Abstract
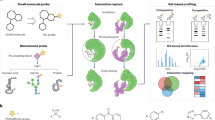
Current tandem mass spectral libraries for lipid annotations in metabolomics are limited in size and diversity. We provide a freely available computer-generated tandem mass spectral library of 212,516 spectra covering 119,200 compounds from 26 lipid compound classes, including phospholipids, glycerolipids, bacterial lipoglycans and plant glycolipids. We show platform independence by using tandem mass spectra from 40 different mass spectrometer types including low-resolution and high-resolution instruments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kind, T. & Fiehn, O. Bioanal. Rev. 2, 23–60 (2010).
Song, H., Hsu, F.F. & Turk, J. J. Am. Soc. Mass Spectrom. 18, 1848–1858 (2007).
Yetukuri, L. et al. BMC Syst. Biol. 1, 12 (2007).
Forrester, J.S., Milne, S.B., Ivanova, P.T. & Brown, H.A. Mol. Pharmacol. 65, 813–821 (2004).
Yang, K., Cheng, H., Gross, R.W. & Han, X. Anal. Chem. 81, 4356–4368 (2009).
Bou Khalil, M. et al. Mass Spectrom. Rev. 29, 877–929 (2010).
Taguchi, R. & Ishikawa, M. J. Chromatogr. A 1217, 4229–4239 (2010).
Herzog, R. et al. PLoS ONE 7, e29851 (2012).
Fahy, E., Sud, M., Cotter, D. & Subramaniam, S. Nucleic Acids Res. 35, W606–W612 (2007).
Sud, M. et al. Nucleic Acids Res. 35, D527–D532 (2007).
Pirok, G. et al. J. Chem. Inf. Model. 46, 563–568 (2006).
Schüller, A., Hähnke, V. & Schneider, G. QSAR Comb. Sci. 26, 407–410 (2007).
Stein, S.E. & Scott, D.R. J. Am. Soc. Mass Spectrom. 5, 859–866 (1994).
Quehenberger, O. et al. J. Lipid Res. 51, 3299–3305 (2010).
Gao, X. et al. Anal. Bioanal. Chem. 402, 2923–2933 (2012).
Sud, M., Fahy, E. & Subramaniam, S. J. Cheminform. 4, 23 (2012).
Subramaniam, S. et al. Chem. Rev. 111, 6452–6490 (2011).
Kangas, L.J. et al. Bioinformatics 28, 1705–1713 (2012).
Sartain, M.J., Dick, D.L., Rithner, C.D., Crick, D.C. & Belisle, J.T. J. Lipid Res. 52, 861–872 (2011).
Layre, E. et al. Chem. Biol. 18, 1537–1549 (2011).
Sheldon, M.T., Mistrik, R. & Croley, T.R. J. Am. Soc. Mass Spectrom. 20, 370–376 (2009).
Stein, S. Anal. Chem. 84, 7274–7282 (2012).
Matyash, V., Liebisch, G., Kurzchalia, T.V., Shevchenko, A. & Schwudke, D. J. Lipid Res. 49, 1137–1146 (2008).
Sandra, K., Pereira Ados, S., Vanhoenacker, G., David, F. & Sandra, P. J. Chromatogr. A 1217, 4087–4099 (2010).
Mayampurath, A.M. et al. Bioinformatics 24, 1021–1023 (2008).
Chambers, M.C. et al. Nat. Biotechnol. 30, 918–920 (2012).
Frank, A.M. et al. J. Proteome Res. 7, 113–122 (2008).
Stein, S.E. & Heller, D.N. J. Am. Soc. Mass Spectrom. 17, 823–835 (2006).
Acknowledgements
We thank the Lipid MAPS consortium and the US National Institute of General Medical Sciences for providing extensive lipid identification and database services; the NIST Mass Spectrometry group for providing the freely available NIST MS Search GUI program and for help with the Lib2NIST converter; ModLab (Universität Frankfurt am Main) for providing the free SMILIB enumeration tool; and ChemAxon for a free research license for the Marvin and Instant-JChem cheminformatics tools. K.-H.L. was supported by the National Research Foundation of Korea, Ministry of Education, Science and Technology (grant 2010-0021368), the Korea Healthcare Technology R&D Project, Ministry of Health and Welfare (grant A103017) and the Cooperative Research Program for Agriculture Science and Technology Development (project PJ00948604), Rural Development Administration, Republic of Korea. T.K. and O.F. were supported by the US National Science Foundation (MCB 1139644) and US National Institutes of Health (P20 HL113452 and U24 DK097154).
Author information
Authors and Affiliations
Contributions
T.K., K.-H.L., D.Y.L. and O.F. designed the experiments. T.K., K.-H.L., B.D., J.K.M. and D.Y.L. performed mass spectrometric experiments. T.K. and K.-H.L. performed mass spectral fragmentation analysis and compound annotations. T.K. created the compound structures and developed the in silico MS/MS libraries and wrote and validated the algorithm. T.K. and O.F. wrote the manuscript in interaction with all contributing authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figure and Note
Supplementary Figure 1–5 and Supplementary Note 1 (PDF 10817 kb)
Supplementary Table 1
Detailed statistics of the LipidBlast MS/MS libraries with detailed lipid compound numbers (XLS 97 kb)
Supplementary Table 2
Complete table of mass spectrometry platforms that can be used with the LipidBlast libraries (XLS 112 kb)
Rights and permissions
About this article
Cite this article
Kind, T., Liu, KH., Lee, D. et al. LipidBlast in silico tandem mass spectrometry database for lipid identification. Nat Methods 10, 755–758 (2013). https://doi.org/10.1038/nmeth.2551
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2551
This article is cited by
-
Novel insight into the lipid network of plasma extracellular vesicles reveal sex-based differences in the lipidomic profile of alcohol use disorder patients
Biology of Sex Differences (2024)
-
MetaboAnalystR 4.0: a unified LC-MS workflow for global metabolomics
Nature Communications (2024)
-
In vivo neuroprotective capacity of a Dunaliella salina extract - comprehensive transcriptomics and metabolomics study
npj Science of Food (2024)
-
Human Cerebrospinal Fluid Sample Preparation and Annotation for Integrated Lipidomics and Metabolomics Profiling Studies
Molecular Neurobiology (2024)
-
Lipidomic landscape of circulating extracellular vesicles isolated from adolescents exposed to ethanol intoxication: a sex difference study
Biology of Sex Differences (2023)