Abstract
Plant microRNAs (miRNAs) regulate the abundance of target mRNAs by guiding their cleavage at the sequence complementary region. We have modified an Arabidopsis thaliana miR159 precursor to express artificial miRNAs (amiRNAs) targeting viral mRNA sequences encoding two gene silencing suppressors, P69 of turnip yellow mosaic virus (TYMV) and HC-Pro of turnip mosaic virus (TuMV). Production of these amiRNAs requires A. thaliana DICER-like protein 1. Transgenic A. thaliana plants expressing amiR-P69159 and amiR-HC-Pro159 are specifically resistant to TYMV and TuMV, respectively. Expression of amiR-TuCP159 targeting TuMV coat protein sequences also confers specific TuMV resistance. However, transgenic plants that express both amiR-P69159 and amiR-HC-Pro159 from a dimeric pre-amiR-P69159/amiR-HC-Pro159 transgene are resistant to both viruses. The virus resistance trait is displayed at the cell level and is hereditable. More important, the resistance trait is maintained at 15 °C, a temperature that compromises small interfering RNA–mediated gene silencing. The amiRNA-mediated approach should have broad applicability for engineering multiple virus resistance in crop plants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kang, B.C., Yeam, I. & Jahn, M.M. Genetics of plant virus resistance. Annu. Rev. Phytopathol. 43, 581–621 (2005).
Carrington, J.C., Kasschau, K.D. & Johansen, L.K. Activation and suppression of RNA silencing by plant viruses. Virology 281, 1–5 (2001).
Sanford, J.C. & Johnston, S.A. The concept of parasite-derived resistance-Deriving resistance genes from the parasite's own genome. J. Theor. Biol. 113, 395–405 (1985).
Abel, P.P. et al. Delay of disease development in transgenic plants that express the Tobacco mosaic virus coat protein gene. Science 232, 738–743 (1986).
Tepfer, M. Risk assessment of virus-resistant transgenic plants. Annu. Rev. Phytopathol. 40, 467–491 (2002).
Palukaitis, P. & Zaitlin, M. Replicase-mediated resistance to plant virus disease. Adv. Virus Res. 48, 349–377 (1997).
Baulcombe, D.C. Mechanisms of pathogen-derived resistance to viruses in transgenic plants. Plant Cell 8, 1833–1844 (1996).
Waterhouse, P.M., Graham, M.W. & Wang, M.B. Virus resistance and gene silencing in plants can be induced by simultaneous expression of sense and antisense RNA. Proc. Natl. Acad. Sci. USA 95, 13959–13964 (1998).
Smith, H.A., Swaney, S.L., Parks, T.D., Wernsman, E.A. & Dougherty, W.G. Transgenic plant virus resistance mediated by untranslatable sense RNAs: expression, regulation, and fate of nonessential RNAs. Plant Cell 6, 1441–1453 (1994).
Helliwell, C.A. & Waterhouse, P.M. Constructs and methods for hairpin RNA-mediated gene silencing in plants. Methods Enzymol. 392, 24–35 (2005).
Xie, Z. et al. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2, 642–652 (2004).
Mourrain, P. et al. Arabidopsis SGS2 and SGS3 genes are required for post-transcriptional gene silencing and natural virus resistance. Cell 101, 533–542 (2000).
Dalmay, T., Horsefield, R., Braunstein, T.H. & Baulcombe, D.C. SDE3 encodes an RNA helicase required for post-transcriptional gene silencing in Arabidopsis. EMBO J. 20, 2069–2078 (2001).
Chen, J., Li, W.X., Xie, D., Peng, J.R. & Ding, S.W. Viral virulence protein suppresses RNA silencing-mediated defense but upregulates the role of microrna in host gene expression. Plant Cell 16, 1302–1313 (2004).
Kasschau, K.D. et al. P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev. Cell 4, 205–217 (2003).
Llave, C., Kasschau, K.D. & Carrington, J.C. Virus-encoded suppressor of post-transcriptional gene silencing targets a maintenance step in the silencing pathway. Proc. Natl. Acad. Sci. USA 97, 13401–13406 (2000).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Jones-Rhoades, M.W., Bartel, D.P. & Bartel, B. MicroRNAs and their regulatory roles in plants. Annu. Rev. Plant Biol. 57, 19–53 (2006).
Vaucheret, H., Vazquez, F., Crete, P. & Bartel, D.P. The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes Dev. 18, 1187–1197 (2004).
Prod'homme, D., Jakubiec, A., Tournier, V., Drugeon, G. & Jupin, I. Targeting of the Turnip yellow mosaic virus 66K replication protein to the chloroplast envelope is mediated by the 140K protein. J. Virol. 77, 9124–9135 (2003).
Mitchell, E.J. & Bond, J.M. Variation in the coat protein sequence of British isolates of Turnip yellow mosaic virus and comparison with previously published isolates. Arch. Virol. 150, 2347–2355 (2005).
