Abstract
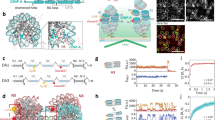
Centromeres direct the assembly of kinetochores, microtubule-attachment sites that allow chromosome segregation on the mitotic spindle. Fundamental differences in size and organization between evolutionarily distant eukaryotic centromeres have in many cases obscured general principles of their function. Here we demonstrate that centromere-binding proteins are highly conserved between budding yeast and humans. We identify the histone-fold protein Cnn1CENP-T as a direct centromere receptor of the microtubule-binding Ndc80 complex. The amino terminus of Cnn1 contains a conserved peptide motif that mediates stoichiometric binding to the Spc24–25 domain of the Ndc80 complex. Consistent with the critical role of this interaction, artificial tethering of the Ndc80 complex through Cnn1 allows mini-chromosomes to segregate in the absence of a natural centromere. Our results reveal the molecular function of CENP-T proteins and demonstrate how the Ndc80 complex is anchored to centromeres in a manner that couples chromosome movement to spindle dynamics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Cheeseman, I. M. & Desai, A. Molecular architecture of the kinetochore-microtubule interface. Nat. Rev. Mol. Cell Biol. 9, 33–46 (2008).
Lampert, F. & Westermann, S. A blueprint for kinetochores—new insights into the molecular mechanics of cell division. Nat. Rev. Mol. Cell Biol. 12, 407–412 (2011).
Santaguida, S. & Musacchio, A. The life and miracles of kinetochores. EMBO J. 28, 2511–2531 (2009).
Liu, D., Vader, G., Vromans, M. J., Lampson, M. A. & Lens, S. M. Sensing chromosome bi-orientation by spatial separation of aurora B kinase from kinetochore substrates. Science 323, 1350–1353 (2009).
Musacchio, A. & Salmon, E. D. The spindle-assembly checkpoint in space and time. Nat. Rev. Mol. Cell Biol. 8, 379–393 (2007).
Cheeseman, I. M., Chappie, J. S., Wilson-Kubalek, E. M. & Desai, A. The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell 127, 983–997 (2006).
Alushin, G. M. et al. The Ndc80 kinetochore complex forms oligomeric arrays along microtubules. Nature 467, 805–810 (2010).
Guse, A., Carroll, C. W., Moree, B., Fuller, C. J. & Straight, A. F. In vitro centromere and kinetochore assembly on defined chromatin templates. Nature 477, 354–358 (2011).
Carroll, C. W., Milks, K. J. & Straight, A. F. Dual recognition of CENP-A nucleosomes is required for centromere assembly. J. Cell Biol. 189, 1143–1155 (2010).
Foltz, D. R. et al. The human CENP-A centromeric nucleosome-associated complex. Nat. Cell Biol. 8, 458–469 (2006).
Okada, M. et al. The CENP-H-I complex is required for the efficient incorporation of newly synthesized CENP-A into centromeres. Nat. Cell Biol. 8, 446–457 (2006).
Cheeseman, I. M. et al. Phospho-regulation of kinetochore-microtubule attachments by the Aurora kinase Ipl1p. Cell 111, 163–172 (2002).
Westermann, S. et al. Architecture of the budding yeast kinetochore reveals a conserved molecular core. J. Cell Biol. 163, 215–222 (2003).
Liu, X., McLeod, I., Anderson, S., Yates, J. R. 3rd & He, X. Molecular analysis of kinetochore architecture in fission yeast. EMBO J. 24, 2919–2930 (2005).
Meraldi, P., McAinsh, A. D., Rheinbay, E. & Sorger, P. K. Phylogenetic and structural analysis of centromeric DNA and kinetochore proteins. Genome Biol. 7, R23 (2006).
Westermann, S., Drubin, D. G. & Barnes, G. Structures and functions of yeast kinetochore complexes. Annu. Rev. Biochem. 76, 563–591 (2007).
Malik, H. S. & Henikoff, S. Major evolutionary transitions in centromere complexity. Cell 138, 1067–1082 (2009).
Hori, T., Okada, M., Maenaka, K. & Fukagawa, T. CENP-O class proteins form a stable complex and are required for proper kinetochore function. Mol. Biol. Cell 19, 843–854 (2008).
De Wulf, P., McAinsh, A. D. & Sorger, P. K. Hierarchical assembly of the budding yeast kinetochore from multiple subcomplexes. Genes Dev. 17, 2902–2921 (2003).
