Key Points
-
Trypanosomatid parasites cause several neglected diseases in humans and animals that range in severity from comparatively mild to nearly invariably fatal. The organisms that are responsible for human diseases include Trypanosoma brucei, which causes human African trypanosomiasis (HAT), Trypanosoma cruzi, which causes Chagas disease, and Leishmania spp., which cause leishmaniasis.
-
The current drugs for the treatment of trypanosomatid diseases are unsatisfactory owing to several reasons: poor efficacy, drug resistance, toxic side effects and parenteral administration. Hence, new drugs are urgently required.
-
The drug discovery process typically takes 10–15 years. This process should be guided by target product profiles (TPPs) that define the key features and requirements for a new drug, such as route of administration, length of treatment, cost and safety margins.
-
The drug discovery process starts with target-based or phenotypic (whole-cell) approaches. In the former, compounds are screened against a molecular target (usually an enzyme); in the latter, compounds are screened directly against the intact parasite growing in culture.
-
In general, there has been a very poor success rate in target-based approaches against trypanosomatids, despite some unique biochemical and metabolic features in trypanosomatid parasites. There are very few robustly validated drug targets.
-
Phenotypic approaches have led to some promising compounds, following the optimization of the initial hits. Some of these are now in clinical development for the treatment of HAT.
-
Another approach for drug discovery is to reposition drugs from other disease areas. In some cases this has been successful.
-
There is still a need to refine the drug discovery pathway for Chagas disease, in cellular and animal models in particular, to improve the identification of compounds that can cure patients.
-
There is still a long way to go, but good progress is being made in drug discovery to find potential new drugs to treat trypanosomatid diseases.
Abstract
The WHO recognizes human African trypanosomiasis, Chagas disease and the leishmaniases as neglected tropical diseases. These diseases are caused by parasitic trypanosomatids and range in severity from mild and self-curing to near invariably fatal. Public health advances have substantially decreased the effect of these diseases in recent decades but alone will not eliminate them. In this Review, we discuss why new drugs against trypanosomatids are required, approaches that are under investigation to develop new drugs and why the drug discovery pipeline remains essentially unfilled. In addition, we consider the important challenges to drug discovery strategies and the new technologies that can address them. The combination of new drugs, new technologies and public health initiatives is essential for the management, and hopefully eventual elimination, of trypanosomatid diseases from the human population.
Similar content being viewed by others
Main
Trypanosomatid parasites cause several neglected diseases in humans and animals, which range in severity from comparatively mild to near invariably fatal1,2. The organisms that are responsible for human diseases are Trypanosoma brucei subsp., which cause human African trypanosomiasis (HAT), Trypanosoma cruzi, which causes Chagas disease, and Leishmania spp., which cause leishmaniasis. Together, these insect-transmitted parasites threaten millions of people. All of these organisms have complex life cycles, with substantial differences in morphology, cell biology and biochemistry between life cycle stages, and, in some cases, between species (Box 1).
The control of trypanosomatid diseases has had a mixed history, although public health campaigns are showing success in many instances. For example, the Southern Cone and Andean initiatives are tackling Chagas disease using a combination of insecticide spraying of dwellings, improved housing, screening of people in endemic zones and blood bank monitoring3. However, in South America there is a substantial number of individuals who are infected with T. cruzi and many infected individuals have migrated to North America and Europe, where the disease is non-endemic. In the case of leishmaniasis, co-infection with Leishmania spp. and HIV can increase disease burden and severity, and recent refugee movements from the Middle East into Europe are likely to increase the prevalence of leishmaniasis in Europe. In the immediate post-colonial period, HAT resurged, but vector control, active case-finding and treatment have all helped to control the disease4. However, many trypanosomatid diseases are zoonotic, which makes eradication extremely unlikely. The current target is elimination, which is still an ambitious goal. Despite progress, trypanosomatid diseases remain a substantial public health problem and there is an urgent need for new drugs to tackle them.
None of the available drugs for the treatment of trypanosomatid diseases (Table 1) is satisfactory and new drugs are required, especially those that are suitable for rural health systems that have limited resources. The current standard of care is monotherapy, with the exception of nifurtimox–eflornithine combination therapy (NECT) for HAT, although various drug combinations are in clinical trials. Importantly, many of the current treatments require parenteral administration5 and also have poor efficacy, major side effects and increasing levels of resistance6,7,8. Most of the drugs that are in use probably have several modes of action, as they act on multiple parasite targets9. Goals for drug discovery include the development of completely new classes of therapeutic, reduced host toxicity, improved administration regimens and the development of combination therapies.
Vaccine development is a powerful approach to disease management but remains challenging in trypanosomatid diseases, owing to efficient immune-evasion mechanisms, such as antigenic variation in African trypanosomes, and the intracellular locations of T. cruzi and Leishmania spp. in the human host. Progress towards human10 and canine11 Leishmania spp. vaccines, and the challenges in developing vaccines for HAT12 and Chagas disease13, have been reviewed recently and will not be discussed here.
In this Review, we discuss the potential for the development of new drug therapies against trypanosomatids. In addition, we highlight unique biological features of these parasites that suggest potential targets, methods that are used to identify bioactive compounds and consider some of the outcomes of recent campaigns. We encourage the reader to consider excellent reviews of the life cycles, genomes, pathogenesis and more general aspects of the biology of trypanosomatids that have been published elsewhere (see Refs 14,15,16,17,18,19).
Drug discovery
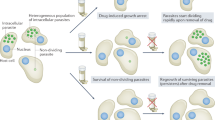
A successful drug discovery campaign typically takes 10–15 years (Fig. 1). High attrition rates, together with relatively few organizations working on drug discovery for trypanosomatid parasites, mean that the number of new compounds in clinical development is very low (Fig. 2) and unlikely to meet the clinical need. Ideally, the pipeline should contain several new agents that are suitable for combination therapy. The advantages of combination therapies are manifold: they can increase the clinical efficacy of treatments; they can decrease side effects by enabling lower dosing of individual agents; and they can decrease the risk of developing resistance. Decreasing resistance is crucial for safeguarding new medicines that emerge from the drug discovery pipeline.
Drug discovery progresses through several stages (hit discovery, lead optimization, preclinical, clinical and registration stages), each of which involves specific steps and regulations. The failure rate at each stage is high, which underscores the need for an active pipeline of drug discovery projects.
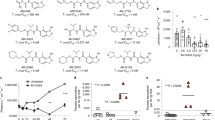
a | Several compounds are currently in preclinical and clinical development for human African trypanosomiasis (HAT), visceral leishmaniasis (VL) and Chagas disease (CD). b | Anti-trypanosomatid compounds that were identified through phenotypic approaches that have been progressed into clinical trials.
Three broad approaches are used for drug discovery against trypanosomatids. First, there are target-based approaches, which involve screening for inhibitors against a purified protein (for example, an enzyme). Compounds identified through the screening (or structure-based) process are subsequently optimized to show efficacy in a cellular model. Second, there are phenotypic approaches, which involve screening for growth inhibitors directly against an intact parasite, usually in an in vitro culture. Last, there is compound re-positioning, which is the re-deployment of compounds that were previously developed for an alternative use as new anti-trypanosomatid therapies.
The drug discovery process is ideally driven by target product profiles (TPPs), which define the properties of a drug that are required for clinical application20,21,22. Such factors include the route of administration (for example, oral, inhalation or intravenous), acceptable dosing regimen and course of treatment, acceptable safety and tolerability levels, cost and shelf-life. TPPs enable the development of compound progression criteria, which define parameters for compounds at each stage of the drug discovery process (for example, hit, validated hit, lead and preclinical candidate; see Fig. 1). Progression criteria include assessment of the physicochemical properties (such as solubility in physiological media, lipophilicity, molecular weight, hydrogen bond donors and acceptors), potency (against the molecular target and intact organism), selectivity, chemical and metabolic stability, pharmacokinetics, efficacy and safety. Additional criteria for parasitic infections can include factors such as cytocidal activity and the rate of parasite killing. The Drugs for Neglected Diseases Initiative (DNDi) is a public–private partnership that focuses on drug discovery and clinical development for these organisms. It has developed TPPs and compound progression criteria for trypanosomatid diseases (see the DNDi website for more information)21.
Target-based approaches
For target-based approaches, the key is careful selection of the most promising molecular targets. A recent review highlights some examples of target-based drug discovery against trypanosomatids23. For neglected diseases in general, including the trypanosomatid diseases, there has been very limited success from target-based approaches. This is often due to a lack of translation from inhibition of the target (enzyme) in a purified cell-free context to inhibition of proliferation of the parasite and/or subsequent activity in an animal disease model. In part, this reflects the absence of robustly validated targets (for example, enzymes that have essential activities for the parasite) and highlights the need for fundamental research into trypanosomatid biology and for thorough genetic and chemical validation of potential targets24. However, this is only part of the problem. As we discuss below, an improved understanding of how to translate compounds that are active in vitro into therapeutics is required, which includes better defining the cellular and animal models (Box 2) that predict clinical efficacy in humans.
We have published some criteria to aid in the selection of molecular targets9,20,24 (Box 3). Many target-based drug discovery programmes can be initially viewed as target validation25. Therefore, it is vital to obtain proof-of-concept (POC) of anti-parasitic activity for new target-derived chemical series at the earliest possible stage, ideally both in cellular and animal models, to minimize the waste of resources if the target fails to progress.
