Abstract
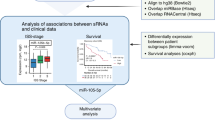
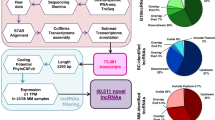
Although long intergenic non-coding RNAs (lincRNA) role in various cancers is described, their significance in Multiple Myeloma (MM) remains poorly defined. Here we have studied the lincRNA profile and their clinical impact in MM. We performed RNA-seq on MM cells from 308 newly diagnosed and uniformly treated patients, 16 normal plasma cells and utilized RNA-seq data from 532 newly diagnosed patients from CoMMpass study to analyze for lincRNAs. We observed 869 differentially expressed lincRNAs in MM compared to normal plasma cells. We identified 14 lincRNAs associated with PFS and calculated a risk score to stratify patients. The median PFS between high vs low-risk groups was 17 months vs not-reached (NR); and OS 30 months vs NR, respectively (p < 0.0001 for both). In the independent validation dataset between high and low-risk groups, PFS was 27 vs 42 months (HR 2.06 [1.44−2.96]; p < 0.0005); and 4-year OS 62% vs 86% (HR 2.76 [1.51–5.05]; p < 0.0005) confirming significant clinical relevance of lincRNA in MM. Importantly, lincRNA signature was able to further identify patients with significant differential outcomes within each low and high-risk categories identified using standard risk categorization including cytogenetic/FISH, ISS, and MRD negative or positive. Our results suggest that lincRNAs have an independent effect on MM outcome and provide a rationale to evaluate its molecular and biological impact.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Morris KV, Mattick JS. The rise of regulatory RNA. Nat Rev Genet. 2014;15:423–37.
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012;22:1775–89.
Harrow J, Frankish A, Gonzalez JM, Tapanari E, Diekhans M, Kokocinski F, et al. GENCODE: the reference human genome annotation for the ENCODE Project. Genome Res. 2012;22:1760–74.
Huarte M. The emerging role of lncRNAs in cancer. Nat Med. 2015;21:1253–61.
Zheng GX, Do BT, Webster DE, Khavari PA, Chang HY. Dicer-microRNA-Myc circuit promotes transcription of hundreds of long noncoding RNAs. Nat Struct Mol Biol. 2014;21:585–90.
Zhang X, Gejman R, Mahta A, Zhong Y, Rice KA, Zhou Y, et al. Maternally expressed gene 3, an imprinted noncoding RNA gene, is associated with meningioma pathogenesis and progression. Cancer Res. 2010;70:2350–8.
Fulciniti M, Amodio N, Bandi RL, Cagnetta A, Samur MK, Acharya C, et al. miR-23b/SP1/c-myc forms a feed-forward loop supporting multiple myeloma cell growth. Blood Cancer J. 2016;6:e380.
Zarate R, Boni V, Bandres E, Garcia-Foncillas J. MiRNAs and LincRNAs: could they be considered as biomarkers in colorectal cancer?. Int J Mol Sci. 2012;13:840–65.
Yang N, Chen J, Zhang H, Wang X, Yao H, Peng Y, et al. LncRNA OIP5-AS1 loss-induced microRNA-410 accumulation regulates cell proliferation and apoptosis by targeting KLF10 via activating PTEN/PI3K/AKT pathway in multiple myeloma. Cell Death Dis. 2017;8:e2975.
Gao D, Lv AE, Li HP, Han DH, Zhang YP. LncRNA MALAT-1 elevates HMGB1 to promote autophagy resulting in inhibition of tumor cell apoptosis in multiple myeloma. J Cell Biochem. 2017;118:3341–8.
Ronchetti D, Manzoni M, Todoerti K, Neri A, Agnelli L. In silico characterization of miRNA and long non-coding RNA interplay in multiple myeloma. Genes (Basel). 2016;29;7(12). pii: E107.
Meng YB, He X, Huang YF, Wu QN, Zhou YC, Hao DJ. Long noncoding RNA CRNDE promotes multiple myeloma cell growth by suppressing miR-451. Oncol Res. 2017;25:1207–14.
Li B, Chen P, Qu J, Shi L, Zhuang W, Fu J, et al. Activation of LTBP3 gene by a long noncoding RNA (lncRNA) MALAT1 transcript in mesenchymal stem cells from multiple myeloma. J Biol Chem. 2014;289:29365–75.
