Abstract
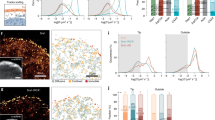
The actin cytoskeleton has been implicated in restricting diffusion of plasma membrane components. Here, simultaneous observations of quantum dot-labelled FcɛRI motion and GFP-tagged actin dynamics provide direct evidence that actin filament bundles define micron-sized domains that confine mobile receptors. Dynamic reorganization of actin structures occurs over seconds, making the location and dimensions of actin-defined domains time-dependent. Multiple FcɛRI often maintain extended close proximity without detectable correlated motion, suggesting that they are co-confined within membrane domains. FcɛRI signalling is activated by crosslinking with multivalent antigen. We show that receptors become immobilized within seconds of crosslinking. Disruption of the actin cytoskeleton results in delayed immobilization kinetics and increased diffusion of crosslinked clusters. These results implicate actin in membrane partitioning that not only restricts diffusion of membrane proteins, but also dynamically influences their long-range mobility, sequestration and response to ligand binding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Dustin, M. L. & Cooper, J. A. The immunological synapse and the actin cytoskeleton: molecular hardware for T cell signaling. Nature Immunol. 1, 23–29 (2000).
Lidke, D. S., Lidke, K. A., Rieger, B., Jovin, T. M. & Arndt-Jovin, D. J. Reaching out for signals: filopodia sense EGF and respond by directed retrograde transport of activated receptors. J. Cell. Biol. 4, 619–626 (2005).
Zhang, J., et al. Characterizing the topography of membrane receptors and signaling molecules from spatial patterns obtained using nanometer-scale electron-dense probes and electron microscopy. Micron 1, 14–34 (2006).
Boniface, J. J., et al. Initiation of signal transduction through the T cell receptor requires the multivalent engagement of peptide/MHC ligands [corrected]. Immunity 4, 459–466 (1998).
Kraft, S. & Kinet, J. P. New developments in FcɛRI regulation, function and inhibition. Nature Rev. Immunol. 5, 365–378 (2007).
Thyagarajan, R., Arunkumar, N. & Song, W. Polyvalent antigens stabilize B cell antigen receptor surface signaling microdomains. J. Immunol. 12, 6099–6106 (2003).
Kusumi, A., Ike, H., Nakada, C., Murase, K. & Fujiwara, T. Single-molecule tracking of membrane molecules: plasma membrane compartmentalization and dynamic assembly of raft-philic signaling molecules. Semin. Immunol. 1, 3–21 (2005).
Murase, K., et al. Ultrafine membrane compartments for molecular diffusion as revealed by single molecule techniques. Biophys. J. 6, 4075–4093 (2004).
Ritchie, K., et al. Detection of non-Brownian diffusion in the cell membrane in single molecule tracking. Biophys. J. 3, 2266–2277 (2005).
Saxton, M. J. & Jacobson, K. Single-particle tracking: applications to membrane dynamics. Annu. Rev. Biophys. Biomol. Struct., 373–399 (1997).
Singer, S. & Nicolson, G. The fluid mosaic model of the structure of cell membranes. Science 4023, 720–731 (1972).
Jacobson, K., Sheets, E. D. & Simson, R. Revisiting the fluid mosaic model of membranes. Science 5216, 1441–1442 (1995).
Kusumi, A., et al. Paradigm shift of the plasma membrane concept from the two-dimensional continuum fluid to the partitioned fluid: high-speed single-molecule tracking of membrane molecules. Annu. Rev. Biophys. Biomol. Struct., 351–378 (2005).
Morone, N., et al. Three-dimensional reconstruction of the membrane skeleton at the plasma membrane interface by electron tomography. J. Cell. Biol. 6, 851–862 (2006).
Douglass, A. D. & Vale, R. D. Single-molecule microscopy reveals plasma membrane microdomains created by protein–protein networks that exclude or trap signaling molecules in T cells. Cell 6, 937–950. (2005).
Draber, P. & Draberova, L. Lipid rafts in mast cell signaling. Mol. Immunol. 16–18, 1247–1252 (2002).
Frankel, D. J., et al. Revealing the topography of cellular membrane domains by combined atomic force microscopy/fluorescence imaging. Biophys. J. 7, 2404–2413 (2006).
