Abstract
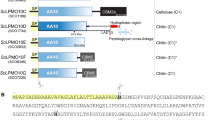
Cellulase E4 from Thermomonospora fusca is unusual in that it has characteristics of both exo- and endo-cellulases. Here we report the crystal structure of a 68K Mr fragment of E4 (E4-68) at 1.9 Å resolution. E4-68 contains both a family 9 catalytic domain, exhibiting an (α/α)6 barrel fold, and a family III cellulose binding domain, having an antiparallel β-sandwich fold. While neither of these folds is novel, E4-68 provides the first cellulase structure having interacting catalytic and cellulose binding domains. The complexes of E4-68 with cellopentaose, cellotriose and cellobiose reveal conformational changes associated with ligand binding and allow us to propose a catalytic mechanism for family 9 enzymes. We also provide evidence that E4 has two novel characteristics: first it combines exo- and endo- activities and second, when it functions as an exo-cellulase, it cleaves off cellotetraose units.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Walker, L. & Wilson, D. Engineering cellulases. Biores Technol 36, 3–14 (1991).
Wyman, C.E., Bain, R.L., Hinman, N.D. & Stevens, D.J. Renewable energy: sources for fuels and electricity. (Island Press, Washington, DC; 1993).
Davies, G. & Henrissat, B. Structures and mechanisms of glycosyl hydrolases. Structure 3, 853–859 (1995).
Tomme, P., Warren, R.A.J., Miller, R.C.J., Kilburn, D.G. & Gilkes, N.R. Cellulose-binding domains: Classification and properties in ACS Symposium Series (eds Saddler, J.N. & Penner, M.H.) 142–163 (American Chemical Society, Washington DC; 1994).
Henrissat, B. & Bairoch, A. A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem. J. 293, 781–788 (1993).
Bairoch, A., Bucher, P. & Hoffmann, K. The PROSITE database. Nucl. Acid. Res. 25 217–221 (1997).
Irwin, D.C., Spezio, M., Walker, L.P. & Wilson, D.B. Activity studies of eight purified cellulases specificity synergism and binding domain effects. Biotechnol. Bioeng. 42, 1002–1013 (1993).
Barr, B.K., Hsieh, Y.L., Ganem, B. & Wilson, D.B. Identification of two functionally different classes of exocellulases. Biochemistry 35, 586–592 (1996).
Juy, M., et al. Three-dimensional structure of a thermostable bacterial cellulase. Nature 357, 89–91 (1992).
Tormo, J., et al. Crystal structure of a bacterial family-Ill cellulose-binding domain: a general mechanism for attachment to cellulose. EMBO J 15, 5739–5751 (1996).
Jauris, S., et al. Sequence analysis of the Clostridium stercorarium celZ gene encoding a thermoactive cellulase (Avicelase I): identification of catalytic and cellulose-binding domains. Mol. Gen. Genet. 223, 258–267 (1990).
Bronnenmeier, K. & Staudenbauer, W.L. Resolution of Clostridium stercorarium cellulase by fast protein liquid chromatography (FPLC). Appl. Microbiol. Biotechnol. 27, 432–436 (1988).
Meinke, A., et al. Unusual sequence organization in CenB, an inverting endoglucanase from Cellulomonas fimi. J Bacteriol 173, 308–314 (1991).
Sander, C. & Schneider, R. Database of homology-derived protein structures and the structural meaning of sequence alignment. Proteins Struct Funct Genet 9, 56–68 (1991).
Diederichs, K. Structural superposition of proteins with unknown alignment and detection of topological similarity using a six-dimensional search algorithm. Proteins Struct. Funct. Gen. 23, 187–195 (1995).
Alzari, P.M., Souchon, H. & Dominguez, R. The crystal structure of endoglucanase CelA, a family 8 glycosyl hydrolase from Clostridium thermocellum. Structure 4, 265–275 (1996).
Aleshin, A., Golubev, A., Firsov, L.M. & Honzatko, R.B. Crystal structure of glucoamylase from Aspergillus awamori var-x100 to 2.2 Å resolution. J. Biol. Chem. 267, 19291–19298 (1992).
Flanagen, K., Walahaw, J., Price, S.L. & Goodfellow, J.M. Solvent interactions with pi ring systems in proteins. Protein Engng. 8, 109–116 (1995).
