnol. Prog. 17, 227–239 (2001).
Sedaghat, A.R., Sherman, A. & Quon, M.J. A mathematical model of metabolic insulin signaling pathways. Am. J. Physiol. Endocrinol. Metab. 283, E1084–E1101 (2002).
Hartwell, L.H., Hopfield, J.J., Leibler, S. & Murray, A.W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
O'Rourke, S.M. & Herskowitz, I. Unique and redundant roles for HOG MAPK pathway components as revealed by whole-genome expression analysis. Mol. Biol. Cell 15, 532–542 (2004).
Camisard, V., Brienne, J.P., Baussart, H., Hammann, J. & Suhr, H. Inline characterization of cell concentration and cell volume in agitated bioreactors using in situ microscopy: application to volume variation induced by osmotic stress. Biotechnol. Bioeng. 78, 73–80 (2002).
Albertyn, J., Hohmann, S. & Prior, B.A. Characterization of the osmotic-stress response in Saccharomyces cerevisiae: osmotic stress and glucose repression regulate glycerol-3-phosphate dehydrogenase independently. Curr. Genet. 25, 12–18 (1994).
Saito, H. & Tatebayashi, K. Regulation of the osmoregulatory HOG MAPK cascade in yeast. J. Biochem. 136, 267–272 (2004).
Rep, M., Albertyn, J., Thevelein, J.M., Prior, B.A. & Hohmann, S. Different signalling pathways contribute to the control of GPD1 gene expression by osmotic stress in Saccharomyces cerevisiae. Microbiol. 145, 715–727 (1999).
Van Wuytswinkel, O. et al. Response of Saccharomyces cerevisiae to severe osmotic stress: evidence for a novel activation mechanism of the HOG MAP kinase pathway. Mol. Microbiol. 37, 382–397 (2000).
Ansell, R., Granath, K., Hohmann, S., Thevelein, J.M. & Adler, L. The two isoenzymes for yeast NAD+-dependent glycerol 3-phosphate dehydrogenase encoded by GPD1 and GPD2 have distinct roles in osmoadaptation and redox regulation. EMBO J. 16, 2179–2187 (1997).
Krantz, M. et al. Anaerobicity prepares Saccharomyces cerevisiae cells for faster adaptation to osmotic shock. Eukaryot. Cell 3, 1381–1390 (2004).
Maeda, T., Wurgler-Murphy, S.M. & Saito, H. A two-component system that regulates an osmosensing MAP kinase cascade in yeast. Nature 369, 242–245 (1994).
Hornberg, J.J. et al. Principles behind the multifarious control of signal transduction. ERK phosphorylation and kinase/phosphatase control. FEBS J. 272, 244–258 (2005).
Lu, J.M., Deschenes, R.J. & Fassler, J.S. Saccharomyces cerevisiae histidine phosphotransferase Ypd1p shuttles between the nucleus and cytoplasm for SLN1-dependent phosphorylation of Ssk1p and Skn7p. Eukaryot. Cell 2, 1304–1314 (2003).
Hwang, I. & Sheen, J. Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413, 383–389 (2001).
Tamas, M.J. et al. Fps1p controls the accumulation and release of the compatible solute glycerol in yeast osmoregulation. Mol. Microbiol. 31, 1087–1104 (1999).
Pettersson, H., Filipsson, C., Becit, E., Brive, L. & Hohmann, S. Aquaporins in yeasts and filamentous fungi. Biol. Cell. 97, 487–500 (2005).
Wurgler-Murphy, S.M., Maeda, T., Witten, E.A. & Saito, H. Regulation of the Saccharomyces cerevisiae HOG1 mitogen-activated protein kinase by the PTP2 and PTP3 protein tyrosine phosphatases. Mol. Cell. Biol. 17, 1289–1297 (1997).
Karlgren, S. et al. Conditional osmotic stress in yeast: a system to study transport through aquaglyceroporins and osmostress signaling. J. Biol. Chem. 280, 7186–7193 (2005).
Rizzi, M., Baltes, M., Theobald, U. & Reuss, M. In vivo analysis of metabolic dynamics in Saccharomyces cerevisiae: II. Mathematical model. Biotechnol. Bioeng. 55, 39–54 (1997).
Martinez de Maranon, I., Marechal, P.A. & Gervais, P. Passive response of Saccharomyces cerevisiae to osmotic shifts: cell volume variations depending on the physiological state. Biochem. Biophys. Res. Commun. 227, 519–523 (1996).
Hynne, F., Dano, S. & Sorensen, P.G. Full-scale model of glycolysis in Saccharomyces cerevisiae. Biophys. Chem. 94, 121–163 (2001).
Teusink, B. et al. Can yeast glycolysis be understood in terms of in vitro kinetics of the constituent enzymes? Testing biochemistry. Eur. J. Biochem. 267, 5313–5329 (2000).
Theobald, U., Mailinger, W., Baltes, M., Rizzi, M. & Reuss, M. In vivo analysis of metabolic dynamics in Saccharomyces cerevisiae: I. Experimental observations. Biotechnol. Bioeng. 55, 305–316 (1997).