Drug targets with the highest degree of validation. The best-validated drug target in T. brucei is ornithine decarboxylase, which is the target of eflornithine, a drug that is used clinically for the treatment of HAT. Eflornithine is a suicide inhibitor that was initially developed for the treatment of cancer, but was subsequently re- purposed for HAT26. Selectivity is thought to arise from the more rapid turnover of human ornithine decarboxylase compared with the trypanosome enzyme27, or due to the inhibition of the biosynthesis of trypanothione28, which is a metabolite that is unique to trypanosomatids.
The enzyme N-myristoyltransferase (NMT) has also been well validated as a molecular target for HAT29,30,31. In a programme that was initiated with a high-throughput screen against NMT, a compound series was identified and subsequently optimized (typified by DDD85646; Fig. 3b) to be active in a mouse model of the first stage of HAT, which does not involve the central nervous system. There was strong evidence that the compounds inhibit NMT in cells and that this inhibition kills parasites, which validates both the target and the mode of action. NMT is also present in humans, but T. brucei is acutely sensitive to NMT inhibition, probably because endocytosis, which occurs at a very high rate in T. brucei, is affected. NMT has also been validated as a target in a second-stage mouse model of HAT (K.D.R., unpublished observations). The challenge with the second-stage disease is that compounds need to penetrate the blood–brain barrier and achieve therapeutic concentrations in the central nervous system without causing host toxicity.
a | Trypanosomatids have unique metabolic pathways and cellular functions that are attractive for drug discovery. Many of the enzymes they produce are divergent from other eukaryotes, and they have unique or highly specialized organelles such as the kinetoplast and the glycosome, respectively. b | For some anti-trypanosomatid compounds the molecular targets are known. DDD85646 targets N-myristoyltransferase (NMT), posaconazole and ravuconazole are CYP51 inhibitors, K777 irreversibly inhibits the cysteine protease cruzipain, and GNF6702 selectively inhibits the trypanosomatid proteasome.
Very recently, the proteasome was shown to have great potential as a target in all three types of trypansomatid32. This study used a phenotypic approach to develop a parasite-specific selective inhibitor (GNF6702) that does not inhibit the human proteasome. This is an excellent example of taking a phenotypic hit and subsequently deconvoluting the target. The initial experiments to determine the mode of action involved generating compound-resistant T. cruzi mutants followed by whole-genome sequencing, which revealed mutations in the β4 subunit of the proteasome. Various additional biochemical experiments have demonstrated that GNF6702 specifically inhibits the chymotrypsin-like activity of the parasite proteasome.
Biological features of trypanosomatids that might be targeted. Trypanosomatids are one of the most evolutionary divergent eukaryotic lineages from mammals, a feature that is reflected in their distinct biology (Fig. 3a). Conversely, there are many similarities between T. brucei, T. cruzi and Leishmania spp., and many molecular mechanisms are conserved between all three lineages. Trypanosomatid-specific metabolic and cellular pathways (discussed below) should represent excellent drug targets as specificity should be an easier criterion to control, but no candidate drugs have been developed that inhibit such targets. In fact, most potential trypanosome-specific targets remain unexplored for drug discovery and/or are of unknown druggability. Ironically, the best-validated targets in trypanosomatids are those repurposed from oncology (ornithine decarboxylase) and two pan-eukaryotic essential targets (NMT and the proteasome), which are discussed above.
Uniquely, trypanosomatids package the first six or seven enzymes of glycolysis into the glycosome, which is a specialized form of peroxisome. Glycolysis is especially important for the bloodstream forms of African trypanosomes, which rely exclusively on this pathway for the production of ATP. The compartmentalization of glycolysis in trypanosomatids is accompanied by fundamental differences in allosteric regulation of the pathway compared with most other eukaryotes. Consequently, phosphofructokinase, for example, is being pursued as a target33. However, computational modelling of glycolysis suggests that there is little prospect of killing trypanosomes by suppressing glycolysis unless inhibition is irreversible or uncompetitive, owing to the enormous glycolytic flux through the system34. Metabolic compartmentalization requires the transport of substrates (glucose), negatively-charged metabolic intermediates (such as 3-phosphoglyceric acid, dihydroxyacetone phosphate and glycerol-3-phosphate) and products (such as pyruvate). The transporters and permeases for these molecules (and other larger charged metabolites and cofactors, such as nucleotide diphosphates and triphosphates, nucleotide sugars and NADH) remain elusive but could represent potential drug targets18. Similarly, the biogenesis of glycosomes might also have unique and druggable features.
With about 180 members, the kinomes of trypanosomatids are extensive but lack predicted receptor tyrosine kinases, or even general tyrosine kinases, and contain disproportionately high numbers of certain enzyme subtypes; for example, STE and NEK kinases35. Chemical biology has demonstrated distinct inhibition profiles for host and parasite kinases36, which suggests that the selective inhibition of parasite kinases is feasible. Furthermore, both genome-wide and kinome-wide RNAi-knockdown screens indicate that several of these enzymes are essential35,37. However, although potent and selective inhibitors against essential protein kinases in cultured parasites have been developed38,39,40, none was sufficiently active in vivo. The repurposing of mammalian kinase inhibitors has shown promise41, with cure of HAT in an animal model reported for one kinase inhibitor42. However, so far, it is unknown which (if any) trypanosomatid kinases are being targeted by the repurposed mammalian kinase inhibitor, and both chemical and genetic validation of this approach are still required. The recent identification of a highly divergent kinetochore in trypanosomes43 may provide new kinase targets in this class, but their druggability remains to be determined.
Trypanosomatids also have other divergent signalling pathways that could provide therapeutic opportunities. For example, whereas trypanosomatids lack identifiable G protein-coupled receptors, they have a large family of membrane-bound adenylate cyclases that modulate the immune response of the host44 and are probably involved in parasite differentiation45 through unconventional downstream cyclic AMP (cAMP) response proteins46,47. Similarly, a family of cAMP phosphodiesterases have attracted interest as potential targets48,49.
The assembly and maintenance of the cell surface is crucial for organisms that interact with, and defend themselves against, their hosts and the immune system. Although the fundamentals of protein and membrane synthesis, transport and recycling are well conserved among eukaryotes, there is substantial specialization between species. For example, trypanosomatids have evolved divergent protein N-glycosylation50,51 and glycosylphosphatidylinositol (GPI) membrane anchor biosynthetic pathways, the latter of which is a validated target for HAT52. Similarly, the machineries for the export of glycoproteins, and for endocytosis and recycling, are highly divergent in trypanosomes, with several canonical components being replaced by novel factors53,54,55. The major surface glycoproteins are also distinct, and although the functions of many of these glycoproteins remain unknown, they are probably crucial for survival in the host56. Furthermore, the endosomal apparatus contains some components that are important for defence against the innate immune response57. All of these peculiarities provide the potential for therapeutic exploitation.
Interestingly, endocytosis and transport mediated by transmembrane proteins are important for drug uptake by trypanosomatids. For example, aqua- glyceroporin 2 from T. brucei is responsible for the uptake of melarsoprol and pentamidine, and the invariant surface glycoprotein 75 is responsible for the uptake of suramin58,59,60.
Divergent gene expression might also be targeted. Transcription in trypanosomatids is almost exclusively polycistronic, and several chromatin modifiers are involved in determining the sites of transcription initiation and termination61. Bromodomain 'readers' in particular, which bind to acetylated histones, are potential targets62, as are histone acetyltransferases (also known as 'writers' (Ref. 63)) and deacetylases (also known as 'erasers' (Ref. 64)). Novel transcription factors are also potentially druggable, such as class I transcription factor A65, which is also, unusually, required for the transcription of genes that encode the major surface glycoproteins by RNA polymerase I (Pol1) in the African trypanosome; Pol1 is restricted to ribosomal RNA transcription in most other eukaryotes.
Protein-coding mRNAs require trans-splicing in trypanosomatids, which is distinct from the cis-splicing that is required to remove introns from the vast majority of mammalian mRNAs. Although the splicing mechanism for cis-splicing and trans-splicing is broadly similar, there are substantial differences in the splicing machinery66. Polycistronic transcription relies on the post- transcriptional control of gene expression and, consistent with this, numerous trypanosomal RNA-binding proteins have key roles in mRNA maturation, stability and translational control67. The process of translation itself also presents novel targets at the level of the ribosome68 and aminoacyl-tRNA synthetases69.
Examples of target-based drug discovery programmes. There are several examples of trypanosomatid-specific targets that have been investigated. One example involves redox metabolism; trypanosomes have a unique di-thiol called trypanothione. Several enzymes that are involved in the synthesis and modulation of the trypanothione redox system, including trypanothione reductase (TryR)70 and synthetase (TryS)71,72, are essential for the survival of the parasite. Numerous attempts have been made to discover drug-like inhibitors of TryR73,74. Multiple series have been identified from several large-scale and medium-scale screens of synthetic libraries and natural products, some of which have been used in structure-based drug design. Unfortunately, so far, none has delivered compounds that are suitable for clinical development. A key reason seems to be the large hydrophobic active site of TryR70, which is difficult to inhibit with a small drug-like molecule. Active compounds have also been designed against the companion biosynthetic enzyme TryS75.