Tomimura, K., Gibbs, A.J., Jenner, C.E., Walsh, J.A. & Ohshima, K. The phylogeny of Turnip mosaic virus; comparisons of 38 genomic sequences reveal a Eurasian origin and a recent 'emergence' in east Asia. Mol. Ecol. 12, 2099–2111 (2003).
Sanchez, F., Martinez-Herrera, D., Aguilar, I. & Ponz, F. Infectivity of Turnip mosaic potyvirus cDNA clones and transcripts on the systemic host Arabidopsis thaliana and local lesion hosts. Virus Res. 55, 207–219 (1998).
Morch, M.D., Boyer, J.C. & Haenni, A.L. Overlapping open reading frames revealed by complete nucleotide sequencing of Turnip yellow mosaic virus genomic RNA. Nucleic Acids Res. 16, 6157–6173 (1988).
Riechmann, J.L., Lain, S. & Garcia, J.A. Highlights and prospects of potyvirus molecular biology. J. Gen. Virol. 73, 1–16 (1992).
Schwab, R., Ossowski, S., Riester, M., Warthmann, N. & Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18, 1121–1133 (2006).
Alvarez, J.P. et al. Endogenous and synthetic microRNAs stimulate simultaneous, efficient, and localized regulation of multiple targets in diverse species. Plant Cell 18, 1134–1151 (2006).
Simon-Mateo, C. & Garcia, J.A. MicroRNA-guided processing impairs Plum pox virus replication, but the virus readily evolves to escape this silencing mechanism. J. Virol. 80, 2429–2436 (2006).
Dickins, R.A. et al. Probing tumor phenotypes using stable and regulated synthetic microRNA precursors. Nat. Genet. 37, 1289–1295 (2005).
Boden, D. et al. Enhanced gene silencing of HIV-1 specific siRNA using microRNA designed hairpins. Nucleic Acids Res. 23, 1154–1158 (2004).
Stegmeier, F., Hu, G., Rickles, R.J., Hannon, G.J. & Elledge, S.J. A lentiviral microRNA-based system for single-copy polymerase II-regulated RNA interference in mammalian cells. Proc. Natl. Acad. Sci. USA 102, 13212–13217 (2005).
Zeng, Y., Wagner, E.J. & Cullen, B.R. Both natural and designed micro RNAs can inhibit the expression of cognate mRNAs when expressed in human cells. Mol. Cell 9, 1327–1333 (2002).
Sun, D., Melegari, M., Sridhar, S., Rogler, C.E. & Zhu, L. Multi-miRNA hairpin method that improves gene knockdown efficiency and provides linked multi-gene knockdown. Biotechniques 41, 59–63 (2006).
Gazzani, S., Lawrenson, T., Woodward, C., Headon, D. & Sablowski, R. A link between mRNA turnover and RNA interference in Arabidopsis. Science 306, 1046–1048 (2004).
Souret, F.F., Kastenmayer, J.P. & Green, P.J. AtXRN4 degrades mRNA in Arabidopsis and its substrates include selected miRNA targets. Mol. Cell 15, 173–183 (2004).
Szittya, G. et al. Low temperature inhibits RNA silencing-mediated defence by the control of siRNA generation. EMBO J. 22, 633–640 (2003).
Overhoff, M. et al. Local RNA target structure influences siRNA efficacy: a systematic global analysis. J. Mol. Biol. 348, 871–881 (2005).
Kameda, T., Ikegami, K., Liu, Y., Terada, K. & Sugiyama, T. A hypothermic-temperature-sensitive gene silencing by the mammalian RNAi. Biochem. Biophys. Res. Commun. 315, 599–602 (2004).
Fortier, E. & Belote, J.M. Temperature-dependent gene silencing by an expressed inverted repeat in Drosophila. Genesis 26, 240–244 (2000).
Rubio, T., Borja, M., Scholthof, H.B. & Jackson, A.O. Recombination with host transgenes and effects on virus evolution: an overview and opinion. Mol. Plant Microbe Interact. 12, 87–92 (1999).
Hammond, J., Lecoq, H. & Raccah, B. Epidemiological risks from mixed virus infections and transgenic plants expressing viral genes. Adv. Virus Res. 54, 189–314 (1999).
Aaziz, R. & Tepfer, M. Recombination in RNA viruses and in virus-resistant transgenic plants. J. Gen. Virol. 80, 1339–1346 (1999).
Falk, B.W. & Bruening, G. Will transgenic crops generate new viruses and new diseases? Science 263, 1395–1396 (1994).
Lin, S.S., Wu, H.W., Jan, F.J., Hou, R.F. & Yeh, S.D. Modifications of the HC-Pro of Zucchini yellow mosaic potyvirus for generation of attenuated mutants for cross protection against severe infection. Phytopathology (in press).