Xu, G.Y., et al. Solution structure of a cellulose-binding domain from Cellulomonas fimi by nuclear magnetic resonance spectroscopy. Biochemistry 34, 6993–7009 (1995).
Tews, I., et al. Bacterial chitobiase structure provides insight into catalytic mechanism and the basis of Tay-Sachs disease. Nature Struct. Biol. 3, 638–648 (1996).
Shimon, L.J.W., et al. A cohesin domain from Clostridium thermocellum: the crystal structure provides new insights into cellulosome assembly. Structure 5, 381–390 (1997).
Davies, G.J., Tolley, S.P., Henrissat, B., Hjort, C. & Schulein, M. Structures of oligosaccharide-bound forms of the endoglucanase V from Humicola insolens at 1.9 Å resolution. Biochemistry 34, 16210–16220 (1995).
Spezio, M., Wilson, D.B. & Karplus, P.A. Crystal structure of the catalytic domain of a thermophilic endocellulase. Biochemistry 32, 9906–9916 (1993).
Rouvinen, J., Bergfors, T., Teeri, T., Knowles, J.K.C. & Jones, T.A. Three-dimensional structure of cellobiohydrolase II from Trichoderma reesei. Science 249, 380–386 (1990).
Jung, E.D., et al. Dna sequences and expression in streptomyces-lividans of an exoglucanase gene and an endoglucanase gene from Thermomonospora fusca. Appl Environ Microbiol 59, 3032–3043 (1993).
Koshland, D.E.J. Stereochemistry and the mechanism of enzymatic reactions. Biol. Rev. 28, 416–436 (1953).
Chauvaux, S., Beguin, P. & Aubert, J.P. Site-directed mutagenesis of essential carboxylic residues in Clostridium thermocellum endoglucanase celd. J Biol Chem 267, 4472–4478 (1992).
Tomme, P., et al. Identification of a histidyl residue in the active center of endoglucanase d from clostridium-thermocellum. J. Biol. Chem. 266, 10313–10318 (1991).
Tomme, P., Van, B.J. & Claeyssens, M. Modification of catalytically important carboxyl residues in endoglucanase D from Clostridium thermocellum. Biochem. J. 285, 319–324 (1992).
Tanaka, Y., Tao, W., Blanchard, J.S. & Hehre, E.J. Transition state structures for the hydrolysis of alpha-D-glucopyranosyl fluoride by retaining and inverting reactions of glycosylases. J. Biol. Chem. 269, 32306–32312 (1994).
Sakon, J., Adney, W.S., Himmel, M.E., Thomas, S.R. & Karplus, P.A. Crystal structure of thermostable family 5 endocellulase E1 from Acidothermus cellulolyticus in complex with cellotetraose. Biochemistry 35, 10648–10660 (1996).
McCarter, J.D. & Withers, S.G. Mechanisms of enzymatic glycoside hydrolysis. Curr. Opin. Struct. Biol. 4, 885–892 (1994).
Sacemski, I.I. & Lienhard, G.E. The effect of gadolinium ion on the binding of inhibitors and substrates to lysozyme. J. Biol. Chem. 249, 2932–2938 (1974).
Poole, D.M., Hazlewood, G.P., Huskisson, N.S., Virden, R. & Gilbert, H.J. The role of conserved tryptophan residues in the interaction of a bacterial cellulose binding domain with its ligand. FEMS Microbiol Lett 106, 77–83 (1993).
Din, N., et al. The cellulose-binding domain of endoglucanase A (CenA) from Cellulomonas fimi: Evidence for the involvement of tryptophan residues in binding. Mol. Microbiol. 11, 747–755 (1994).
Linder, M., et al. Identification of functionally important amino acids in the cellulose-binding domain of Trichoderma reesei cellobiohydrolase I. Prot. Sci. 4, 1056–1064 (1995).
Barr, B.K. & Wilson, D.B. unpublished result. (1997). [AUTHOR: this cannot be cited in the reference list. Must be cited in the text as (B.K. Barr & D.B. Wilson, unpublished results) and removed from here. You must then renumber the following refs in the ref. list and throughout the text]
Zhang, S., Lao, G. & Wilson, D.B. Characterization of a Thermomonospora fusca exocellulase. Biochemistry 34, 3386–3395 (1995).
Cudney, B., Patel, S., Weisgraber, K., Newhouse, Y. & McPherson, A. Screening and optimization strategies for macromolecular crystal growth. Acta Crystallogr. Sect D 50, 414–423 (1994).