Hori, T. et al. CCAN makes multiple contacts with centromeric DNA to provide distinct pathways to the outer kinetochore. Cell 135, 1039–1052 (2008).
Gascoigne, K. E. et al. Induced ectopic kinetochore assembly bypasses the requirement for CENP-A nucleosomes. Cell 145, 410–422 (2011).
Nishino, T. et al. CENP-T-W-S-X forms a unique centromeric chromatin structure with a histone-like fold. Cell 148, 487–501 (2012).
Burley, S. K., Xie, X., Clark, K. L. & Shu, F. Histone-like transcription factors in eukaryotes. Curr. Opin. Struct. Biol. 7, 94–102 (1997).
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Sandman, K. & Reeve, J. N. Archaeal histones and the origin of the histone fold. Curr. Opin. Microbiol. 9, 520–525 (2006).
Talbert, P. B. & Henikoff, S. Histone variants–ancient wrap artists of the epigenome. Nat. Rev. Mol. Cell Biol. 11, 264–275 (2010).
Tanaka, K., Chang, H. L., Kagami, A. & Watanabe, Y. CENP-C functions as a scaffold for effectors with essential kinetochore functions in mitosis and meiosis. Dev. Cell 17, 334–343 (2009).
Amano, M. et al. The CENP-S complex is essential for the stable assembly of outer kinetochore structure. J. Cell Biol. 186, 173–182 (2009).
Yan, Z. et al. A histone-fold complex and FANCM form a conserved DNA-remodeling complex to maintain genome stability. Mol. Cell 37, 865–878 (2010).
McAinsh, A. D. & Meraldi, P. The CCAN complex: linking centromere specification to control of kinetochore-microtubule dynamics. Semin. Cell Dev. Biol. 22, 946–952 (2011).
Petrovic, A. et al. The MIS12 complex is a protein interaction hub for outer kinetochore assembly. J. Cell Biol. 190, 835–852 (2010).
Maskell, D. P., Hu, X. W. & Singleton, M. R. Molecular architecture and assembly of the yeast kinetochore MIND complex. J. Cell Biol. 190, 823–834 (2010).
Hornung, P. et al. Molecular architecture and connectivity of the budding yeast Mtw1 kinetochore complex. J. Mol. Biol. 405, 548–559 (2011).
Ciferri, C. et al. Implications for kinetochore-microtubule attachment from the structure of an engineered Ndc80 complex. Cell 133, 427–439 (2008).
Kiermaier, E., Woehrer, S., Peng, Y., Mechtler, K. & Westermann, S. A Dam1-based artificial kinetochore is sufficient to promote chromosome segregation in budding yeast. Nat. Cell Biol. 11, 1109–1115 (2009).
Lacefield, S., Lau, D. T. & Murray, A. W. Recruiting a microtubule-binding complex to DNA directs chromosome segregation in budding yeast. Nat. Cell Biol. 11, 1116–1120 (2009).
Westermann, S. et al. The Dam1 kinetochore ring complex moves processively on depolymerizing microtubule ends. Nature 440, 565–569 (2006).
Wigge, P. A. et al. Analysis of the Saccharomyces spindle pole by matrix-assisted laser desorption/ionization (MALDI) mass spectrometry. J. Cell Biol. 141, 967–977 (1998).
Screpanti, E. et al. Direct binding of Cenp-C to the Mis12 complex joins the inner and outer kinetochore. Curr. Biol. 21, 391–398 (2011).
Suzuki, A. et al. Spindle microtubules generate tension-dependent changes in the distribution of inner kinetochore proteins. J. Cell Biol. 193, 125–140 (2011).
Akiyoshi, B. et al. Tension directly stabilizes reconstituted kinetochore-microtubule attachments. Nature 468, 576–579 (2010).
Bock, L. J. et al. Cnn1 inhibits the interactions between the KMN complexes of the yeast kinetochore. Nat. Cell Biol.http://dx.doi.org/10.1038/ncb2495 (2012).
Powers, A. F. et al. The Ndc80 kinetochore complex forms load-bearing attachments to dynamic microtubule tips via biased diffusion. Cell 136, 865–875 (2009).
Kiyomitsu, T., Obuse, C. & Yanagida, M. Human Blinkin/AF15q14 is required for chromosome alignment and the mitotic checkpoint through direct interaction with Bub1 and BubR1. Dev. Cell 13, 663–676 (2007).