Tamas, M.J., Rep, M., Thevelein, J.M. & Hohmann, S. Stimulation of the yeast high osmolarity glycerol (HOG) pathway: evidence for a signal generated by a change in turgor rather than by water stress. FEBS Lett. 472, 159–165 (2000).
Gervais, P. & Beney, L. Osmotic mass transfer in the yeast Saccharomyces cerevisiae. Cell Mol. Biol. (Noisy-le-grand) 47, 831–839 (2001).
Gervais, P., Molin, P., Marechal, P.A. & Herail-Foussereau, C. Thermodynamics of yeast cell osmoregulation: passive mechanisms. J. Biol. Phys. 22, 73–86 (1996).
Kedem, O. & Katchalsky, A. Thermodynamic analysis of the permeability of biological membranes to non-electrolytes. Biochim. Biophys. Acta 27, 229–246 (1958).
Caley, S.D., Guttman, H.J. & Record, M.T.J. Biophysical characterization of changes in amounts and activity of Escherichia coli cell and compartment water and turgor pressure in response to osmotic stress. Biophys. J. 78, 1748–1764 (2000).
Zimmermann, U. Physics of turgor- and osmoregulation. Ann. Rev. Plant Physiol. 29, 121–148 (1978).
Reed, R.H., Chudek, J.A., Foster, R. & Gadd, G.M. Osmotic significance of glycerol accumulation in exponentially growing yeasts. Appl. Environ. Microbiol. 53, 2119–2123 (1987).
Acknowledgements
E.K. is supported by the Berlin Center of Genome Based Bioinformatics (BCB), financed by the German Federal Ministry for Education and Research (BMBF, grant 031U109C). B.N. and P.G. are PhD students of the National Research School for Genomics and Bioinformatics, Göteborg, supported by the Swedish Ministry for Education and Research. S.H. holds a research position of the Swedish Research Council. Research in his lab is supported by the European Commission (contracts QLK3-CT2000-00778 and QLK1-CT2001-01066) and the Human Frontier Science Program. Systems Biology of yeast osmoregulation is supported by the European Commission (the QUASI project, contract LSHG-CT2003-503230 to S.H. and E.K.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Standard experiment (one single osmotic shock in wild type cells with 0.5 M NaCl at time zero) used for parameter determination. (PDF 24 kb)
Supplementary Fig. 2
Parameter dependence of the behavior of the MAPK cascade. (PDF 23 kb)
Supplementary Fig. 3
Sensitivity S of the Euclidean distance D between experimental data and simulations with respect to parameter variation. (PDF 36 kb)
Supplementary Fig. 4
Dependence of characteristic system features on parameter values. (PDF 44 kb)
Supplementary Fig. 5
Simulation of osmotic shocks with increasing level of NaCl. (PDF 35 kb)
Supplementary Fig. 6
Open Fps1. (PDF 18 kb)
Supplementary Fig. 7
Yeast cells overproducing glycerol. (PDF 24 kb)
Supplementary Fig. 8
Overproduction of protein phosphatase Ptp2 in wild type cells and cells with unregulated Fps1. (PDF 91 kb)
Supplementary Fig. 9
Profile of intracellular glycerol levels in wild type cells. (PDF 18 kb)
Supplementary Fig. 10
Simulations of enhanced expression of activated alleles of Ssk2 (dark dashed lines) and Hog1 (gray dashed lines). (PDF 15 kb)
Supplementary Table 1
Equations governing the dynamics of the phosphorelay module. (PDF 30 kb)
Supplementary Table 2
Equations governing the dynamics of the MAP kinase cascade. (PDF 36 kb)
Supplementary Table 3
Equations governing the dynamics of transcription and translation. (PDF 26 kb)
Supplementary Table 4
Equations governing the dynamics of the carbohydrate metabolism. (PDF 46 kb)
Supplementary Table 5
Set of equations governing the changes of volume and osmotic pressure. (PDF 60 kb)
Supplementary Table 6
Integrative model of the response of yeast cells to osmotic shock. (PDF 65 kb)
Rights and permissions
About this article
Cite this article
Klipp, E., Nordlander, B., Krüger, R. et al. Integrative model of the response of yeast to osmotic shock. Nat Biotechnol 23, 975–982 (2005). https://doi.org/10.1038/nbt1114
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1114
This article is cited by
-
Role of ultrasensitivity in biomolecular circuitry for achieving homeostasis
Nonlinear Dynamics (2024)
-
Positive feedback induces switch between distributive and processive phosphorylation of Hog1
Nature Communications (2023)
-
Biologia futura: combinatorial stress responses in fungi
Biologia Futura (2022)
-
Intrinsically disordered protein biosensor tracks the physical-chemical effects of osmotic stress on cells
Nature Communications (2021)
-
Rapid and reversible cell volume changes in response to osmotic stress in yeast
Brazilian Journal of Microbiology (2021)