Lillemeier, B. F., Pfeiffer, J. R., Surviladze, Z., Wilson, B. S. & Davis, M. M. Plasma membrane-associated proteins are clustered into islands attached to the cytoskeleton. Proc. Natl Acad. Sci. USA. 50, 18992–18997 (2006).
Garman, S. C., Wurzburg, B. A., Tarchevskaya, S. S., Kinet, J. P. & Jardetzky, T. S. Structure of the Fc fragment of human IgE bound to its high-affinity receptor FcɛRI α. Nature 6793, 259–266. (2000).
Metzger, H. The interaction of IgE with rat basophilic leukemia cells. II. Quantitative aspects of the binding reaction. J. Exp. Med. 6, 1676–1695 (1974).
Hartwig, J. H. & DeSisto, M. The cytoskeleton of the resting human blood platelet: structure of the membrane skeleton and its attachment to actin filaments. J. Cell. Biol. 3, 407–425 (1991).
Ortega, E., Schweitzer-Stenner, R. & Pecht, I. Possible orientational constraints determine secretory signals induced by aggregation of IgE receptors on mast cells. EMBO J. 13, 4101–4109 (1988).
Giepmans, B. N., Deerinck, T. J., Smarr, B. L., Jones, Y. Z. & Ellisman, M. H. Correlated light and electron microscopic imaging of multiple endogenous proteins using Quantum dots. Nature Methods 10, 743–749. (2005).
Wilson, B. S., Pfeiffer, J. R. & Oliver, J. M. Observing FcɛRI signaling from the inside of the mast cell membrane. J. Cell. Biol. 5, 1131–1142 (2000).
Barisas, B. G., et al. Compartmentalization of the Type I Fcɛ receptor and MAFA on mast cell membranes. Biophys. Chem. 1–3, 209–217 (2007).
Feder, T. J., Brust-Mascher, I., Slattery, J. P., Baird, B. & Webb, W. W. Constrained diffusion or immobile fraction on cell surfaces: a new interpretation. Biophys. J. 6, 2767–2773 (1996).
Suzuki, K., Ritchie, K., Kajikawa, E., Fujiwara, T. & Kusumi, A. Rapid hop diffusion of a G-protein-coupled receptor in the plasma membrane as revealed by single-molecule techniques. Biophys. J. 5, 3659–3680 (2005).
Pyenta, P., Schwille, P., Webb, W. W., Holowka, D. & Baird, B. Lateral diffusion of membrane lipid-anchored probes before and after aggregation of cell surface IgE-receptors. J. Phys. Chem. A, 107, 8310–8318 (2003).
Schlessinger, J., Webb, W. W., Elson, E. L. & Metzger, H. Lateral motion and valence of Fc receptors on rat peritoneal mast cells. Nature 5586, 550–552 (1976).
Thomas, J. L., Feder, T. J. & Webb, W. W. Effects of protein concentration on IgE receptor mobility in rat basophilic leukemia cell plasma membranes. Biophys. J. 5, 1402–1412 (1992).
Daumas, F., et al. Confined diffusion without fences of a g-protein-coupled receptor as revealed by single particle tracking. Biophys. J. 1, 356–366 (2003).
Jacquier, V., Prummer, M., Segura, J. M., Pick, H. & Vogel, H. Visualizing odorant receptor trafficking in living cells down to the single-molecule level. Proc. Natl Acad. Sci. USA, (2006).
Kusumi, A., Sako, Y. & Yamamoto, M. Confined lateral diffusion of membrane receptors as studied by single particle tracking (nanovid microscopy). Effects of calcium-induced differentiation in cultured epithelial cells. Biophys. J. 5, 2021–2040 (1993).
Pfeiffer, J. R. & Oliver, J. M. Tyrosine kinase-dependent assembly of actin plaques linking Fc-ɛ-R1 crosslinking to increased cell substrate adhesion in Rbl-2h3 tumor mast-cells. J. Immunol. 1, 270–279 (1994).
Destainville, N. & Salome, L. Quantification and correction of systematic errors due to detector time-averaging in single-molecule tracking experiments. Biophys. J. 2, L17–L19 (2006).
Frigeri, L. & Apgar, J. R. The role of actin microfilaments in the downregulation of the degranulation response in RBL-2H3 mast cells. J. Immunol. 4, 2243–2250. (1999).
