Abstract
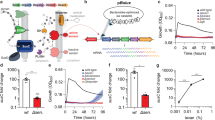
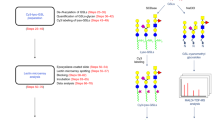
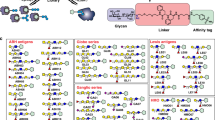
Glycosylation of bacterial cell surfaces is emerging as a critical factor in symbiosis, pathogenesis, cell-cell interactions and immune evasion1,2,3. The lack of high-throughput analytical tools to examine bacterial glycans has been a major obstacle to the field and has hindered closer examination of the dynamics of carbohydrate variation. We have recently developed a lectin microarray for the analysis of glycoproteins4. Herein we present a rapid analytical system based on this technology for the examination of bacterial glycans. The glycosylation pattern observed distinguishes closely related Escherichia coli strains from one another, providing a facile means of fingerprinting bacteria. In addition, dynamic alterations in the carbohydrate coat of a pathogenic E. coli strain are readily observed. The fast evaluation of real-time alterations in surface-carbohydrate epitopes allows examination of the dynamic role of bacterial sugars in response to external stimuli such as the immune system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Schmidt, M.A., Riley, L.W. & Benz, I. Sweet new world: glycoproteins in bacterial pathogens. Trends Microbiol. 11, 554–561 (2003).
Lerouge, I. & Vanderleyden, J. O-antigen structural variation: mechanisms and possible roles in animal/plant-microbe interactions. FEMS Microbiol. Rev. 26, 17–47 (2002).
Weintraub, A. Immunology of bacterial polysaccharide antigens. Carbohydr. Res. 338, 2539–2547 (2003).
Pilobello, K.T., Krishnamoorthy, L., Slawek, D. & Mahal, L.K. Development of a lectin microarray for the rapid analysis of protein glycopatterns. ChemBioChem 6, 985–989 (2005).
Caroff, M. & Karibian, D. Structure of bacterial lipopolysaccharides. Carbohydr. Res. 338, 2431–2447 (2003).
van der Woude, M.W. & Baumler, A.J. Phase and antigenic variation in bacteria. Clin. Microbiol. Rev. 17, 581–611 (2004).
Whitfield, C. & Roberts, I.S. Structure, assembly and regulation of expression of capsules in Escherichia coli. Mol. Microbiol. 31, 1307–1319 (1999).
Roberts, I.S. The biochemistry and genetics of capsular polysaccharide production in bacteria. Annu. Rev. Microbiol. 50, 285–315 (1996).
Harvey, D.J. Matrix-assisted laser desorption/ionization mass spectrometry of carbohydrates. Mass Spectrom. Rev. 18, 349–450 (1999).
Szymanski, C.M. et al. Detection of conserved N-linked glycans and phase-variable lipooligosaccharides and capsules from campylobacter cells by mass spectrometry and high resolution magic angle spinning NMR spectroscopy. J. Biol. Chem. 278, 24509–24520 (2003).
Sletmoen, M., Maurstad, G., Sikorski, P., Paulsen, B.S. & Stokke, B.T. Characterisation of bacterial polysaccharides: steps towards single-molecular studies. Carbohydr. Res. 338, 2459–2475 (2003).
Buttke, T.M. & Ingram, L.O. Comparison of lipopolysaccharides from Agmenellum quadruplicatum to Escherichia coli and Salmonella typhimurium by using thin-layer chromatography. J. Bacteriol. 124, 1566–1573 (1975).
Angeloni, S. et al. Glycoprofiling with micro-arrays of glycoconjugates and lectins. Glycobiology 15, 31–41 (2005).
Zheng, T., Peelen, D. & Smith, L.M. Lectin arrays for profiling cell surface carbohydrate expression. J. Am. Chem. Soc. 127, 9982–9983 (2005).
Kuno, A. et al. Evanescent-field fluorescence-assisted lectin microarray: a new strategy for glycan profiling. Nat. Methods 2, 851–856 (2005).
Aabenhus, R., Hynes, S.O., Permin, H., Moran, A.P. & Andersen, L.P. Lectin typing of Campylobacter concisus. J. Clin. Microbiol. 40, 715–717 (2002).
Hynes, S.O., Hirmo, S., Wadstrom, T. & Moran, A.P. Differentiation of Helicobacter pylori isolates based on lectin binding of cell extracts in an agglutination assay. J. Clin. Microbiol. 37, 1994–1998 (1999).
Ertl, P. & Mikkelsen, S.R. Electrochemical biosensor array for the identification of microorganisms based on lectin-lipopolysaccharide recognition. Anal. Chem. 73, 4241–4248 (2001).
Annuk, H., Hynes, S.O., Hirmo, S., Mikelsaar, M. & Wadstrom, T. Characterisation and differentiation of lactobacilli by lectin typing. J. Med. Microbiol. 50, 1069–1074 (2001).
Disney, M.D. & Seeberger, P.H. The use of carbohydrate microarrays to study carbohydrate-cell interactions and to detect pathogens. Chem. Biol. 11, 1701–1707 (2004).