Cheeseman, I. M. et al. A conserved protein network controls assembly of the outer kinetochore and its ability to sustain tension. Genes Dev. 18, 2255–2268 (2004).
Cole, C., Barber, J. D. & Barton, G. J. The Jpred 3 secondary structure prediction server. Nucleic Acids Res. 36, W197–W201 (2008).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 39, D38–D51 (2011).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Johnson, L. S., Eddy, S. R. & Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinformatics 11, 431 (2010).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Soding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005).
Finn, R. D. et al. The Pfam protein families database. Nucleic Acids Res. 38, D211–D222 (2010).
Letunic, I., Doerks, T. & Bork, P. SMART 6: recent updates and new developments. Nucleic Acids Res. 37, D229–D232 (2009).
Lupas, A., Van Dyke, M. & Stock, J. Predicting coiled coils from protein sequences. Science 252, 1162–1164 (1991).
Wootton, J. C. & Federhen, S. Analysis of compositionally biased regions in sequence databases. Methods Enzymol. 266, 554–571 (1996).
Katoh, K. & Toh, H. Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform. 9, 286–298 (2008).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Hayashi, T. et al. Mis16 and Mis18 are required for CENP-A loading and histone deacetylation at centromeres. Cell 118, 715–729 (2004).
Remmert, M., Biegert, A., Hauser, A. & Soding, J. HHblits: lightning-fast iterative protein sequence searching by HMM–HMM alignment. Nat. Methods 9, 173–175 (2012).
Lampert, F., Hornung, P. & Westermann, S. The Dam1 complex confers microtubule plus end-tracking activity to the Ndc80 kinetochore complex. J. Cell Biol. 189, 641–649 (2010).
Rodal, A. A., Manning, A. L., Goode, B. L. & Drubin, D. G. Negative regulation of yeast WASp by two SH3 domain-containing proteins. Curr. Biol. 13, 1000–1008 (2003).
Mendoza, M. A., Panizza, S. & Klein, F. Analysis of protein-DNA interactions during meiosis by quantitative chromatin immunoprecipitation (qChIP). Methods Mol. Biol. 557, 267–283 (2009).
Acknowledgements
The authors thank all members of the Westermann laboratory for discussions and J. M. Peters for critical reading of the manuscript. We thank T. Burkard for suggestions, W. Lugmayr for support with high-performance computing, O. Hudecz for help with presentation of the mass spectrometry results and M. Madalinski for peptide synthesis. We thank P. De Wulf (European Institute of Oncology, Milan, Italy) for the Cnn1–3GFP strain and communicating results before publication. Research in the Westermann laboratory receives funding from the European Research Council under the European Community’s Seventh Framework Programme (S.W. FP7/2007-2013)/ERC grant agreement no. 203499, and from the Austrian Science Fund FWF (S.W., SFB F34-B03).
Author information
Authors and Affiliations
Contributions
A.S. performed bioinformatic sequence analysis. M.M. purified CCAN proteins and performed characterization of proteins. P.H. performed interaction studies. F.L. conducted ChIP and biochemical experiments. K.M. guided the mass spectrometry analysis. S.W. guided the study and performed biochemical and genetic experiments supported by G.L. All authors discussed results and analysed data. S.W. and A.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 614 kb)
Supplementary Table 1
Supplementary Information (XLS 132 kb)
Supplementary Table 2
Supplementary Information (XLS 33 kb)
Supplementary Table 3
Supplementary Information (XLS 500 kb)
Supplementary Table 4
Supplementary Information (XLS 22 kb)
Supplementary Table 5
Supplementary Information (XLS 28 kb)
Rights and permissions
About this article
Cite this article
Schleiffer, A., Maier, M., Litos, G. et al. CENP-T proteins are conserved centromere receptors of the Ndc80 complex. Nat Cell Biol 14, 604–613 (2012). https://doi.org/10.1038/ncb2493
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2493
This article is cited by
-
Bridgin connects the outer kinetochore to centromeric chromatin
Nature Communications (2021)
-
Multiple phosphorylations control recruitment of the KMN network onto kinetochores
Nature Cell Biology (2018)
-
Regulation of kinetochore configuration during mitosis
Current Genetics (2018)
-
Drosophila Nnf1 paralogs are partially redundant for somatic and germ line kinetochore function
Chromosoma (2017)
-
The molecular basis for centromere identity and function
Nature Reviews Molecular Cell Biology (2016)