Zhang, X., Garreton, V. & Chua, N.H. The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Genes Dev. 19, 1532–1543 (2005).
Skotnicki, M.L., Mackenzie, A.M. & Gibbs, A.J. Turnip yellow mosaic virus variants produced from DNA clones encoding their genomes. Arch. Virol. 127, 25–35 (1992).
Chung, M.H., Chen, M.K. & Pan, S.M. Floral spray transformation can efficiently generate Arabidopsis transgenic plants. Transgenic Res. 9, 471–476 (2000).
Voinnet, O., Lederer, C. & Baulcombe, D.C. A viral movement protein prevents spread of the gene silencing signal in Nicotiana benthamiana. Cell 103, 157–167 (2000).
Chen, C.C. et al. Identification of Turnip mosaic virus isolates causing yellow stripe and spot on Calla Lily. Plant Dis. 87, 901–905 (2003).
Acknowledgements
S.-S.L. was supported by a fellowship from Ministry of Education, Taiwan. K.-C.C. and H.-W.W. are visiting students from the National Chung-Hsing University, Taiwan. We thank Jun Chen for TYMV, Chin-Chih Chen for TuMV and TuMV-GFP; Mengdai Xu for technical assistance, Chan-Sen Wang and Xuning Wang for assistance with microarray analysis and statistical treatment of the results; and Enno Krebbers, Richard Broglie, Karen Broglie and Barbara Mazur for helpful suggestions and stimulating discussions. This work was supported by a grant from DUPONT to N.-H.C.
Author information
Authors and Affiliations
Contributions
N.-H.C. and J.L.R. first conceived the idea of using amiRNAs to regulate gene expression. Q.-W.N., S.-S.L. and J.L.R. designed the amiRNAs. Q.-W.N. generated the transgenic plants. S.-S.L, Q.-W.N., K.-C.C. and H.-W.N. performed the virus challenge and related experiments. S.-D.Y. provided specific strains of TuMV and TuMV-GFP and advice on experimental design. All authors discussed the results and commented on the manuscript, which was written by N.-H.C. and S.-S.L.
Corresponding author
Ethics declarations
Competing interests
Q.W.N., S.-S.L., J.L.R. & N.-H.C. are co-inventors on a patent application on the use of microRNAs to regulate plant gene expression and to confer virus resistance. The results described in this manuscript were used to support claims in the patent application.
Supplementary information
Supplementary Fig. 1
Transcript analysis of 6 predicted target genes of amiRNAs. (PDF 123 kb)
Supplementary Fig. 2
Transgenic plants expressing amiR-P69159 are resistant to TYMV infection. (PDF 64 kb)
Supplementary Fig. 3
Transgenic plants expressing amiR-HC-Pro159 are resistant to TuMV infection. (PDF 91 kb)
Supplementary Fig. 4
Expression of amiRNAs, endogenous miRNAs and ta-siRNA in WT and transgenic plants at 15 °C and 24 °C. (PDF 68 kb)
Supplementary Fig. 5
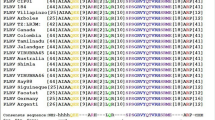
Alignment of amiR-P69159 and amiR-HC-Pro159 with P69 and HC-Pro sequences of different TYMV and TuMV strains. (PDF 274 kb)
Supplementary Table 1
Comparative microarray analysis of gene expression in WT and amiRNA transgenic plants. (PDF 65 kb)
Supplementary Table 2
Infectivity assay of amiR-TuCP159 transgenic plants challenged with TYMV or TuMV inocula. (PDF 17 kb)
Supplementary Table 3
Break-down of specific resistance amiR-P69159 and amiR-HC-Pro159 transgenic plants co-inoculated with TYMV and TuMV at 15 °C or 24 °C (PDF 17 kb)
Rights and permissions
About this article
Cite this article
Niu, QW., Lin, SS., Reyes, J. et al. Expression of artificial microRNAs in transgenic Arabidopsis thaliana confers virus resistance. Nat Biotechnol 24, 1420–1428 (2006). https://doi.org/10.1038/nbt1255
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1255
This article is cited by
-
Silencing P25, HC-Pro and Brp1 of Potato Virus (Viroid) Using Artificial microRNA Confers Resistance to PVX, PVY and PSTVd in Transgenic Potato
Potato Research (2023)
-
Systematic identification of miRNA-regulatory networks unveils their potential roles in sugarcane response to Sorghum mosaic virus infection
BMC Plant Biology (2022)
-
Tools for engineering resistance against pathogens in plants
Journal of Plant Biochemistry and Biotechnology (2022)
-
Towards plant resistance to viruses using protein-only RNase P
Nature Communications (2021)
-
Artificial miRNA mediated resistance in tobacco against Jatropha leaf curl Gujarat virus by targeting RNA silencing suppressors
Scientific Reports (2021)