Abstract
Background
African Americans have the highest pancreatic cancer incidence of any racial/ethnic group in the United States. The oral microbiome was associated with pancreatic cancer risk in a recent study, but no such studies have been conducted in African Americans. Poor oral health, which can be a cause or effect of microbial populations, was associated with an increased risk of pancreatic cancer in a single study of African Americans.
Methods
We prospectively investigated the oral microbiome in relation to pancreatic cancer risk among 122 African-American pancreatic cancer cases and 354 controls. DNA was extracted from oral wash samples for metagenomic shotgun sequencing. Alpha and beta diversity of the microbial profiles were calculated. Multivariable conditional logistic regression was used to estimate odds ratios (ORs) and 95% confidence intervals (CIs) for associations between microbes and pancreatic cancer risk.
Results
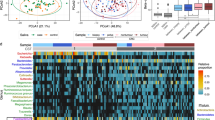
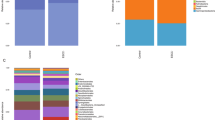
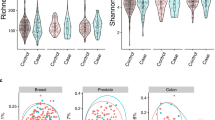
No associations were observed with alpha or beta diversity, and no individual microbial taxa were differentially abundant between cases and control, after accounting for multiple comparisons. Among never smokers, there were elevated ORs for known oral pathogens: Porphyromonas gingivalis (OR = 1.69, 95% CI: 0.80–3.56), Prevotella intermedia (OR = 1.40, 95% CI: 0.69–2.85), and Tannerella forsythia (OR = 1.36, 95% CI: 0.66–2.77).
Conclusions
Previously reported associations between oral taxa and pancreatic cancer were not present in this African-American population overall.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Data availability
The oral microbiome data produced in the current study are available via the database of Genotypes and Phenotypes (dpGaP, accession number: phs002454.v1.p1).
References
Surveillance, Epidemiology, and End Results (SEER) Program. SEER*Stat Database: Incidence—SEER 18 Regs Research Data, Nov 2018 Sub (2000–2015) <Katrina/Rita Population Adjustment>—Linked To County Attributes—Total U.S., 1969–2017 Counties, National Cancer Institute, DCCPS, Surveillance Research Program, released April 2019, based on the November 2018 submission. www.seer.cancer.gov.
Gerlovin H, Michaud DS, Cozier YC, Palmer JR. Oral health in relation to pancreatic cancer risk in African American women. Cancer Epidemiol Biomark Prev. 2019;28:675–9.
Michaud DS, Joshipura K, Giovannucci E, Fuchs CS. A prospective study of periodontal disease and pancreatic cancer in US male health professionals. J Natl Cancer Inst. 2007;99:171–5.
Hiraki A, Matsuo K, Suzuki T, Kawase T, Tajima K. Teeth loss and risk of cancer at 14 common sites in Japanese. Cancer Epidemiol Biomark Prev. 2008;17:1222–7.
Stolzenberg-Solomon RZ, Dodd KW, Blaser MJ, Virtamo J, Taylor PR, Albanes D. Tooth loss, pancreatic cancer, and Helicobacter pylori. Am J Clin Nutr. 2003;78:176–81.
Hujoel PP, Drangsholt M, Spiekerman C, Weiss NS. An exploration of the periodontitis-cancer association. Ann Epidemiol. 2003;13:312–6.
Arora M, Weuve J, Fall K, Pedersen NL, Mucci LA. An exploration of shared genetic risk factors between periodontal disease and cancers: a prospective co-twin study. Am J Epidemiol. 2010;171:253–9.
Ahn J, Segers S, Hayes RB. Periodontal disease, Porphyromonas gingivalis serum antibody levels and orodigestive cancer mortality. Carcinogenesis. 2012;33:1055–8.
Maisonneuve P, Amar S, Lowenfels AB. Periodontal disease, edentulism, and pancreatic cancer: a meta-analysis. Ann Oncol. 2017;28:985–95.
Fan X, Alekseyenko AV, Wu J, Peters BA, Jacobs EJ, Gapstur SM, et al. Human oral microbiome and prospective risk for pancreatic cancer: a population-based nested case-control study. Gut. 2018;67:120–7.