In Chagas disease, the biosynthesis of sterols has been the focus of several drug discovery programmes. Several molecular targets have been investigated, including sterol 14α-demethylase (CYP51)76 and squalene synthase77. Much of this effort has involved repurposing compounds that were developed as antifungals or as cholesterol- lowering agents. Clinical trials have tested two CYP51 inhibitors, posaconazole78 and fosravuconazole (also known as E1224; a prodrug of ravuconazole; Fig. 3b). Although there was an initial clearance of parasites with posaconazole and fosravuconazole, disease recurred after treatment ceased, which indicates that neither agent is suitable for treatment, at least as a monotherapy. The reasons for these failures are not fully understood but they highlight the need for animal models (Box 2) that can distinguish between compounds that are efficacious in humans and those, such as posaconazole, that are not79.
Another substantially progressed target for Chagas disease is cruzipain, which is a protease that has similarities to cathepsin L. A vinyl sulfone irreversible inhibitor of cruzipain (K777) was advanced to preclinical development80,81 but was abandoned owing to poor tolerability in primates and dogs, even at a low dose.
Folate metabolism has also been the subject of extensive drug discovery programmes, in particular the enzymes dihydrofolate reductase and a trypanosome- specific target, pteridine reductase 1 (PTR1). Both of these enzymes are thought to be essential, at least in T. brucei82,83. There are similarities between the substrates for these enzymes, and inhibitors have been identified that inhibit both enzymes84. Despite extensive work in this area, for reasons that are not fully understood, there is little correlation between activity against the enzyme and activity against the parasite85. As far as we understand, no inhibitors for these targets have progressed to preclinical development.
Trypanosomatids lack purine biosynthetic pathways and take up purines from the host. In leishmaniasis, this dependence on external purines has been targeted with allopurinol. Allopurinol is taken up by the parasites and is then phosphoribosylated to the corresponding nucleotide, which then acts as a cellular poison86. It is used for the treatment of leishmaniasis in dogs and has been in clinical trials in humans but has not progressed.
Phenotypic approaches
To circumvent the challenges of target-based drug discovery, phenotypic approaches have been widely used for most neglected disease agents, including for the trypanosomatids87. In this regard, the key requirements are appropriate chemical libraries for screening5,88, robust assays and appropriate screening cascades.
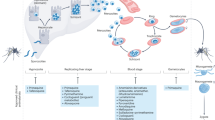
Screening cascades. Many different cellular assays are available for the analysis of trypanosome responses to compounds (Fig. 4). It is especially important to establish that compounds are effective against the clinically relevant life cycle stages, which can be problematic for the intracellular life cycle stages of Leishmania spp. and T. cruzi. Compounds must cross multiple membranes to reach targets with the parasite in cellular assays; three in the case of Leishmania spp. amastigotes that reside inside acidic (pH ∼5.5) parasitophorous vacuoles within macrophages. In animal models the situation is more complex still, as there are additional barriers to cross.
Various life cycle stages can be used for the purpose of hit discovery that range from insect forms to host-stage forms in animal models. The different technologies that can be used for phenotypic assays depend on the parasite form and stage, and have specific advantages and disadvantages. Examples of compounds the anti-trypanosomal activities of which were detected using insect forms, in vitro host-stage forms and animal models are shown (edelfosine168, GNF6702 (Ref. 32), SCYX-7158 (Ref. 109) and sodium antimony tartrate169). T. brucei, Trypanosoma brucei; T. cruzi, Trypanosoma cruzi.
To identify molecules that are suitable for drug discovery, it is essential to use an appropriate combination of assays to build confidence in the chemical start points (referred to as hits). For example, initial hit finding generally requires a high-throughput assay to access chemical diversity, followed by confirmation by more physiologically relevant (but lower throughput) assays (Fig. 4). Additional cellular assays that are representative of the in vivo situation are important to support combined pharmacokinetic and pharmacodynamic analyses in animal models. These provide an indication of the concentration and exposure time of a compound that are required to kill the parasites in animals and to predict the situation in humans. The best combination and the optimal order of phenotypic assays depend on the parasite in question.
For T. brucei, typical high-throughput screens identify not only favoured cytocidal compounds but also proliferation-slowing and cytostatic compounds. Therefore, a secondary assay is required to select those hits that are cytocidal, either using washout experiments to demonstrate a lack of reversibility29,89,90 or direct cell viability assays91. A further issue for HAT is that compounds need to penetrate the blood–brain barrier to be active against second-stage disease. Currently, there are no reliable in vitro (cell-based) assays for predicting penetration of the blood–brain barrier. However, the physicochemical properties of compounds that are likely to penetrate this barrier have been analysed92,93,94, which can assist in the selection of compounds for screening.
T. cruzi usually replicates well in intracellular amastigote assays95, which enables the identification of both cidal and static compounds. However, as T. cruzi evades the immune system during chronic infection, cidal compounds are probably essential for cure. Therefore, hits need to be followed up in a cidality assay. There is now also a drive to remove compounds that target CYP51 (see above), and assays that directly assess activity against CYP51 (Refs 78,96) need to be added to the screening cascade.
For Leishmania spp., many of the intracellular assays only report cytocidal compounds, as intracellular amastigotes replicate relatively slowly97,98. Although this eliminates the need for further cidality assays, the hit rates are low21 and throughput can be relatively poor. Furthermore, it is challenging to identify potentially valuable but weak or poorly selective hits. One solution to the low throughput is to use an axenic (free- growing) amastigote assay as the primary screen. Axenic amastigotes do not occur naturally, so care must be taken in interpreting the data. Such assays also need to be designed to only identify cytocidal compounds to prevent false-positives, as we have recently reported99. Hits can then be confirmed in an intracellular assay.
For all trypanosomatids, as with other areas of anti-infective drug discovery, it is also crucial to measure activity against a panel of clinical isolates before progressing compound series too far, to be sure that activity is not specific to laboratory strains. For all cell-based assays, replication rate, starting density and rate of killing are key factors that are required to correctly interpret compound potency; it is important to define these parameters as clearly as possible before interpreting data on new hits.
To date, phenotypic approaches have been more successful in discovering new developable series than target-based screens. In the case of HAT, the two compounds that are currently in clinical trials, fexinidazole and the oxaborole SCYX-7158, were both derived from phenotypic approaches (Fig. 2b).
Recognizing that nitroheterocycles have anti- trypanosomal activity, DNDi sourced and screened numerous nitroheterocycles and re-discovered fexinidazole, a compound that had been investigated preclinically by Hoechst but was then abandoned100. Nitroheterocycles can be genotoxic; therefore, counter-screening for genotoxicity at an early stage was a key selection criterion101. Fexinidazole, similarly to nifurtimox, is a prodrug that requires activation by a nitroreductase102. Fexinidazole has also been shown to have potential for the treatment of Chagas disease103 and leishmaniasis. Sulfoxide and the sulfone metabolites of fexinidazole, rather than the parent drug, are the active compounds against the intra-macrophage form of Leishmania spp.104. Results of a phase II proof-of-concept clinical trial against visceral leishmaniasis are expected soon. For Chagas disease, the metabolites are more active than the parent compound105 and a phase II trial was initiated. Unfortunately, the doses that were used in this trial caused safety and tolerability issues and the trial was stopped.
Another nitroheterocycle, DNDi-VL-2098, showed activity in animal models of leishmaniasis106 and was selected for further development from a series of nitroimidazooxazoles being investigated preclinically by DNDi. Unfortunately, toxic effects were noted and the progression of the compound was stopped. A backup for this compound (DNDi-0690) has now been selected and is in preclinical development (Fig. 2a). The antitubercular drug delamanid, which belongs to the same chemical class, has also been proposed as a possible candidate107. A novel nitroreductase (NTR2) has been identified as the activating enzyme for these bicyclic nitroheterocycles in Leishmania spp.108.
From a library of oxaboroles, the benzoxaborole 6-carboxamides were particularly active against T. brucei and, following a lead optimization programme, SCYX-7158 was selected as a clinical candidate for HAT109. The mode of action of oxaboroles against HAT is still not understood, but may include polypharmacology110. Another oxaborole, DNDi-6148, has recently been moved into preclinical development with DNDi for visceral leishmaniasis.
A series of diamidines showed potent activity against HAT, one of which (pafuramidine) was taken into clinical trials. Pafuramidine is a prodrug that is metabolized by the host into the active compound diamidine DB75. Although the precise mode of action is unknown, similar to other diamidines, the drug is selectively concentrated within parasites111. However, clinical trials were unsuccessful and were stopped owing to safety concerns112.
Sitamaquine, which is an orally bioavailable 8-aminoquinoline, was discovered by the Walter Reed Army Institute of Research and has progressed into clinical trials for the treatment of visceral leishmaniasis by GlaxoSmithKline113,114. The mechanism of its action is not fully understood115.
Importantly, the modes of action of all of the aforementioned compound series were unknown during the drug discovery process and up to candidate selection, and indeed, remain at best incompletely characterized. Thus, although the absence of a clear understanding of the mode of action does not preclude clinical development, it does represent a major gap in knowledge that can hinder the further optimization and development of back-up series.
