Abstract
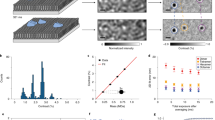
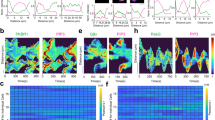
Photoactivated localization microscopy (PALM) is a powerful approach for investigating protein organization, yet tools for quantitative, spatial analysis of PALM datasets are largely missing. Combining pair-correlation analysis with PALM (PC-PALM), we provide a method to analyze complex patterns of protein organization across the plasma membrane without determination of absolute protein numbers. The approach uses an algorithm to distinguish a single protein with multiple appearances from clusters of proteins. This enables quantification of different parameters of spatial organization, including the presence of protein clusters, their size, density and abundance in the plasma membrane. Using this method, we demonstrate distinct nanoscale organization of plasma-membrane proteins with different membrane anchoring and lipid partitioning characteristics in COS-7 cells, and show dramatic changes in glycosylphosphatidylinositol (GPI)-anchored protein arrangement under varying perturbations. PC-PALM is thus an effective tool with broad applicability for analysis of protein heterogeneity and function, adaptable to other single-molecule strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kusumi, A. et al. Paradigm shift of the plasma membrane concept from the two-dimensional continuum fluid to the partitioned fluid: High-speed single-molecule tracking of membrane molecules. Annu. Rev. Biophys. Biomol. Struct. 34, 351–378 (2005).
Eggeling, C. et al. Direct observation of the nanoscale dynamics of membrane lipids in a living cell. Nature 457, 1159–1162 (2009).
Simons, K. & Gerl, M.J. Revitalizing membrane rafts: new tools and insights. Nat. Rev. Mol. Cell Biol. 11, 688–699 (2010).
Lingwood, D. & Simons, K. Lipid rafts as a membrane-organizing principle. Science 327, 46–50 (2010).
Lillemeier, B.F., Pfeiffer, J.R., Surviladze, Z., Wilson, B.S. & Davis, M.M. Plasma membrane-associated proteins are clustered into islands attached to the cytoskeleton. Proc. Natl. Acad. Sci. USA 103, 18992–18997 (2006).
Prior, I.A., Muncke, C., Parton, R.G. & Hancock, J.F. Direct visualization of Ras proteins in spatially distinct cell surface microdomains. J. Cell Biol. 160, 165–170 (2003).
van Zanten, T.S. et al. Hotspots of GPI-anchored proteins and integrin nanoclusters function as nucleation sites for cell adhesion. Proc. Natl. Acad. Sci. USA 106, 18557–18562 (2009).
Goswami, D. et al. Nanoclusters of GPI-anchored proteins are formed by cortical actin-driven activity. Cell 135, 1085–1097 (2008).
Glebov, O.O. & Nichols, B.J. Lipid raft proteins have a random distribution during localized activation of the T-cell receptor. Nat. Cell Biol. 6, 238–243 (2004).
Tanaka, K.A.K. et al. Membrane molecules mobile even after chemical fixation. Nat. Methods 7, 865–866 (2010).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S.T., Girirajan, T.P.K. & Mason, M.D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M.J., Bates, M. & Zhuang, X.W. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Folling, J. et al. Fluorescence nanoscopy by ground-state depletion and single-molecule return. Nat. Methods 5, 943–945 (2008).
Wombacher, R. et al. Live-cell super-resolution imaging with trimethoprim conjugates. Nat. Methods 7, 717–719 (2010).
Lillemeier, B.F. et al. TCR and Lat are expressed on separate protein islands on T cell membranes and concatenate during activation. Nat. Immunol. 11, 90–96 (2010).
Hess, S.T. et al. Dynamic clustered distribution of hemagglutinin resolved at 40 nm in living cell membranes discriminates between raft theories. Proc. Natl. Acad. Sci. USA 104, 17370–17375 (2007).
Fuchs, J. et al. A photoactivatable marker protein for pulse-chase imaging with superresolution. Nat. Methods 7, 627–630 (2010).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
Owen, D.M. et al. PALM imaging and cluster analysis of protein heterogeneity at the cell surface. J. Biophotonics 3, 446–454 (2010).
