Abstract
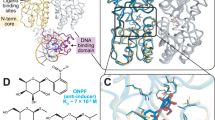
The side chains of Arg 31, Glu 36 and Arg 40 in Arc repressor form a buried salt-bridge triad. The entire salt-bridge network can be replaced by hydrophobic residues in combinatorial randomization experiments resulting in active mutants that are significantly more stable than wild type. The crystal structure of one mutant reveals that the mutant side chains pack against each other in an otherwise wild-type fold. Thus, simple hydrophobic interactions provide more stabilizing energy than the buried salt bridge and confer comparable conformational specificity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Barlow, D.J. & Thornton, J.M. Ion-pairs in proteins. J. molec. Biol. 168, 867–885(1983).
Perutz, M. & Raidt, H. Stereochemical basis of heat stability in bacterial ferredoxins and in haemoglobin A2. Nature 255, 256–259 (1975).
Perutz, M. Electrostatic effects in proteins. Science 201, 1187–1191 (1978).
Ruegg, C., Ammer, D. & Lerch, K. Comparison of amino acid sequence and thermostability of tyrosinase from three wild type strains of Neurospora crassa. J. biol. Chem. 257, 6420–6426 (1982).
Fersht, A.R. Conformational equilibria in α- and δ-chymotrypsin. The energetics and importance of the salt bridge. J. molec. Biol. 64, 497–509 (1972).
Anderson, D.E., Becktel, W.J. & Dahlquist, F.W. pH-induced denaturation of proteins: A single salt bridge contributes 3–5 kcal/mol to the free energy of folding of T4 lysozyme. Biochemistry 29, 2403–2408 (1990).
Horovitz, A., Serrano, L., Avron, B., Bycroft, M. & Fersht, A.R. Strength and co-operativity of contributions of surface salt bridges to protein stability. J. molec. Biol. 216, 1031–1044 (1990).
Erwin, C.R., Barnett, B.L., Oliver, J.D. & Sullivan, J.F. Effects of engineered salt bridges on the stability of subtilisin BPN'. Prot. Engng. 4, 87–97 (1990).
Sali, D., Bycroft, M. & Fersht, A.R. Surface electrostatic interactions contribute little to stability of barnase. J. molec. Biol. 220, 779–778 (1991).
Dao-Pin, S., Sauer, U., Nicholson, H. & Matthews, B.W. Contributions of engineered surface salt bridges to the stability of T4 lysozyme determined by directed mutagenesis. Biochemistry 30, 7142–7153 (1991).
Hendsch, Z.S. & Tidor, B. Do salt bridges stabilize proteins? A continuum electrostatic analysis. Prot. Sci. 3, 211–226 (1994).
Raumann, B.E., Rould, M.A., Pabo, C.O. & Sauer, R.T. DNA recognition by β-sheets in the Arc repressor-operator crystal structure. Nature 367, 754–757 (1994).
Vershon, A.K., Bowie, J.U., Karplus, T.M. & Sauer, R.T. Isolation and analysis of Arc repressor mutants: Evidence for an unusual mechanism of DNA binding. Proteins 1, 302–311 (1986).
Bowie, J.U. & Sauer, R.T. Identifying determinants of folding and activity for a protein of unknown structure. Proc. natn. Acad. Sci. U.S.A. 86, 2152–2156 (1989).
Milla, M.E., Brown, B.M. & Sauer, R.T. Protein stability effects of a complete set of alanine substitutions in Arc repressor. Nature struct. Biol. 1, 518–523 (1994).
Brown, B.M., Milla, M.E., Smith, T.L. & Sauer, R.T. Scanning mutagenesis of the Arc repressor as a functional probe of operator recognition. Nature struct. Biol. 1, 164–168 (1994).
Horovitz, A. & Fersht, A.R. Strategy for analysing the co-operativity of intramolecular interactions in peptides and proteins. J. molec. Biol. 214, 613–617 (1990).
Bowie, J.U. & Sauer, R.T. Equilibrium dissociation and unfolding of the Arc repressor dimer. Biochemistry 28, 7139–7143 (1989).
Chothia, C. The nature of the accessible and buried surfaces in proteins. J molec. Biol. 105, 1–14 (1976).
O'Shea, E.K., Klemm, J.D., Kim, P.S. & Alber, T. X-ray structure of the GCN4 leucine zipper, a two-stranded, parallel coiled coil. Science 254, 539–544 (1991).
Harbury, P.B., Zhang, T., Kim, P.S. & Alber, T. A switch between two-, three-, and four-stranded coiled coils in GCN4 leucine zipper mutants. Science 262, 1401–1407 (1993).
Milla, M.E., Brown, B.M. & Sauer, R.T. P22 Arc repressor: Enhanced expression of unstable mutants by addition of polar C-terminal sequences. Prot. Sci. 2, 2198–2205 (1993).
Bowie, J.U. & Sauer, R.T. Identification of C-terminal extensions that protect proteins from intracellular proteolysis. J. biol. Chem. 264, 7596–7602 (1989).
Brown, B.M. & Sauer, R.T. Assembly of the Arc repressor-operator complex: Cooperative interactions between DNA-bound dimers. Biochemistry 32, 1354–1363 (1993).
Milla, M.E. & Sauer, R.T. P22 Arc repressor: Folding kinetics of a single-domain, dimeric protein. Biochemistry 33, 1125–1133 (1994).
Breg, J.N., van Opheusden, J.H.J., Burgering, M.J., Boelens, R. & Kaptein, R. Structure of Arc repressor in solution: Evidence for a family of β-sheet DNA-binding proteins. Nature 346, 586–589 (1990).
Bonvin, A.M.J.J., Vis, H., Breg, J.N., Burgering, M.J.M., Boelens, R. & Kaptein, R. Nuclear magnetic resonance solution structure of the Arc repressor using relaxation matrix calculations. J. molec. Biol. 236, 328–341 (1994).
Chakrabartty, A., Kortemme, T., Padmanabhan, S. & Baldwin, R.L. Aromatic side-chain contribution to far-ultraviolet circular dichroism of helical peptides and its effect on measurement of helix propensities. Biochemistry. 32, 5560–5565 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Waldburger, C., Schildbach, J. & Sauer, R. Are buried salt bridges important for protein stability and conformational specificity?. Nat Struct Mol Biol 2, 122–128 (1995). https://doi.org/10.1038/nsb0295-122
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0295-122
This article is cited by
-
Loss of a conserved salt bridge in bacterial glycosyl hydrolase BgIM-G1 improves substrate binding in temperate environments
Communications Biology (2018)
-
Salt Bridge in Aqueous Solution: Strong Structural Motifs but Weak Enthalpic Effect
Scientific Reports (2018)
-
A new structural arrangement in proteins involving lysine NH3 + group and carbonyl
Scientific Reports (2017)
-
DBAC: A simple prediction method for protein binding hot spots based on burial levels and deeply buried atomic contacts
BMC Systems Biology (2011)
-
Adhesive water networks facilitate binding of protein interfaces
Nature Communications (2011)