Compound repositioning
Recently, there has been considerable interest in repurposing or repositioning drugs and drug-leads for many diseases116,117,118,119,120,121,122. However, the concept is not new and many drugs that are currently used for the treatment of neglected tropical diseases were 'repositioned' from anticancer, antibacterial, antifungal and anti-helminthic indications. These include the antifungal amphotericin B, the anticancer agent miltefosine and the antibiotic paromomycin, all of which were repurposed for the treatment of visceral leishmaniasis123. Other examples have already been mentioned above. More recently, the nitrofuran drug nifurtimox, which was originally developed in the 1960s for the treatment of Chagas disease, was repositioned as a combination therapy with eflornithine (NECT; mentioned previously) to decrease the cost and duration of treatment of late-stage HAT124. Unfortunately, not all repurposing efforts have been successful; for example, the CYP51 inhibitors against T. cruzi.
Drug repurposing is not without its drawbacks. For example, the drugs may have been optimized for a different human disease and the initial therapeutic activity may become an undesirable side effect that needs to be reduced or eliminated. A second problem is that repurposed drugs often do not fit the TPP for neglected diseases and many are not fit for purpose in resource-poor settings. High cost, marginal safety windows, the need for hospitalization or prolonged treatment, poor stability in conditions of high temperature and high humidity, and lack of oral bioavailability are just some of the issues that must be addressed. Nonetheless, the adage of Sir James Black, “the most fruitful basis for the discovery of a new drug is to start with an old drug”, still has substantial value, as the success with NECT, amphotericin B, paromomycin and other drugs attest125.
Target deconvolution
Phenotypic screening of chemical libraries and existing drugs has produced many chemical start points, particularly for the treatment of infections with T. brucei and T. cruzi, and, to a lesser extent, for Leishmania spp.21. However, chemical optimization of these phenotypic start points can be challenging owing, for example, to pharmacokinetic issues, insufficient potency or off-target toxicity. Without target deconvolution (that is, the identification of the molecular target), target-based screening cannot be used to find alternative chemical scaffolds that might overcome these issues, and structure-based drug design cannot be used for compound optimization126. In addition, although not essential, knowledge of the mode of action can facilitate the design of combination therapies, surveillance for the emergence and spread of resistance, and assessment of the risk of resistance.
Target deconvolution has proved very successful in many therapeutic areas127, in particular for malaria, for which several new targets have been identified recently from phenotypic hits, including PfATP4 (Ref. 128), PfPI4K129, PfeEF2 (Ref. 130), PfCARL131 and PfPheRS132. Another recent example of validating a trypanosomatid target (the proteasome) through deconvolution of an optimized phenotypic hit was discussed above32.
Although several approaches to target deconvolution exist133, further development is required for the trypanosomatids. Small molecules have many potential cellular targets, and unbiased screening approaches can be extremely powerful in identifying genetic, biochemical or metabolic associations with their modes of action. Genetic screens perturb gene expression by knockdown, knockout or overexpression. A particularly powerful approach for T. brucei is RNA interference target sequencing (RIT-seq)37, which has successfully identified genes that contribute to anti-trypanosomal drug action58. The CRISPR–Cas9 genome-editing approach is established as a powerful alternative to RNAi for genome-scale loss-of-function screening134 and is functional in T. cruzi135,136 and in Leishmania spp.137,138. Gain-of-function screens have also been used for drug target identification in T. brucei139 and Leishmania spp.140, and similar technology is available for T. cruzi141, but these approaches have not yet been widely applied to target deconvolution.
Chemical proteomics is also useful for target deconvolution. Essentially, proteins from a cell extract are isolated based on their affinity for immobilized small-molecule drug leads and are then identified by mass spectrometry142, an approach that has been used to identify potential target kinases in T. brucei36. Other approaches, such as the cellular thermal shift assay, also use chemical proteomic profiling but do not require immobilization of the inhibitor on beads143, which can be problematic for maintaining binding to the target protein. In addition, metabolomics can detect the depletion of metabolic products and the accumulation of substrates, which can indicate specific target enzymes144. Cellular approaches can also contribute to target deconvolution by revealing morphological defects in the cellular compartments that are primarily affected by a drug lead. Similarly, computational approaches may be used for structure-based target prediction.
A combination of largely unbiased orthogonal approaches to target deconvolution (from those outlined above) represents a powerful new strategy to alleviate current bottlenecks in the progression of compounds that are developed from trypanosomatid phenotypic screens.
Perspectives
In the past decade, drug discovery efforts against neglected tropical diseases have increased. Importantly, some pharmaceutical companies have become more engaged during this time period and several academic centres have established powerful drug discovery capabilities. Public–private partnerships, such as DNDi and various charitable and government funding agencies, have made major financial and other contributions to enable activities to proceed on the scale that is required for drug discovery. It is exciting that new compounds are undergoing clinical trials for HAT, although attrition in the drug discovery process suggests that there is no room for complacency. However, there are currently no new classes of drug in the clinical development pipeline for leishmaniasis or Chagas disease, and there is still a great need for new (ideally oral) drugs to treat each trypanosomatid disease. By definition, combination therapies to improve efficacy and decrease the risk of resistance require two or more drugs, preferably that have distinct modes of action, and place even more pressure on the development pipeline. Hence, more work is still required.
There are several reasons why the drug discovery process has not yet yielded new drugs for trypanosomatid diseases. There is a lack of well-validated molecular targets in the trypanosomatids, which has hampered traditional target-based approaches. Target-based assays have been replaced by more successful phenotypic screens. However, phenotypic screens have their own challenges. For HAT, compounds need to penetrate the blood–brain barrier to treat second-stage disease, which limits the compounds that should be screened or progressed. For Chagas disease, many of the hits target CYP51, which is a very promiscuous target and was unsuccessful in the clinical trials of posaconazole and fosravuconazole. For leishmaniasis, there is a very low hit-rate against the clinically relevant intra-macrophage form, for reasons that are not well understood.
One of the key challenges of phenotypic drug discovery is how to address issues, such as potency, toxicity and pharmacokinetic problems, that arise during the hit optimization process. Scaffold hopping and vector optimization become more problematic without knowledge of the molecular target. The identification of the targets of phenotypic hits should facilitate progression of these compounds and also enable more high-value target-based drug discovery in the future.
Another major challenge is defining the relevant cellular and animal models (Box 2) that closely mimic human clinical conditions. This is problematic for trypanosomatid diseases, as there are very few clinically active compounds that can be used to define these models and many of the clinically active compounds are unconventional: they are reactive (for example, nitro drugs); they are selective as a result of active transport (for example, melarsoprol and pentamidine); they are active through polypharmacological actions (for example, arsenicals and antimonials); or they are covalent inhibitors (for example, eflornithine). It is possible that in vitro cellular assays, which more closely mimic animal and human leishmanial infections, could have a higher hit rate in phenotypic screening than current assays. For Chagas disease, we need cellular and animal models that can distinguish between compounds that are active in humans (for example, benznidazole) and those that are not (for example, posaconazole). Each new compound that is taken into the clinic can provide valuable pharmacodynamic insights, which should be fed back into the drug discovery process to refine all of these models.
Despite the aforementioned challenges, the development of new in vivo and in vitro technologies, and superior methods for genetic manipulation of parasites, and increased collaborations between the pharmaceutical industry, academic laboratories, charities and other non-government organizations will start to fill the drug pipeline against these devastating and global diseases.
Change history
05 June 2017
In figure 2b of this article, the structures of Pafuramidine and DB75 were incorrect. The mistakes have been corrected in the PDF and online. The authors apologize to readers for any confusion caused.
11 September 2018
The structures of nifurtimox in Table 1 were incorrect and have been updated in the pdf and online. The authors apologize for any confusion caused.
References
McCall, L. I. & McKerrow, J. H. Determinants of disease phenotype in trypanosomatid parasites. Trends Parasitol. 30, 342–349 (2014).
World Health Organization. Investing to Overcome the Global Impact of Neglected Tropical Diseases: Third WHO Report on Neglected Diseases (World Health Organisation, 2015).
Dias, J. C. Southern Cone Initiative for the elimination of domestic populations of Triatoma infestans and the interruption of transfusional Chagas' disease. Historical aspects, present situation, and perspectives. Mem. Inst. Oswaldo Cruz 102 (Suppl. 1), 11–18 (2007).
Khyatti, M. et al. Infectious diseases in North Africa and North African immigrants to Europe. Eur. J. Public Health 24 (Suppl. 1), 47–56 (2014).
Gilbert, I. H. Target-based drug discovery for human African trypanosomiasis: selection of molecular target and chemical matter. Parasitology 141, 28–36 (2014).
Singh, N., Kumar, M. & Singh, R. K. Leishmaniasis: current status of available drugs and new potential drug targets. Asian Pac. J. Trop. Med. 5, 485–497 (2012).
Clayton, J. Chagas' disease: pushing through the pipeline. Nature 465, S12–S15 (2010).
Fairlamb, A. H., Gow, N. A. R., Matthews, K. R. & Waters, A. P. Drug resistance in eukaryotic microorganisms. Nat. Microbiol. 1, e16092 (2016).
Gilbert, I. H. Drug discovery for neglected diseases: molecular target-based and phenotypic approaches. J. Med. Chem. 56, 7719–7726 (2013). This paper compares molecular target-based and phenotypic approaches for drug discovery and highlights the strengths and weaknesses of each approach in the context of neglected diseases.