Menon, A. K., Holowka, D., Webb, W. W. & Baird, B. Cross-linking of receptor-bound IgE to aggregates larger than dimers leads to rapid immobilization. J. Cell. Biol. 2, 541–550 (1986).
Dahl, S. C., Geib, R. W., Fox, M. T., Edidin, M. & Branton, D. Rapid capping in α-spectrin-deficient MEL cells from mice afflicted with hereditary hemolytic anemia. J. Cell. Biol. 5, 1057–1065 (1994).
Tang, Q. & Edidin, M. Lowering the barriers to random walks on the cell surface. Biophys. J. 1, 400–407 (2003).
Pike, L. J. Rafts defined: a report on the Keystone Symposium on Lipid Rafts and Cell Function. J. Lipid Res. 7, 1597–1598 (2006).
Viola, A. & Gupta, N. Tether and trap: regulation of membrane-raft dynamics by actin-binding proteins. Nature Rev. Immunol. 11, 889–896 (2007).
Mao, S. Y., Varin-Blank, N., Edidin, M. & Metzger, H. Immobilization and internalization of mutated IgE receptors in transfected cells. J. Immunol. 3, 958–966 (1991).
Dahan, M., et al. Diffusion dynamics of glycine receptors revealed by single-quantum dot tracking. Science 5644, 442–445 (2003).
Liu, F. T., et al. Monoclonal dinitrophenyl-specific murine IgE antibody: preparation, isolation, and characterization. J. Immunol. 6, 2728–2737 (1980).
Arndt-Jovin, D. J., et al. In vivo cell imaging with semiconductor quantum dots and noble-metal nanodots. Proc. SPIE, 6096, 1–10 (2006).
Martin, D. S., Forstner, M. B. & Kas, J. A. Apparent subdiffusion inherent to single particle tracking. Biophys. J. 4, 2109–2117 (2002).
Acknowledgements
This work was supported by NIH grants R01 GM49814, R01 AI051575 and P20 GM 67594, the Oxnard Foundation, ACS IRG 192 and by the Sandia LDRD and SURP programs. Sandia is a multi-program laboratory operated by Sandia Corporation, a Lockheed Martin Company, for the United States Department of Energy under contract DE-AC04-94AL85000. N.A. was supported by NSF IGERT DGE-0549500 and the UNM-SOM MD/PhD Program. We thank Sheli Ryan for assistance with cell culture, Jun Zhang for EM image analysis, and Jonas Anderson and Bernd Rieger for assistance with image processing. The UNM Cancer Center Fluorescence Microscopy Facility received support from NIH grants S10 RR14668, S10 RR19287, S10 RR016918, P20 RR11830 and P30 CA118100 and from NSF grant MCB9982161. Electron micrographs were generated in the University of New Mexico Electron Microscopy Facility, which received support from NIH grants P20 GM067594, S10 RRI5734 and RR022493.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1, S2, S3, S4, S5, S6, S7, S8 and Supplementary Methods (PDF 2693 kb)
Supplementary Information
Supplementary Movie 1 (MOV 1177 kb)
Supplementary Information
Supplementary Movie 2 (MOV 308 kb)
Supplementary Information
Supplementary Movie 3 (MOV 433 kb)
Supplementary Information
Supplementary Movie 4 (MOV 1040 kb)
Supplementary Information
Supplementary Movie 5 (MOV 727 kb)
Rights and permissions
About this article
Cite this article
Andrews, N., Lidke, K., Pfeiffer, J. et al. Actin restricts FcɛRI diffusion and facilitates antigen-induced receptor immobilization. Nat Cell Biol 10, 955–963 (2008). https://doi.org/10.1038/ncb1755
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1755
This article is cited by
-
CPB-3 and CGH-1 localize to motile particles within dendrites in C. elegans PVD sensory neurons
BMC Research Notes (2021)
-
Direct observation of Thermomyces lanuginosus lipase diffusional states by Single Particle Tracking and their remodeling by mutations and inhibition
Scientific Reports (2019)
-
Assessing metastatic potential of breast cancer cells based on EGFR dynamics
Scientific Reports (2019)
-
Conventional analysis of movement on non-flat surfaces like the plasma membrane makes Brownian motion appear anomalous
Communications Biology (2019)
-
Recent advances in optical microscopic methods for single-particle tracking in biological samples
Analytical and Bioanalytical Chemistry (2019)