Abstract
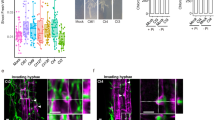
Plant infections caused by fungi are often associated with an increase in the pH of the surrounding host tissue1. Extracellular alkalinization is thought to contribute to fungal pathogenesis, but the underlying mechanisms are poorly understood. Here, we show that the root-infecting fungus Fusarium oxysporum uses a functional homologue of the plant regulatory peptide RALF (rapid alkalinization factor)2,3 to induce alkalinization and cause disease in plants. An upshift in extracellular pH promotes infectious growth of Fusarium by stimulating phosphorylation of a conserved mitogen-activated protein kinase essential for pathogenicity4,5. Fungal mutants lacking a functional Fusarium (F)-RALF peptide failed to induce host alkalinization and showed markedly reduced virulence in tomato plants, while eliciting a strong host immune response. Arabidopsis plants lacking the receptor-like kinase FERONIA, which mediates the RALF-triggered alkalinization response6, displayed enhanced resistance against Fusarium. RALF homologues are found across a number of phylogenetically distant groups of fungi, many of which infect plants. We propose that fungal pathogens use functional homologues of alkalinizing peptides found in their host plants to increase their infectious potential and suppress host immunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Prusky, D. & Yakoby, N. Pathogenic fungi: leading or led by ambient pH? Mol. Plant Pathol. 4, 509–516 (2003).
Murphy, E. & De Smet, I. Understanding the RALF family: a tale of many species. Trends Plant Sci. 19, 664–671 (2014).
Pearce, G., Moura, D. S., Stratmann, J. & Ryan, C. A. Jr . RALF, a 5-kDa ubiquitous polypeptide in plants, arrests root growth and development. Proc. Natl Acad. Sci. USA 98, 12843–12847 (2001).
Di Pietro, A., Garcia-Maceira, F. I., Meglecz, E. & Roncero, M. I. A MAP kinase of the vascular wilt fungus Fusarium oxysporum is essential for root penetration and pathogenesis. Mol. Microbiol. 39, 1140–1152 (2001).
Turra, D., Segorbe, D. & Di Pietro, A. Protein kinases in plant pathogenic fungi: conserved regulators of infection. Annu. Rev. Phytopathol. 52, 267–288 (2014).
Haruta, M., Sabat, G., Stecker, K., Minkoff, B. B. & Sussman, M. R. A peptide hormone and its receptor protein kinase regulate plant cell expansion. Science 343, 408–411 (2014).
Cessna, S. G., Sears, V. E., Dickman, M. B. & Low, P. S. Oxalic acid, a pathogenicity factor for Sclerotinia sclerotiorum, suppresses the oxidative burst of the host plant. Plant Cell 12, 2191–2200 (2000).
Prusky, D., McEvoy, J. L., Leverentz, B. & Conway, W. S. Local modulation of host pH by Colletotrichum species as a mechanism to increase virulence. Mol. Plant Microbe Interact. 14, 1105–1113 (2001).
Dean, R. et al. The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414–430 (2012).
Mulkey, T. J. & Evans, M. L. Geotropism in corn roots: evidence for its mediation by differential acid efflux. Science 212, 70–71 (1981).
Xu, J. R. & Hamer, J. E. MAP kinase and cAMP signaling regulate infection structure formation and pathogenic growth in the rice blast fungus Magnaporthe grisea. Genes Dev. 10, 2696–2706 (1996).
Lopez-Berges, M. S., Rispail, N., Prados-Rosales, R. C. & Di Pietro, A. A nitrogen response pathway regulates virulence functions in Fusarium oxysporum via the protein kinase TOR and the bZIP protein MeaB. Plant Cell 22, 2459–2475 (2010).
Srivastava, R., Liu, J. X., Guo, H., Yin, Y. & Howell, S. H. Regulation and processing of a plant peptide hormone, AtRALF23, in Arabidopsis. Plant J. 59, 930–939 (2009).
Pearce, G., Yamaguchi, Y., Munske, G. & Ryan, C. A. Structure–activity studies of RALF, rapid alkalinization factor, reveal an essential—YISY—motif. Peptides 31, 1973–1977 (2010).
Soanes, D. & Richards, T. A. Horizontal gene transfer in eukaryotic plant pathogens. Annu. Rev. Phytopathol. 52, 583–614 (2014).
Ludin, P., Nilsson, D. & Mäser, P. Genome-wide identification of molecular mimicry candidates in parasites. PLoS ONE 6, e17546 (2011).