Srivastava, S., Shankar, P., Mishra, J. & Singh, S. Possibilities and challenges for developing a successful vaccine for leishmaniasis. Parasit. Vectors 9, 277 (2016).
Gharbi, M. et al. Leishmaniosis (Leishmania infantum infection) in dogs. Rev. Sci. Tech. 34, 613–626 (2015).
La Greca, F. & Magez, S. Vaccination against trypanosomiasis: can it be done or is the trypanosome truly the ultimate immune destroyer and escape artist? Hum. Vaccin. 7, 1225–1233 (2011).
Rodriguez-Morales, O. et al. Experimental vaccines against Chagas' disease: a journey through history. J. Immunol. Res. 2015, 489758 (2015).
Langousis, G. & Hill, K. L. Motility and more: the flagellum of Trypanosoma brucei. Nat. Rev. Microbiol. 12, 505–518 (2014).
Cardoso, M. S., Reis-Cunha, J. L. & Bartholomeu, D. C. Evasion of the immune response by Trypanosoma cruzi during acute infection. Front. Immunol. 6, 659 (2015).
Welburn, S. C., Molyneux, D. H. & Maudlin, I. Beyond tsetse — implications for research and control of human African trypanosomiasis epidemics. Trends Parasitol. 32, 230–241 (2016).
Imhof, S. & Roditi, I. The social life of African trypanosomes. Trends Parasitol. 31, 490–498 (2015).
Haanstra, J. R., Gonzalez-Marcano, E. B., Gualdron-Lopez, M. & Michels, P. A. Biogenesis, maintenance and dynamics of glycosomes in trypanosomatid parasites. Biochim. Biophys. Acta 1863, 1038–1048 (2016).
McConville, M. J. & Naderer, T. Metabolic pathways required for the intracellular survival of Leishmania. Annu. Rev. Microbiol. 65, 543–561 (2011).
Wyatt, P. G., Gilbert, I. H., Read, K. D. & Fairlamb, A. H. Target validation: linking target and chemical properties to desired product profile. Curr. Top. Med. Chem. 11, 1275–1283 (2011). This study is a key paper that discusses the selection of molecular targets for drug discovery in neglected diseases.
Don, R. & Ioset, J.-R. Screening strategies to identify new chemical diversity for drug development to treat kinetoplastid infections. Parasitology 141, 140–146 (2014). This study summarizes screening strategies for drug discovery in kinetoplastids.
Nwaka, S. & Hudson, A. Innovative lead discovery strategies for tropical diseases. Nat. Rev. Drug Discov. 5, 941–955 (2006).
Nagle, A. S. et al. Recent developments in drug discovery for Leishmaniasis and human African trypanosomiasis. Chem. Rev. 114, 11305–11347 (2014). This study provides a comprehensive survey of some target-based drug discovery programmes against trypanosomatids.
Frearson, J. A., Wyatt, P. G., Gilbert, I. H. & Fairlamb, A. H. Target assessment for antiparasitic drug discovery. Trends Parasitol. 23, 589–595 (2007).
Workman, P. & Collins, I. Probing the probes: fitness factors for small molecule tools. Chem. Biol. 17, 561–577 (2010). This study provides a detailed discussion on what is required for a chemical probe.
Priotto, G. et al. Safety and effectiveness of first line eflornithine for Trypanosoma brucei gambiense sleeping sickness in Sudan: cohort study. BMJ 336, 705–708 (2008).
Brun, R., Don, R., Jacobs, R. T., Wang, M. Z. & Barrett, M. P. Development of novel drugs for human African trypanosomiasis. Future Microbiol. 6, 677–691 (2011).
Fairlamb, A. H., Henderson, G. B., Bacchi, C. J. & Cerami, A. In vivo effects of difluoromethylornithine on trypanothione and polyamine levels in blood-stream forms of Trypanosoma brucei. Mol. Biochem. Parasitol. 24, 185–191 (1987).
Frearson, J. A. et al. N-Myristoyltransferase inhibitors as new leads to treat sleeping sickness. Nature 464, 728–732 (2010). This study validates N -myristoyltransferase as a drug target for T. brucei.
Brand, S. et al. Discovery of a novel class of orally active trypanocidal N-myristoyltransferase inhibitors. J. Med. Chem. 55, 140–152 (2012).
Brand, S. et al. Lead optimization of a pyrazole sulfonamide series of Trypanosoma brucei N-myristoyltransferase inhibitors: identification and evaluation of CNS penetrant compounds as potential treatments for stage 2 human African trypanosomiasis. J. Med. Chem. 57, 9855–9869 (2014).
Khare, S. et al. Proteasome inhibition for treatment of leishmaniasis, Chagas' disease and sleeping sickness. Nature 537, 229–233 (2016). This study validates the proteasome as a drug target for trypanosomatids.
Brimacombe, K. R. et al. Identification of ML251, a potent inhibitor of T. brucei and T. cruzi phosphofructokinase. ACS Med. Chem. Lett. 5, 12–17 (2014).
Eisenthal, R. & Cornish-Bowden, A. Prospects for antiparasitic drugs. The case of Trypanosoma brucei, the causative agent of African sleeping sickness. J. Biol. Chem. 273, 5500–5505 (1998).
Jones, N. G. et al. Regulators of Trypanosoma brucei cell cycle progression and differentiation identified using a kinome-wide RNAi screen. PLoS Pathog. 10, e1003886 (2014).
Urbaniak, M. D. et al. Chemical proteomic analysis reveals the drugability of the kinome of Trypanosoma brucei. ACS Chem. Biol. 7, 1858–1865 (2012).
Alsford, S. et al. High throughput phenotyping using parallel sequencing of RNA interference targets in the African trypanosome. Genome Res. 21, 915–924 (2011). This study describes a genome-scale loss-of-function RNAi approach that facilitates drug-target prioritization for a trypanosomatid.
Woodland, A. et al. From on-target to off-target activity: identification and optimisation of Trypanosoma brucei GSK3 inhibitors and their characterisation as anti-Trypanosoma brucei drug discovery lead molecules. ChemMedChem 8, 1127–1137 (2013).
Urich, R. et al. The design and synthesis of potent and selective inhibitors of Trypanosoma brucei glycogen synthase kinase 3 for the treatment of human African trypanosomiasis. J. Med. Chem. 57, 7536–7549 (2014).
Ma, J. et al. Nuclear DBF-2-related kinases are essential regulators of cytokinesis in bloodstream stage Trypanosoma brucei. J. Biol. Chem. 285, 15356–15368 (2010).
Amata, E. et al. Identification of “preferred” human kinase inhibitors for sleeping sickness lead discovery. Are some kinases better than others for inhibitor repurposing? ACS Infect. Dis. 2, 180–186 (2016).
Diaz, R. et al. Identification and characterization of hundreds of potent and selective inhibitors of Trypanosoma brucei growth from a kinase-targeted library screening campaign. PLoS Negl. Trop. Dis. 8, e3253 (2014).
Akiyoshi, B. & Gull, K. Discovery of unconventional kinetochores in kinetoplastids. Cell 156, 1247–1258 (2014).
Salmon, D. et al. Adenylate cyclases of Trypanosoma brucei inhibit the innate immune response of the host. Science 337, 463–466 (2012).
Mony, B. M. et al. Genome-wide dissection of the quorum sensing signalling pathway in Trypanosoma brucei. Nature 505, 681–685 (2014).
Gould, M. K. et al. Cyclic AMP effectors in African trypanosomes revealed by genome-scale RNA interference library screening for resistance to the phosphodiesterase inhibitor CpdA. Antimicrob. Agents Chemother. 57, 4882–4893 (2013).
Tagoe, D. N., Kalejaiye, T. D. & de Koning, H. P. The ever unfolding story of cAMP signaling in trypanosomatids: vive la difference! Front. Pharmacol. 6, 185 (2015).
de Koning, H. P. et al. Pharmacological validation of Trypanosoma brucei phosphodiesterases as novel drug targets. J. Infect. Dis. 206, 229–237 (2012).
Veerman, J. et al. Synthesis and evaluation of analogs of the phenylpyridazinone NPD-001 as potent trypanosomal TbrPDEB1 phosphodiesterase inhibitors and in vitro trypanocidals. Bioorg. Med. Chem. 24, 1573–1581 (2016).
Izquierdo, L. et al. Distinct donor and acceptor specificities of Trypanosoma brucei oligosaccharyltransferases. EMBO J. 28, 2650–2661 (2009).
Damerow, M. et al. Identification and functional characterization of a highly divergent N-acetylglucosaminyltransferase I (TbGnTI) in Trypanosoma brucei. J. Biol. Chem. 289, 9328–9339 (2014).
Smith, T. K., Crossman, A., Brimacombe, J. S. & Ferguson, M. A. J. Chemical validation of GPI biosynthesis as a drug target against African sleeping sickness. EMBO J. 23, 4701–4708 (2004).
Adung'a, V. O., Gadelha, C. & Field, M. C. Proteomic analysis of clathrin interactions in trypanosomes reveals dynamic evolution of endocytosis. Traffic 14, 440–457 (2013).
Manna, P. T., Gadelha, C., Puttick, A. E. & Field, M. C. ENTH and ANTH domain proteins participate in AP2-independent clathrin-mediated endocytosis. J. Cell Sci. 128, 2130–2142 (2015).