Gamian, A., Kenne, L., Mieszala, M., Ulrich, J. & Defaye, J. Structure of the Escherichia coli O24 and O56 O-specific sialic-acid-containing polysaccharides and linkage of these structures to the core region in lipopolysaccharides. Eur. J. Biochem. 225, 1211–1220 (1994).
Yao, Z. & Valvano, M.A. Genetic analysis of the O-specific lipopolysaccharide biosynthesis region (rfb) of Escherichia coli K-12 W3110: identification of genes that confer group 6 specificity to Shigella flexneri serotypes Y and 4a. J. Bacteriol. 176, 4133–4143 (1994).
Wu, A.M., Wu, J.H., Singh, T., Liu, J.H. & Herp, A. Lectinochemical studies on the affinity of Anguilla anguilla agglutinin for mammalian glycotopes. Life Sci. 75, 1085–1103 (2004).
Baldus, S.E. et al. Characterization of the binding specificity of Anguilla anguilla agglutinin (AAA) in comparison to Ulex europaeus agglutinin I (UEA-I). Glycoconj. J. 13, 585–590 (1996).
Alvarez, R.A., Lee, A., Davis, C., Hoffmann, J. & Blixt, O. Defining the binding specificity of commercially available plant lectins using a printed glycan array. Glycobiology 15, 1207 (2005).
Coyne, M.J., Reinap, B., Lee, M.M. & Comstock, L.E. Human symbionts use a host-like pathway for surface fucosylation. Science 307, 1778–1781 (2005).
Vimr, E.R., Kalivoda, K.A., Deszo, E.L. & Steenbergen, S.M. Diversity of microbial sialic acid metabolism. Microbiol. Mol. Biol. Rev. 68, 132–153 (2004).
Badger, J.L. & Kim, K.S. Environmental growth conditions influence the ability of Escherichia coli K1 to invade brain microvascular endothelial cells and confer serum resistance. Infect. Immun. 66, 5692–5697 (1998).
Neidhardt, F.C., Bloch, P.L. & Smith, D.F. Culture medium for enterobacteria. J. Bacteriol. 119, 736–747 (1974).
Karlsson, K.A. The human gastric colonizer Helicobacter pylori: a challenge for host-parasite glycobiology. Glycobiology 10, 761–771 (2000).
Friis, L.M., Pin, C., Pearson, B.M. & Wells, J.M. In vitro cell culture methods for investigating Campylobacter invasion mechanisms. J. Microbiol. Methods 61, 145–160 (2005).
Manimala, J.C., Li, Z., Jain, A., VedBrat, S. & Gildersleeve, J.C. Carbohydrate array analysis of anti-Tn antibodies and lectins reveals unexpected specificities: implications for diagnostic and vaccine development. ChemBioChem 6, 2229–2241 (2005).
Acknowledgements
The authors thank D.E. Graham, Department of Chemistry and Biochemistry, University of Texas at Austin, for JM101; S.M. Payne, Department of Molecular Genetics and Microbiology, University of Texas at Austin, for LT2 and HB101; R. Silver and L. Wright, University of Rochester Medical Center, for RS218; I.J. Molineux, Department of Molecular Genetics and Microbiology, University of Texas at Austin; and the Beckman Young Investigator Award for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Growth curve of RS218 in EZ Rich Media. (PDF 24 kb)
Supplementary Fig. 2
Dye uptake as a function of OD600 for cells grown in both EZ Rich Media (Rich Media) and Luria Broth. (PDF 21 kb)
Supplementary Fig. 3
Experimental for 2-color growth-dependent variation of RS218 glycosylation. (PDF 20 kb)
Supplementary Fig. 4
2-color growth dependent variation of RS218 glycosylation (EZ-Rich Defined Media). (PDF 66 kb)
Supplementary Fig. 5
2-color growth dependent variation of RS218 glycosylation (Luria Broth (LB)). (PDF 83 kb)
Supplementary Table 1
Lectin panel. (PDF 23 kb)
Supplementary Table 2
Mean fluorescence and standard deviations. (PDF 10 kb)
Supplementary Table 3
Agglutination data. (PDF 10 kb)
Rights and permissions
About this article
Cite this article
Hsu, KL., Pilobello, K. & Mahal, L. Analyzing the dynamic bacterial glycome with a lectin microarray approach. Nat Chem Biol 2, 153–157 (2006). https://doi.org/10.1038/nchembio767
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio767
This article is cited by
-
Comprehensive analysis of glycosphingolipid glycans by lectin microarrays and MALDI-TOF mass spectrometry
Nature Protocols (2021)
-
Differential recognition of Haemophilus influenzae whole bacterial cells and isolated lipooligosaccharides by galactose-specific lectins
Scientific Reports (2018)
-
Glycointeractions in bacterial pathogenesis
Nature Reviews Microbiology (2018)
-
Lectin binding of human sperm associates with DEFB126 mutation and serves as a potential biomarker for subfertility
Scientific Reports (2016)
-
Pre-B cell receptor binding to galectin-1 modifies galectin-1/carbohydrate affinity to modulate specific galectin-1/glycan lattice interactions
Nature Communications (2015)