Abstract
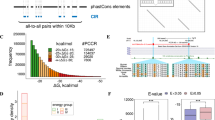
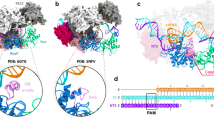
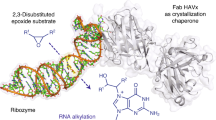
The hammerhead ribozyme (HHRz) is a small, naturally occurring ribozyme that site-specifically cleaves RNA and has long been considered a potentially useful tool for gene silencing. The minimal conserved HHRz motif derived from natural sequences consists of three helices that intersect at a highly conserved catalytic core of 11 nucleotides. The presence of this motif is sufficient to support cleavage at high Mg2+ concentrations, but not at the low Mg2+ concentrations characteristic of intracellular environments. Here we demonstrate that natural HHRzs require the presence of additional nonconserved sequence elements outside of the conserved catalytic core to enable intracellular activity. These elements may stabilize the HHRz in a catalytically active conformation via tertiary interactions. HHRzs stabilized by these interactions cleave efficiently at physiological Mg2+ concentrations and are functional in vivo. The proposed role of these tertiary interacting motifs is supported by mutational, functional, structural and molecular modeling analysis of natural HHRzs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Ruffner, D.E., Stormo, G.D. & Uhlenbeck, O.C. Sequence requirements of the hammerhead RNA self-cleavage reaction. Biochemistry 29, 10695–10702 (1990).
Darnell, J., Lodish, H.F. & Baltimore, D. Molecular Cell Biology (W.H. Freeman, New York, 1986).
Deshler, J.O., Li, H., Rossi, J.J. & Castanotto, D. Ribozymes expressed within the loop of a natural antisense RNA form functional transcription terminators. Gene 155, 35–43 (1995).
Buzayan, J.M., Hampel, A. & Bruening, G. Nucleotide sequence and newly formed phosphodiester bond of spontaneously ligated satellite tobacco ringspot virus RNA. Nucleic Acids Res. 14, 9729–9743 (1986).
Forster, A.C. & Symons, R.H. Self-cleavage of virusoid RNA is performed by the proposed 55-nucleotide active site. Cell 50, 9–16 (1987).
Hernandez, C. & Flores, R. Plus and minus RNAs of peach latent mosaic viroid self-cleave in vitro via hammerhead structures. Proc. Natl. Acad. Sci. USA 89, 3711–3715 (1992).
Long, D.M. & Uhlenbeck, O.C. Kinetic characterization of intramolecular and intermolecular hammerhead RNAs with stem II deletions. Proc. Natl. Acad. Sci. USA 91, 6977–6981 (1994).
Long, D.M., LaRiviere, F.J. & Uhlenbeck, O.C. Divalent metal ions and the internal equilibrium of the hammerhead ribozyme. Biochemistry 34, 14435–14440 (1995).
Pley, H.W., Flaherty, K.M. & McKay, D.B. Three-dimensional structure of a hammerhead ribozyme. Nature 372, 68–74 (1994).
Scott, W.G., Finch, J.T. & Klug, A. The crystal structure of an all-RNA hammerhead ribozyme: a proposed mechanism for RNA catalytic cleavage. Cell 81, 991–1002 (1995).
Hainzl, T., Huang, S. & Sauer-Eriksson, A.E. Structure of the SRP19 RNA complex and implications for signal recognition particle assembly. Nature 417, 767–771 (2002).
Oubridge, C., Kuglstatter, A., Jovine, L. & Nagai, K. Crystal structure of SRP19 in complex with the S domain of SRP RNA and its implication for the assembly of the signal recognition particle. Mol. Cell 9, 1251–1261 (2002).
Berman, H.M. et al. The nucleic acid database. A comprehensive relational database of three-dimensional structures of nucleic acids. Biophys. J. 63, 751–759 (1992).
Massire, C. & Westhof, E. MANIP: an interactive tool for modelling RNA. J. Mol. Graph. Mod. 16, 197–205 (1999).