Manna, P. T., Kelly, S. & Field, M. C. Adaptin evolution in kinetoplastids and emergence of the variant surface glycoprotein coat in African trypanosomatids. Mol. Phylogenet. Evol. 67, 123–128 (2013).
Gadelha, C. et al. Architecture of a host–parasite interface: complex targeting mechanisms revealed through proteomics. Mol. Cell. Proteomics 14, 1911–1926 (2015).
Bart, J. M. et al. Localization of serum resistance-associated protein in Trypanosoma brucei rhodesiense and transgenic Trypanosoma brucei brucei. Cell. Microbiol. 17, 1523–1535 (2015).
Alsford, S. et al. High-throughput decoding of antitrypanosomal drug efficacy and resistance. Nature 482, 232–236 (2012). This study uses a genome-scale loss-of-function RNAi approach to identify mechanisms of drug-resistance and potential target pathways in T. brucei.
Zoltner, M., Leung, K. F., Alsford, S., Horn, D. & Field, M. C. Modulation of the surface proteome through multiple ubiquitylation pathways in African trypanosomes. PLoS Pathog. 11, e1005236 (2015).
Song, J. et al. Pentamidine is not a permeant but a nanomolar inhibitor of the Trypanosoma brucei aquaglyceroporin-2. PLoS Pathog. 12, e1005436 (2016).
Siegel, T. N. et al. Four histone variants mark the boundaries of polycistronic transcription units in Trypanosoma brucei. Genes Dev. 23, 1063–1076 (2009).
Schulz, D. et al. Bromodomain proteins contribute to maintenance of bloodstream form stage identity in the African trypanosome. PLoS Biol. 13, e1002316 (2015).
Kawahara, T. et al. Two essential MYST-family proteins display distinct roles in histone H4K10 acetylation and telomeric silencing in trypanosomes. Mol. Microbiol. 69, 1054–1068 (2008).
Wang, Q. et al. Targeting lysine deacetylases (KDACs) in parasites. PLoS Negl. Trop. Dis. 9, e0004026 (2015).
Brandenburg, J. et al. Multifunctional class I transcription in Trypanosoma brucei depends on a novel protein complex. EMBO J. 26, 4856–4866 (2007).
Gunzl, A. The pre-mRNA splicing machinery of trypanosomes: complex or simplified? Eukaryot. Cell 9, 1159–1170 (2010).
Clayton, C. E. Networks of gene expression regulation in Trypanosoma brucei. Mol. Biochem. Parasitol. 195, 96–106 (2014).
Hashem, Y. et al. High-resolution cryo-electron microscopy structure of the Trypanosoma brucei ribosome. Nature 494, 385–389 (2013).
Kalidas, S. et al. Genetic validation of aminoacyl-tRNA synthetases as drug targets in Trypanosoma brucei. Eukaryot. Cell 13, 504–516 (2014).
Patterson, S. et al. Dihydroquinazolines as a novel class of Trypanosoma brucei trypanothione reductase inhibitors: discovery, synthesis, and characterization of their binding mode by protein crystallography. J. Med. Chem. 54, 6514–6530 (2011).
Wyllie, S. et al. Dissecting the essentiality of the bifunctional trypanothione synthetase-amidase in Trypanosoma brucei using chemical and genetic methods. Mol. Microbiol. 74, 529–540 (2009).
Torrie, L. S. et al. Chemical validation of trypanothione synthetase. J. Biol. Chem. 284, 36137–36145 (2009).
Bernardes, L. S., Zani, C. L. & Carvalho, I. Trypanosomatidae diseases: from the current therapy to the efficacious role of trypanothione reductase in drug discovery. Curr. Med. Chem. 20, 2673–2696 (2013).
Beig, M., Oellien, F., Krauth Siegel, R. L. & Selzer, P. M. in Trypanosomatid Diseases: Molecular Routes to Drug Discovery (eds Jager, T., Koch, O. & Flohe, L.) 385–404 (Wiley, 2013).
Spinks, D. et al. Design, synthesis and biological evaluation of Trypanosoma brucei trypanothione synthetase inhibitors. ChemMedChem 7, 95–106 (2012).
Urbina, J. A. Chemotherapy of Chagas' disease: the how and the why. J. Mol. Med. 77, 332–338 (1999).
Urbina, J. A., Concepcion, J. L., Rangel, S., Visbal, G. & Lira, R. Squalene synthase as a chemotherapeutic target in Trypanosoma cruzi and Leishmania mexicana. Mol. Biochem. Parasitol. 125, 35–45 (2002).
Molina, I. et al. Randomized trial of posaconazole and benznidazole for chronic Chagas' disease. 370, 1899–1908 (2014). This study details a clinical trial of posaconazole for the treatment of Chagas disease.
Francisco, A. F. et al. Limited ability of posaconazole to cure both acute and chronic Trypanosoma cruzi infections revealed by highly sensitive in vivo imaging. Antimicrob. Agents Chemother. 59, 4653–4661 (2015).
Choy, J. W. et al. Chemical–biological characterization of a cruzain inhibitor reveals a second target and a mammalian off-target. Beilstein J. Org. Chem. 9, 15–25 (2013).
Barr, S. C. et al. A cysteine protease inhibitor protects dogs from cardiac damage during infection by Trypanosoma cruzi. Antimicrob. Agents Chemother. 49, 5160–5161 (2005).
Sienkiewicz, N., Jaroslawski, S., Wyllie, S. & Fairlamb, A. H. Chemical and genetic validation of dihydrofolate reductase-thymidylate synthase as a drug target in African trypanosomes. Mol. Microbiol. 69, 520–533 (2008).
Sienkiewicz, N., Ong, H. B. & Fairlamb, A. H. Trypanosoma brucei pteridine reductase 1 is essential for survival in vitro and for virulence in mice. Mol. Microbiol. 77, 658–671 (2010).
Hardy, L. W., Matthews, W., Nare, B. & Beverley, S. M. Biochemical and genetic tests for inhibitors of Leishmania pteridine pathways. Exp. Parasitol. 87, 157–169 (1997).
Gibson, M. W., Dewar, S., Ong, H. B., Sienkiewicz, N. & Fairlamb, A. H. Trypanosoma brucei DHFR-TS revisited: characterisation of a bifunctional and highly unstable recombinant dihydrofolate reductase-thymidylate synthase. PLoS Negl. Trop. Dis. 10, e0004714 (2016).
Looker, D. L., Marr, J. J. & Berens, R. L. Mechanisms of action of pyrazolopyrimidines in Leishmania donovani. J. Biol. Chem. 261, 9412–9415 (1986).
Gilbert, I. H., Leroy, D. & Frearson, J. A. Finding new hits in neglected disease projects: target or phenotypic based screening? Curr. Top. Med. Chem. 11, 1284–1291 (2011).
Brenk, R. et al. Lessons learnt from assembling screening libraries for drug discovery for neglected diseases. ChemMedChem 3, 435–444 (2008).
Mercer, L. et al. 2,4-Diaminopyrimidines as potent inhibitors of Trypanosoma brucei and identification of molecular targets by a chemical proteomics approach. PLoS Negl. Trop. Dis. 5, e956 (2011).
Shibata, S. et al. Selective inhibitors of methionyl-tRNA synthetase have potent activity against Trypanosoma brucei infection in mice. Antimicrob. Agents Chemother. 55, 1982–1989 (2011).
De Rycker, M. et al. A static–cidal assay for Trypanosoma brucei to aid hit prioritisation for progression into drug discovery programmes. PLoS Negl. Trop. Dis. 6, e1932 (2012).
Wager, T. T. et al. Defining desirable central nervous system drug space through the alignment of molecular properties, in vitro ADME, and safety attributes. ACS Chem. Neurosci. 1, 420–434 (2010).
Wager, T. T., Hou, X., Verhoest, P. R. & Villalobos, A. Moving beyond rules: the development of a Central Nervous System Multiparameter Optimization (CNS MPO) approach to enable aignment of drug-like properties. ACS Chem. Neurosci. 1, 435–449 (2010).
Wager, T. T., Villalobos, A., Verhoest, P. R., Hou, X. J. & Shaffer, C. L. Strategies to optimize the brain availability of central nervous system drug candidates. Expert. Opin. Drug Discov. 6, 371–381 (2011).
Ley, V., Andrews, N. W., Robbins, E. S. & Nussenzweig, V. Amastigotes of Trypanosoma cruzi sustain an infective cycle in mammalian cells. J. Exp. Med. 168, 649–659 (1988).
Riley, J. et al. Development of a fluorescence-based Trypanosoma cruzi CYP51 inhibition assay for effective compound triaging in drug discovery programmes for Chagas' disease. PLoS Negl. Trop. Dis. 9, e0004014 (2015).
De Rycker, M. et al. Comparison of a high-throughput high-content intracellular Leishmania donovani assay with an axenic amastigote assay. Antimicrob. Agents Chemother. 57, 2913–2922 (2013).
Berman, J. D., Dwyer, D. M. & Wyler, D. J. Multiplication of Leishmania in human macrophages in vitro. Infect. Immun. 26, 375–379 (1979).
Nuhs, A. et al. Development and validation of a novel Leishmania donovani screening cascade for high-throughput screening using a novel axenic assay with high predictivity of leishmanicidal intracellular activity. PLoS Negl. Trop. Dis. 9, e0004094 (2015).