Veatch, S.L. et al. Critical fluctuations in plasma membrane vesicles. ACS Chem. Biol. 3, 287–293 (2008).
Balagopalan, L., Coussens, N.P., Sherman, E., Samelson, L.E. & Sommers, C.L. The LAT story: a tale of cooperativity, coordination, and choreography. Cold Spring Harbor Perspect. Biol. 2, a005512 (2010).
Ingley, E. Src family kinases: regulation of their activities, levels and identification of new pathways. BBA Proteins Proteomics 1784, 56–65 (2008).
Zagouras, P. & Rose, J.K. Dynamic equilibrium between vesicular stomatitis-virus glycoprotein monomers and trimers in the Golgi and at the cell-surface. J. Virol. 67, 7533–7538 (1993).
Baumgart, T. et al. Large-scale fluid/fluid phase separation of proteins and lipids in giant plasma membrane vesicles. Proc. Natl. Acad. Sci. USA 104, 3165–3170 (2007).
Schutze, S., Tchikov, V. & Schneider-Brachert, W. Regulation of TNFR1 and CD95 signalling by receptor compartmentalization. Nat. Rev. Mol. Cell Biol. 9, 655–662 (2008).
London, M. & London, E. Ceramide selectively displaces cholesterol from ordered lipid domains (rafts)—implications for lipid raft structure and function. J. Biol. Chem. 279, 9997–10004 (2004).
Romer, W. et al. Shiga toxin induces tubular membrane invaginations for its uptake into cells. Nature 450, 670–675 (2007).
Lagerholm, B.C., Weinreb, G.E., Jacobson, K. & Thompson, N.L. Detecting microdomains in intact cell membranes. Annu. Rev. Phys. Chem. 56, 309–336 (2005).
Batista, F.D., Treanor, B. & Harwood, N.E. Visualizing a role for the actin cytoskeleton in the regulation of B-cell activation. Immunol. Rev. 237, 191–204 (2010).
Kenworthy, A.K. et al. Dynamics of putative raft-associated proteins at the cell surface. J. Cell Biol. 165, 735–746 (2004).
Acknowledgements
We thank G. Patterson (US National Institute of Biomedical Imaging and Bioengineering) for providing plasmid constructs; E. Ambroggio (US National Institute of Child Health and Development) for providing purified PAGFP protein; H. Hess and G. Stengel for providing the Peak Selector software and valuable discussion; S. Manley for help in setting up the PALM microscope; B. Baird and D. Holowka for valuable discussions. S.L.V. was supported by US National Institutes of Health grant R00GM87810.
Author information
Authors and Affiliations
Contributions
P.S. and T.J-T. conceived and designed experiments, developed experimental techniques, performed experiments, developed the analytical method, analyzed data and wrote the paper; D.S. helped develop the analytical method; M.R. contributed to characterization of PA-FP and imaging regime; S.L.V. contributed equations used to analyze data; J.L-S. conceived and designed the experiments, and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Notes 1–2 (PDF 3504 kb)
Rights and permissions
About this article
Cite this article
Sengupta, P., Jovanovic-Talisman, T., Skoko, D. et al. Probing protein heterogeneity in the plasma membrane using PALM and pair correlation analysis. Nat Methods 8, 969–975 (2011). https://doi.org/10.1038/nmeth.1704
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1704
This article is cited by
-
Motion of VAPB molecules reveals ER–mitochondria contact site subdomains
Nature (2024)
-
Aptamer-based Membrane Protein Analysis and Molecular Diagnostics
Chemical Research in Chinese Universities (2024)
-
Maximum-likelihood model fitting for quantitative analysis of SMLM data
Nature Methods (2023)
-
Stepwise requirements for polymerases δ and θ in theta-mediated end joining
Nature (2023)
-
The non-catalytic role of DNA polymerase epsilon in replication initiation in human cells
Nature Communications (2022)