Ronchetti D, Agnelli L, Taiana E, Galletti S, Manzoni M, Todoerti K, et al. Distinct lncRNA transcriptional fingerprints characterize progressive stages of multiple myeloma. Oncotarget. 2016;7:14814–30.
Cho SF, Chang YC, Chang CS, Lin SF, Liu YC, Hsiao HH, et al. MALAT1 long non-coding RNA is overexpressed in multiple myeloma and may serve as a marker to predict disease progression. BMC Cancer. 2014;14:809.
Du Z, Sun T, Hacisuleyman E, Fei T, Wang X, Brown M, et al. Integrative analyses reveal a long noncoding RNA-mediated sponge regulatory network in prostate cancer. Nat Commun. 2016;7:10982.
Beck D, Thoms JAI, Palu C, Herold T, Shah A, Olivier J, et al. A four-gene LincRNA expression signature predicts risk in multiple cohorts of acute myeloid leukemia patients. Leukemia. 2018;32(2):263–272.
Mendell JT. Targeting a long noncoding RNA in breast cancer. N Engl J Med. 2016;374:2287–9.
Attal M, Lauwers-Cances V, Hulin C, Leleu X, Caillot D, Escoffre M, et al. Lenalidomide, bortezomib, and dexamethasone with transplantation for myeloma. N Engl J Med. 2017;376:1311–20.
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods. 2017;14:417–9.
Soneson C, Love MI, Robinson MD. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res. 2015;4:1521.
Robinson MD, Oshlack A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010;11:R25.
Law CW, Chen Y, Shi W, Smyth GK. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014;15:R29.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
Samur MK, Shah PK, Wang X, Minvielle S, Magrangeas F, Avet-Loiseau H, et al. The shaping and functional consequences of the dosage effect landscape in multiple myeloma. BMC Genom. 2013;14:672.
Kuiper R, Broyl A, de Knegt Y, van Vliet MH, van Beers EH, van der Holt B, et al. A gene expression signature for high-risk multiple myeloma. Leukemia. 2012;26:2406–13.
Munshi NC, Avet-Loiseau H, Rawstron AC, Owen RG, Child JA, Thakurta A, et al. Association of minimal residual disease with superior survival outcomes in patients with multiple myeloma: a meta-analysis. JAMA Oncol. 2017;3:28–35.
Shaughnessy JD Jr, Zhan F, Burington BE, Huang Y, Colla S, Hanamura I, et al. A validated gene expression model of high-risk multiple myeloma is defined by deregulated expression of genes mapping to chromosome 1. Blood. 2007;109:2276–84.
Mulligan G, Mitsiades C, Bryant B, Zhan F, Chng WJ, Roels S, et al. Gene expression profiling and correlation with outcome in clinical trials of the proteasome inhibitor bortezomib. Blood. 2007;109:3177–88.
Decaux O, Lode L, Magrangeas F, Charbonnel C, Gouraud W, Jezequel P, et al. Prediction of survival in multiple myeloma based on gene expression profiles reveals cell cycle and chromosomal instability signatures in high-risk patients and hyperdiploid signatures in low-risk patients: a study of the Intergroupe Francophone du Myelome. J Clin Oncol. 2008;26:4798–805.
Acknowledgements
NIH grants P01-155258 and P50-100707, Department of Veterans Affairs Merit Review Award I01 BX001584-01.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Samur, M.K., Minvielle, S., Gulla, A. et al. Long intergenic non-coding RNAs have an independent impact on survival in multiple myeloma. Leukemia 32, 2626–2635 (2018). https://doi.org/10.1038/s41375-018-0116-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0116-y
This article is cited by
-
LITTIP/Lgr6/HnRNPK complex regulates cementogenesis via Wnt signaling
International Journal of Oral Science (2023)
-
In-depth analysis of alternative splicing landscape in multiple myeloma and potential role of dysregulated splicing factors
Blood Cancer Journal (2022)
-
Functional dissection of inherited non-coding variation influencing multiple myeloma risk
Nature Communications (2022)
-
Integrative multi-omics identifies high risk multiple myeloma subgroup associated with significant DNA loss and dysregulated DNA repair and cell cycle pathways
BMC Medical Genomics (2021)
-
Dysregulated APOBEC3G causes DNA damage and promotes genomic instability in multiple myeloma
Blood Cancer Journal (2021)