Torreele, E. et al. Fexinidazole — a new oral nitroimidazole drug candidate entering clinical development for the treatment of sleeping sickness. PLoS Negl. Trop. Dis. 4, e923 (2010). The study details the development of fexinidazole for the treatment of HAT.
Tweats, D., Bourdin Trunz, B. & Torreele, E. Genotoxicity profile of fexinidazole — a drug candidate in clinical development for human African trypanomiasis (sleeping sickness). Mutagenesis 27, 523–532 (2012).
Wyllie, S. et al. Nitroheterocyclic drug-resistance mechanisms in Trypanosoma brucei. J. Antimicrob. Chemother. 71, 625–634 (2016).
Bahia, M. T. et al. Fexinidazole: a potential new drug candidate for Chagas' disease. PLoS Negl. Trop. Dis. 6, e1870 (2012).
Wyllie, S. et al. The anti-trypanosome drug fexinidazole shows potential for treating visceral leishmaniasis. Sci. Transl Med. 4, 119re1 (2012). This study repositions fexinidazole for the treatment of leishmaniasis.
Bahia, M. T. et al. Antitrypanosomal activity of fexinidazole metabolites, potential new drug candidates for Chagas' disease. Antimicrob. Agents Chemother. 58, 4362–4370 (2014).
Gupta, S. et al. Nitroimidazo-oxazole compound DNDI-VL-2098: an orally effective preclinical drug candidate for the treatment of visceral leishmaniasis. J. Antimicrob. Chemother. 70, 518–527 (2015). This study provides details of a preclinical candidate for visceral leishmaniasis.
Patterson, S. et al. The anti-tubercular drug delamanid as a potential oral treatment for visceral leishmaniasis. eLife 5, e09744 (2016).
Wyllie, S. et al. Activation of bicyclic nitro-drugs by a novel nitroreductase (NTR2) in Leishmania. PLoS Pathog. 12, e1005971 (2016).
Jacobs, R. T. et al. SCYX-7158, an orally-active benzoxaborole for the treatment of stage 2 human African trypanosomiasis. PLoS Negl. Trop. Dis. 5, e1151 (2011). This study provides details of a clinical candidate for HAT.
Jones, D. C. et al. Genomic and proteomic studies on the mode of action of oxaboroles against the African trypanosome. PLoS Negl. Trop. Dis. 9, e0004299 (2015).
Mathis, A. M. et al. Accumulation and intracellular distribution of antitrypanosomal diamidine compounds DB75 and DB820 in African trypanosomes. Antimicrob. Agents Chemother. 50, 2185–2191 (2006).
Barrett, M. P. & Croft, S. L. Management of trypanosomiasis and leishmaniasis. Br. Med. Bull. 104, 175–196 (2012).
Loiseau, P. M., Cojean, S. & Schrevel, J. Sitamaquine as a putative antileishmanial drug candidate: from the mechanism of action to the risk of drug resistance. Parasite 18, 115–119 (2011).
Sundar, S. et al. Pharmacokinetics of oral sitamaquine taken with or without food and safety and efficacy for treatment of visceral leishmaniais: a randomized study in Bihar, India. Am. J. Trop. Med. Hyg. 84, 892–900 (2011).
Coimbra, E. S. et al. Mechanism of interaction of sitamaquine with Leishmania donovani. J. Antimicrob. Chemother. 65, 2548–2555 (2010).
Ashburn, T. T. & Thor, K. B. Drug repositioning: identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 3, 673–683 (2004).
Jin, G. & Wong, S. T. Toward better drug repositioning: prioritizing and integrating existing methods into efficient pipelines. Drug Discov. Today 19, 637–644 (2014).
Nzila, A., Ma, Z. & Chibale, K. Drug repositioning in the treatment of malaria and TB. Future Med. Chem. 3, 1413–1426 (2011).
Cavalla, D. Predictive methods in drug repurposing: gold mine or just a bigger haystack? Drug Discov. Today 18, 523–532 (2013).
Cragg, G. M., Grothaus, P. G. & Newman, D. J. New horizons for old drugs and drug leads. J. Nat. Prod. 77, 703–723 (2014).
Cavalla, D. Therapeutic switching: a new strategic approach to enhance R&D productivity. IDrugs 8, 914–918 (2005).
O'Connor, O. A. Developing new drugs for the treatment of lymphoma. Eur. J. Haematol. Suppl. 75, 150–158 (2005).
Croft, S. L., Sundar, S. & Fairlamb, A. H. Drug resistance in leishmaniasis. Clin. Microbiol. Rev. 19, 111–126 (2006).
Priotto, G. et al. Nifurtimox–eflornithine combination therapy for second-stage African Trypanosoma brucei gambiense trypanosomiasis: a multicentre, randomised, phase III, non-inferiority trial. Lancet 374, 56–64 (2009).
Raju, T. N. The Nobel chronicles. Lancet 355, 1022 (2000).
Skinner-Adams, T. S. et al. Defining the targets of antiparasitic compounds. Drug Discov. Today 21, 725–739 (2016).
Swinney, D. C. & Anthony, J. How were new medicines discovered? Nat. Rev. Drug Discov. 10, 507–519 (2011).
Rottman, M. et al. Spiroindolones, a potent compound class for the treatment of malaria. Science 329, 1175–1180 (2010).
McNamara, C. W. et al. Targeting Plasmodium PI(4)K to eliminate malaria. Nature 504, 248–253 (2013).
Baragana, B. et al. A novel multiple-stage antimalarial agent that inhibits protein synthesis. Nature 522, 315–320 (2015).
Meister, S. et al. Imaging of Plasmodium liver stages to drive next-generation antimalarial drug discovery. Science 334, 1372–1377 (2011).
Kato, N. et al. Diversity-oriented synthesis yields novel multistage antimalarial inhibitors. Nature 538, 344–349 (2016).
Horn, D. & Duraisingh, M. T. Antiparasitic chemotherapy: from genomes to mechanisms. Annu. Rev. Pharmacol. Toxicol. 54, 71–94 (2014).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR–Cas9 system. Science 343, 80–84 (2014).
Peng, D., Kurup, S. P., Yao, P. Y., Minning, T. A. & Tarleton, R. L. CRISPR–Cas9-mediated single-gene and gene family disruption in Trypanosoma cruzi. mBio 6, e02097–14 (2015).
Lander, N., Li, Z. H., Niyogi, S. & Docampo, R. CRISPR/Cas-9-induced disruption of paraflagellar rod protein 1 and 2 genes in Trypanosoma cruzi reveals their role in flagellar attachment. mBio 6, e01012 (2015).
Sollelis, L. et al. First efficient CRISPR–Cas9-mediated genome editing in Leishmania parasites. Cell. Microbiol. 17, 1405–1412 (2015).
Zhang, W. W. & Matlashewski, G. CRISPR–Cas9- mediated genome editing in Leishmania parasites. mBio 21, e00861 (2015).
Begolo, D., Erben, E. & Clayton, C. Drug target identification using a trypanosome overexpression library. Antimicrob. Agents Chemother. 58, 6260–6264 (2014). This study validates a high-throughput gain-of-function genetic approach for drug-target identification in T. brucei.
Gazanion, E., Fernandez-Prada, C., Papadopoulou, B., Leprohon, P. & Ouellette, M. Cos-Seq for high-throughput identification of drug target and resistant mechanisms in the protozoan parasite Leishmania. Proc. Natl Acad. Sci. USA 113, E3012–E3021 (2016). The study validates a high-throughput gain-of-function genetic approach for drug-target identification in Leishmania spp.
Kelly, J. M., Das, P. & Tomas, A. M. An approach to functional complementation by introduction of large DNA fragments into Trypanosoma cruzi and Leishmania donovani using a cosmid shuttle vector. Mol. Biochem. Parasitol. 65, 51–62 (1994).
Wang, K. et al. Chemistry-based functional proteomics for drug target deconvolution. Expert Rev. Proteomics 9, 293–310 (2012).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Creek, D. J., Anderson, J., McConville, M. J. & Barrett, M. P. Metabolomic analysis of trypanosomatid protozoa. Mol. Biochem. Parasitol. 181, 73–84 (2012).
Trindade, S. et al. Trypanosoma brucei parasites occupy and functionally adapt to the adipose tissue in mice. Cell Host Microbe 19, 837–848 (2016).
Capewell, P. et al. The skin is a significant but overlooked anatomical reservoir for vector-borne African trypanosomes. eLife 5, e17716 (2016).
Desquesnes, M. et al. Trypanosoma evansi and surra: a review and perspectives on origin, history, distribution, taxonomy, morphology, hosts, and pathogenic effects. Biomed. Res. Int. 2013, 194176 (2013).
Carnes, J. et al. Genome and phylogenetic analyses of Trypanosoma evansi reveal extensive similarity to T. brucei and multiple independent origins for dyskinetoplasty. PLoS Negl. Trop. Dis. 9, e3404 (2015).
Auty, H., Torr, S. J., Michoel, T., Jayaraman, S. & Morrison, L. J. Cattle trypanosomosis: the diversity of trypanosomes and implications for disease epidemiology and control. Rev. Sci. Tech. 34, 587–598 (2015).
Bern, C. Chagas' disease N. Engl. J. Med. 373, 456–466 (2015).