Jancarik, J. & Kim, S.H. Sparse matrix sampling: a screening method for crystallization of proteins. J. Appl. Crystallogr. 24, 409–411 (1991).
Hamlin, R. Multiwire area X-ray diffractometers in Methods in Enzymology-Diffraction Methods for Biological Macromolecules (eds. Wyckoff, H.W., Hirs, C.H.W. & Timasheff, S.N.) 416–451 (Academic Press, Orlando, Florida; 1985).
Otwinowski, Z. Oscillation data reduction program. 1-56-62 (SERC Daresbury Laboratory, Warrington, UK; 1993).
Sheldrick, G.M. Phase annealing in SHELX-90: direct methods for larger structures. Acta Crystallogr Sec A 46, 467–473 (1990).
Furey, W. & Swaminathan, S. PHASES-95: a program package for the processing and analysis of diffraction data from macromolecules in Methods in Enzymology: macromolecular crystallography (eds Carter, C. & Sweet, R.) in the press (Academic Press, Orlando, Florida; 1996).
Sack, J.S. Chain - a crystallographic modeling program. J. Mol. Graph. 6, 224–225 (1988).
Brünger, A.T., Kuriyan, J. & Karplus, M. Crystallographic R factor refinement by molecular dynamics. Science 235, 458–460 (1987).
Engh, R.A. & Huber, R. Accurate bond and angle parameters for X-ray protein-structure refinement. Acta Crystallogr. Sect A 47, 392–400 (1991).
Lao, G., Ghangas, G.S., Jung, E.D. & Wilson, D.B. DNA sequences of three beta-1 4-endoglucanase genes from Thermomonospora fusca. J Bacteriol 173, 3397–3407 (1991).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Bagnara-Tardif, C., et al. Sequence analysis of a gene cluster encoding cellulases from Clostridium cellulolyticum. Gene 119, 17–28 (1992).
Hazlewood, G.P., Davidson, K., Laurie, J.I., Huskisson, N.S. & Gilbert, H.J. Gene sequence and properties of Cell, a family E endoglucanase from Clostridium thermocellum. J. Gen. Microbiol. 139, 307–316 (1993).
Navarro, A., Chebrou, M.C., Beguin, P. & Aubert, J.P. Nucleotide sequence of the cellulase gene celF of Clostridium thermocellum. Res. Microbiol. 142, 927–936 (1991).
Shoseyov, O., Hamamoto, T., Foong, F. & Doi, R.H. Cloning of Clostridium cellulovorans endo-1,4-beta-glucanase genes. Biochem. Biophys. Res. Commun. 169, 667–672 (1990).
Kraulis, P. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
IUPAC-IUB. Symbols for specifying the conformation of polysaccharide chains. Eur. J. Biochem. 131, 5–7 (1983).
Gessler, K., et al. Crystal structure of beta-D-cellotetraose hemihydrate with implications for the structure of cellulose II. Science 266, 1027–1029 (1994).
Taylor, J.S., Teo, B., Wilson, D.B. & Brady, J.W. Conformational modeling of substrate binding to endocellulase E2 from Thermomonospora fusca. Protein Engng. 8, 1145–1152 (1995).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281–296 (1991).
Diederichs, K. & Karplus, P.A. Improved R-factors for diffraction data analysis in macromolecular crystallography. Nature Struct. Biol. 4, 269–275 (1997).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Sakon, J., Irwin, D., Wilson, D. et al. Structure and mechanism of endo/exocellulase E4 from Thermomonospora fusca. Nat Struct Mol Biol 4, 810–818 (1997). https://doi.org/10.1038/nsb1097-810
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb1097-810
This article is cited by
-
Mapping the deformability of natural and designed cellulosomes in solution
Biotechnology for Biofuels and Bioproducts (2022)
-
Identification and Characterization of a Glycoside Hydrolase Family 9 Member from the Digestive Gland of the Snail Achatina fulica
BioEnergy Research (2022)
-
A processive GH9 family endoglucanase of Bacillus licheniformis and the role of its carbohydrate-binding domain
Applied Microbiology and Biotechnology (2022)
-
Impacts of cotton linter pulp characteristics on the processivity of glycoside hydrolase family 5 endoglucanase from Volvariella Volvacea
Cellulose (2021)
-
New thermostable endoglucanase from Spirochaeta thermophila and its mutants with altered substrate preferences
Applied Microbiology and Biotechnology (2021)