McKay, D.B. Structure and function of the hammerhead ribozyme: an unfinished story. RNA 2, 395–403 (1996).
Hammann, C., Norman, D.G. & Lilley, D.M. Dissection of the ion-induced folding of the hammerhead ribozyme using 19F NMR. Proc. Natl. Acad. Sci. USA 98, 5503–5508 (2001).
Esteban, J.A., Banerjee, A.R. & Burke, J.M. Kinetic mechanism of the hairpin ribozyme. Identification and characterization of two nonexchangeable conformations. J. Biol. Chem. 272, 13629–13639 (1997).
Zhao, Z.-Y. et al. The folding of the hairpin ribozyme: dependence on the loops and the junction. RNA 6, 1833–1846 (2000).
Walter, N.G., Burke, J.M. & Millar, D.P. Stability of hairpin ribozyme tertiary structure is governed by the interdomain junction. Nat. Struct. Biol. 6, 544–549 (1999).
Stage-Zimmermann, T.K. & Uhlenbeck, O.C. A covalent crosslink converts the hammerhead ribozyme from a ribonuclease to an RNA ligase. Nat. Struct. Biol. 8, 863–867 (2001).
Blount, K.F. & Uhlenbeck, O.C. Internal equilibrium of the hammerhead ribozyme is altered by the length of certain covalent cross-links. Biochemistry 41, 6834–6841 (2002).
Fedor, M.J. Tertiary structure stabilization promotes hairpin ribozyme ligation. Biochemistry 38, 11040–11050 (1999).
Pan, J., Thirumalai, D. & Woodson, S.A. Magnesium-dependent folding of self-splicing RNA: exploring the link between cooperativity, thermodynamics, and kinetics. Proc. Natl. Acad. Sci. USA 96, 6149–6154 (1999).
Pan, J. & Woodson, S.A. The effect of long-range loop-loop interactions on folding of the Tetrahymena self-splicing RNA. J. Mol. Biol. 294, 955–965 (1999).
Lehnert, V., Jaeger, L., Michel, F. & Westhof, E. New loop-loop tertiary interactions in self-splicing introns of subgroup IC and ID: a complete 3D model of the Tetrahymena thermophila ribozyme. Chem. Biol. 3, 993–1009 (1996).
Collins, M.L. et al. A branched DNA signal amplification assay for quantification of nucleic acid targets below 100 molecules/ml. Nucleic Acids Res. 25, 2979–2984 (1997).
Milligan, J.F. & Uhlenbeck, O.C. Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol. 180, 51–62 (1989).
Stage-Zimmermann, T.K. & Uhlenbeck, O.C. Hammerhead ribozyme kinetics. RNA 4, 875–889 (1998).
Massire, C., Gaspin, C. & Westhof, E. DRAWNA: a program for drawing schematic views of nucleic acids. J. Mol. Graph. 12, 201–206 (1994).
Hertel, K.J. et al. Numbering system for the hammerhead. Nucleic Acids Res. 20, 12 (1992).
Gorodkin, J., Knudsen, B., Zwieb, C. & Samuelsson, T. SRPDB (Signal Recognition Particle Database). Nucleic Acids Res. 29, 169–170 (2001).
Leontis, N.B. & Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 7, 499–512 (2001).
Acknowledgements
We thank O. Uhlenbeck, A. Wolfson, J. Rossi and D. Lilley for insightful discussions and critical reading of this manuscript, S. Suggs for comments and encouragement and A. Reynolds, M. Brewer and W. Marshall for their help and support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Khvorova, A., Lescoute, A., Westhof, E. et al. Sequence elements outside the hammerhead ribozyme catalytic core enable intracellular activity. Nat Struct Mol Biol 10, 708–712 (2003). https://doi.org/10.1038/nsb959
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb959
This article is cited by
-
Highly efficient expression of circular RNA aptamers in cells using autocatalytic transcripts
Nature Biotechnology (2019)
-
Massively parallel RNA device engineering in mammalian cells with RNA-Seq
Nature Communications (2019)