Gascon, J., Bern, C. & Pinazo, M. J. Chagas' disease in Spain, the United States and other non-endemic countries. Acta Trop. 115, 22–27 (2010).
Bonney, K. M. & Engman, D. M. Autoimmune pathogenesis of Chagas' heart disease: looking back, looking ahead. Am. J. Pathol. 185, 1537–1547 (2015).
Zingales, B. et al. The revised Trypanosoma cruzi subspecific nomenclature: rationale, epidemiological relevance and research applications. Infect. Genet. Evol. 12, 240–253 (2012).
Cunha-Neto, E. & Chevillard, C. Chagas' disease cardiomyopathy: immunopathology and genetics. Mediators Inflamm. 2014, 683230 (2014).
Hussain, H., Al-Harrasi, A., Al-Rawahi, A., Green, I. R. & Gibbons, S. Fruitful decade for antileishmanial compounds from 2002 to late 2011. Chem. Rev. 114, 10369–10428 (2014).
McLatchie, A. P. et al. Highly sensitive in vivo imaging of Trypanosoma brucei expressing “red-shifted” luciferase. PLoS Negl. Trop. Dis. 7, e2571 (2013). This study details a key mouse model for HAT.
Burrell-Saward, H., Rodgers, J., Bradley, B., Croft, S. L. & Ward, T. H. A sensitive and reproducible in vivo imaging mouse model for evaluation of drugs against late-stage human African trypanosomiasis. J. Antimicrob. Chemother. 70, 510–517 (2015).
Giroud, C. et al. Murine models for Trypanosoma brucei gambiense disease progression — from silent to chronic infections and early brain tropism. PLoS Negl. Trop. Dis. 3, e509 (2009).
Jennings, F. W., Urquhart, G. M., Murray, P. K. & Miller, B. M. Treatment with suranim and 2-substituted 5-nitroimidazoles of chronic murine Trypanosoma brucei infections with central nervous system involvement. Trans. R. Soc. Trop. Med. Hyg. 77, 693–698 (1983).
Myburgh, E. et al. In vivo imaging of trypanosome–brain interactions and development of a rapid screening test for drugs against CNS stage trypanosomiasis. PLoS Negl. Trop. Dis. 7, e2384 (2013).
Romanha, A. J. et al. In vitro and in vivo experimental models for drug screening and development for Chagas' disease. Mem. Inst. Oswaldo Cruz 105, 233–238 (2010).
Bustamante, J. M. & Tarleton, R. L. Methodological advances in drug discovery for Chagas' disease. Expert Opin. Drug Discov. 6, 653–661 (2011).
Lewis, M. D. et al. Bioluminescence imaging of chronic Trypanosoma cruzi infections reveals tissue-specific parasite dynamics and heart disease in the absence of locally persistent infection. Cell. Microbiol. 16, 1285–1300 (2014). This study details a key mouse model for Chagas disease.
Bustamante, J. M., Craft, J. M., Crowe, B. D., Ketchie, S. A. & Tarleton, R. L. New, combined, and reduced dosing treatment protocols cure Trypanosoma cruzi infection in mice. J. Infect. Dis. 209, 150–162 (2014). This study details another key mouse model for Chagas disease.
Canavaci, A. M. et al. In vitro and in vivo high-throughput assays for the testing of anti-Trypanosoma cruzi compounds. PLoS Negl. Trop. Dis. 4, e740 (2010).
Lewis, M. D., Francisco, A. F., Taylor, M. C. & Kelly, J. M. A new experimental model for assessing drug efficacy against Trypanosoma cruzi infectin based on highly sensitive in vivo imaging. J. Biomol. Screen. 20, 36–43 (2015).
Gupta, S. Visceral leishmaniaisis: experimental models for drug discovery. Indian J. Med. Res. 133, 27–39 (2011).
Achterberg, V. & Gercken, G. Cytotoxicity of ester and ether lysophospholipids on Leishmania donovani promastigotes. Mol. Biochem. Parasitol. 23, 117–122 (1987).
Plimmer, H. G. & Thomson, J. D. Further results of the experimental treatment of trypanosomiasis in rats; being a progress report of a committee of the Royal Society. Proc. R. Soc. Lond. B Biol. Sci. 80, 1–10 (1908).
Acknowledgements
The authors are grateful to the following funding bodies for financial support: The Wellcome Trust (strategic grants 077705, 083481, 092340, 100476 and 105021), Wellcome Trust Senior Investigator Awards to D.H., M.F. and M.A.J.F. (grants 100320/Z/12/Z, 204697/Z/16/Z and 101842/Z/13/Z) and a Wellcome Trust Principal Research Fellowship to A.H.F (grant 079838); Drugs for Neglected Diseases Initiative (DNDi); and the UK Medical Research Council for grants to M.C.F. (MR/L018853/1 and MR/P009018/1), D.H. (MR/K000500/1) and to M.C.F and D.H. (MR/K008749/1). The authors thank A. Baliani with help in the preparation of figure 3.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- Trypanosomatid
-
A member of the order Kinetoplastida (suborder Trypanosomatida), a group of protozoan flagellates that includes many pathogenic species. Trypanosomatid is frequently used interchangeably with kinetoplastid.
- Eradication
-
The permanent reduction of the global incidence of infection or disease to zero.
- Elimination
-
Zero incidence of infection or disease in a defined geographical area.
- Parenteral administration
-
Drug administration by routes other than through the gastrointestinal tract, generally by injection.
- Insect vectors
-
Pathogenic trypanosomatids are commonly transmitted by insect species that are specific for the respective parasite. The geographical distribution of these insects restricts the range of parasite transmission.
- Chemical series
-
A series of chemicals that have closely related chemical structures.
- Suicide inhibitor
-
A compound that is activated by an enzyme to give a reactive intermediate that irreversibly inhibits the enzyme through covalent bonding.
- Druggable
-
A protein that can be inhibited or its function modulated by a drug-like molecule.
- Kinomes
-
All protein kinases in certain organisms.
- Kinetochore
-
A protein complex that assembles at the centromeres of chromosomes and is important for chromosome segregation during cell division.
- Polycistronic
-
Polycistronic transcription produces an mRNA that encodes several polypeptides in one molecule, which is then processed into individual polypeptide mRNAs.
- trans-splicing
-
A process that is similar to cis-splicing but, in this case, two different transcripts are spliced together.
- cis-splicing
-
A step in pre-mRNA maturation during which exons are spliced together and introns are removed.
- Structure-based drug design
-
The use of 3D structures of the inhibitors or modulators that are bound to a target protein (derived from X-ray crystallography or NMR) and computational chemistry to aid the design and optimization of lead compounds.
- Drug-like molecule
-
A molecule that has the potential to be an oral drug. Such a molecule will generally follow Lipinskis rule of five: have a molecular weight of less than 500, a cLogP (measure of hydrophobicity) value of less than 5, less than 5 hydrogen bond donors and less than 10 hydrogen bond acceptors.
- Amastigotes
-
The forms of Trypanosoma cruzi and Leishmania spp. that resides within cells of a human host. Amastigotes are rounded and lack a free flagellum.
- Pharmacokinetic
-
Relating to the effect that the body has on a drug.
- Pharmacodynamic
-
Relating to the effects of drugs in the body.
- Polypharmacology
-
Drugs that act through the inhibition or modulation of more than one molecular target or disease pathway.
- Phenotypic screening
-
An approach that uses a whole-cell screen that is designed to identify the effects of a compound on a target cell or pathogen without a need to understand the underlying mode of action. Through the use of high-content screening, several phenotypes can be detected simultaneously; for example, the effects on intracellular parasite viability and host cell viability (that is, toxicity).
- CRISPR–Cas9
-
A prokaryotic immune system that has been repurposed for genome editing in eukaryotic cells; in prokaryotes, the system comprises clustered regularly interspaced short palindromic repeats and the programmable Cas9 nuclease.
- Scaffold hopping
-
The modification of the essential core of a molecule to produce a new core molecule that has broadly similar, but slightly different, properties. This is an approach that is generally used to optimize a hit or lead, improving features such as biological activity, solubility or metabolic stability.
- Vector optimization
-
The modification of substituents on the core of a molecule (vectors) to improve a property, or properties, of a lead molecule (for example biological activity or solubility). It may also encompass optimizing the position on the core scaffold at which a substituent is placed.
Rights and permissions
About this article
Cite this article
Field, M., Horn, D., Fairlamb, A. et al. Anti-trypanosomatid drug discovery: an ongoing challenge and a continuing need. Nat Rev Microbiol 15, 217–231 (2017). https://doi.org/10.1038/nrmicro.2016.193
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2016.193
This article is cited by
-
Anti-trypanosomatid drug discovery: progress and challenges
Nature Reviews Microbiology (2023)
-
In vitro trypanocidal activities and structure–activity relationships of ciprofloxacin analogs
Molecular Diversity (2023)
-
Menthol carbonates as potent antiparasitic agents: synthesis and in vitro studies along with computer-aided approaches
BMC Complementary Medicine and Therapies (2022)
-
Impact of pulmonary African trypanosomes on the immunology and function of the lung
Nature Communications (2022)
-
Phenylbenzothiazole derivatives: effects against a Trypanosoma cruzi infection and toxicological profiles
Parasitology Research (2021)