Johnson, L. S., Eddy, S. R. & Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinformatics 11, 431 (2010).
de Jonge, R., Bolton, M. D. & Thomma, B. P. How filamentous pathogens co-opt plants: the ins and outs of fungal effectors. Curr. Opin. Plant Biol. 14, 400–406 (2011).
van Kan, J. A., Joosten, M. H., Wagemakers, C. A., van den Berg-Velthuis, G. C. & de Wit, P. J. Differential accumulation of mRNAs encoding extracellular and intracellular PR proteins in tomato induced by virulent and avirulent races of Cladosporium fulvum. Plant Mol. Biol. 20, 513–527 (1992).
Vera, P., Tornero, P. & Conejero, V. Cloning and expression analysis of a viroid-induced peroxidase from tomato plants. Mol. Plant Microbe Interact. 6, 790–794 (1993).
Jabs, T., Dietrich, R. A. & Dangl, J. L. Initiation of runaway cell death in an Arabidopsis mutant by extracellular superoxide. Science 273, 1853–1856 (1996).
Luna, E. et al. Callose deposition: a multifaceted plant defense response. Mol. Plant Microbe Interact. 24, 183–193 (2011).
Escobar-Restrepo, J. M. et al. The FERONIA receptor-like kinase mediates male–female interactions during pollen tube reception. Science 317, 656–660 (2007).
The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Hou, S. et al. The secreted peptide PIP1 amplifies immunity through receptor-like kinase 7. PLoS Pathogens 10, e1004331 (2014).
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Kessler, S. A. et al. Conserved molecular components for pollen tube reception and fungal invasion. Science 330, 968–971 (2010).
Keinath, N. F. et al. PAMP (pathogen-associated molecular pattern)-induced changes in plasma membrane compartmentalization reveal novel components of plant immunity. J. Biol. Chem. 285, 39140–39149 (2010).
Kistler, H., Bosland, P., Benny, U., Leong, S. & Williams, P. Relatedness of strains of Fusarium oxysporum from crucifers measured by examination of mitochondrial and ribosomal DNA. Phytopathology 77, 1289–1293 (1987).
Perez-Nadales, E. & Di Pietro, A. The membrane mucin Msb2 regulates invasive growth and plant infection in Fusarium oxysporum. Plant Cell 23, 1171–1185 (2011).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Felix, G., Grosskopf, D. G., Regenass, M., Basse, C. W. & Boller, T. Elicitor-induced ethylene biosynthesis in tomato cells: characterization and use as a bioassay for elicitor action. Plant Physiol. 97, 19–25 (1991).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5, 113 (2004).
Gouy, M., Guindon, S. & Gascuel, O. SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 27, 221–224 (2010).
Richards, T. A. et al. Phylogenomic analysis demonstrates a pattern of rare and ancient horizontal gene transfer between plants and fungi. Plant Cell 21, 1897–1911 (2009).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Acknowledgements
The authors thank E. Martínez Aguilera for technical assistance and A. Ramiro for help with Arabidopsis experiments. This work was supported by grant no. BIO2013-47870-R from the Spanish Ministerio de Innovación y Competitividad (MINECO) to A.D.P. S.M. has received an undergraduate student fellowship from MINECO. D.S. was supported by project BIO2008-04479 from MINECO/ERA-NET PathoGenoMics. M.L.R. was supported by project BIO296 from the Junta de Andalucia. M.E.G. was supported by a Marie Curie Initial Training Network (ITN) ARIADNE (FP7-PEOPLE-ITN-237936) grant from the European Commission. U.F. was supported by the German Federal Ministry of Education and Research–Knowledge-Based Bio-Economy in Europe (BMBF–KBBE) project 031A328 and G.F. was funded by the Deutsche Forschungsgemeinschaft through CRC 1101.
Author information
Authors and Affiliations
Contributions
A.D.P., D.T., T.A.R. and G.F. designed the experiments. S.M., D.S., D.T., M.L.R., U.F., M.E.G., G.L. and T.A.R. carried out the experiments and analysed the data. A.D.P., G.F. and T.A.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Supplementary Table 3. (PDF 9961 kb)
Supplementary Table 1
Genomes searched (XLSX 34 kb)
Supplementary Table 2
Results of JackHMMEr searches (XLS 267 kb)
Rights and permissions
About this article
Cite this article
Masachis, S., Segorbe, D., Turrà, D. et al. A fungal pathogen secretes plant alkalinizing peptides to increase infection. Nat Microbiol 1, 16043 (2016). https://doi.org/10.1038/nmicrobiol.2016.43
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.43
This article is cited by
-
Genetic Mapping of the Root Mycobiota in Rice and its Role in Drought Tolerance
Rice (2023)
-
Why is FERONIA pleiotropic?
Nature Plants (2023)
-
How plants manage pathogen infection
EMBO Reports (2023)
-
Mobile Signaling Peptides: Secret Molecular Messengers with a Mighty Role in Plant Life
Journal of Plant Growth Regulation (2023)
-
Rapid alkalinization factor: function, regulation, and potential applications in agriculture
Stress Biology (2023)