Como DH, Stein Duker LI, Polido JC, Cermak SA. The persistence of oral health disparities for African American children: a scoping review. Int J Environ Res Public Health. 2019;16:710. https://doi.org/10.3390/ijerph16050710.
Rosenberg L, Adams-Campbell L, Palmer JR. The Black Women’s Health Study: a follow-up study for causes and preventions of illness. J Am Med Women’s Assoc. 1995;50:56–8.
Signorello LB, Hargreaves MK, Blot WJ. The Southern Community Cohort Study: investigating health disparities. J Health Care Poor Underserved. 2010;21:26–37.
Signorello LB, Hargreaves MK, Steinwandel MD, Zheng W, Cai Q, Schlundt DG, et al. Southern community cohort study: establishing a cohort to investigate health disparities. J Natl Med Assoc. 2005;97:972–9.
Cozier YC, Palmer JR, Rosenberg L. Comparison of methods for collection of DNA samples by mail in the Black Women’s Health Study. Ann Epidemiol. 2004;14:117–22.
Garcia-Closas M, Egan KM, Abruzzo J, Newcomb PA, Titus-Ernstoff L, Franklin T, et al. Collection of genomic DNA from adults in epidemiological studies by buccal cytobrush and mouthwash. Cancer Epidemiol Biomark Prev. 2001;10:687–96.
Yang Y, Cai Q, Shu XO, Steinwandel MD, Blot WJ, Zheng W, et al. Prospective study of oral microbiome and colorectal cancer risk in low-income and African American populations. Int J Cancer. 2019;144:2381–9.
Yang Y, Zheng W, Cai Q, Shrubsole MJ, Pei Z, Brucker R, et al. Racial differences in the oral microbiome: data from low-income populations of African ancestry and European ancestry. mSystems. 2019;4:e00639–19. https://doi.org/10.1128/mSystems.00639-19.
Adams-Campbell LL, Dash C, Palmer JR, Wiedemeier MV, Russell CW, Rosenberg L, et al. Predictors of biospecimen donation in the Black Women’s Health Study. Cancer Causes Control. 2016;27:797–803.
Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, Blekhman R, et al. Human genetics shape the gut microbiome. Cell. 2014;159:789–99.
McIver LJ, Abu-Ali G, Franzosa EA, Schwager R, Morgan XC, Waldron L, et al. bioBakery: a meta’omic analysis environment. Bioinformatics. 2018;34:1235–7.
Segata N, Waldron L, Ballarini A, Narasimhan V, Jousson O, Huttenhower C. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat Methods. 2012;9:811–4.
Caspi R, Billington R, Keseler IM, Kothari A, Krummenacker M, Midford PE, et al. The MetaCyc database of metabolic pathways and enzymes - a 2019 update. Nucleic Acids Res. 2020;48:D445–53.
Sinha R, Abu-Ali G, Vogtmann E, Fodor AA, Ren B, Amir A, et al. Assessment of variation in microbial community amplicon sequencing by the Microbiome Quality Control (MBQC) project consortium. Nat Biotechnol. 2017;35:1077–86.
Salter SJ, Cox MJ, Turek EM, Calus ST, Cookson WO, Moffatt MF, et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 2014;12:87.
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, et al. Vegan: Community Ecology Package. R package version 2.5-6. 2019, https://cran.r-project.org/web/packages/vegan/index.html. Accessed October 15, 2020.
Hadley W. Ggplot2. New York, NY: Springer Science+Business Media, LLC; 2016.
Holt SC, Ebersole JL. Porphyromonas gingivalis, Treponema denticola, and Tannerella forsythia: the “red complex”, a prototype polybacterial pathogenic consortium in periodontitis. Periodontology 2000. 2005;38:72–122.
Aruni AW, Mishra A, Dou Y, Chioma O, Hamilton BN, Fletcher HM. Filifactor alocis—a new emerging periodontal pathogen. Microbes Infect. 2015;17:517–30.
Aykut B, Pushalkar S, Chen R, Li Q, Abengozar R, Kim JI, et al. The fungal mycobiome promotes pancreatic oncogenesis via activation of MBL. Nature. 2019;574:264–7.
Rothman KJ, Greenland S, Lash TL. Modern epidemiology. 3rd ed. Philadelphia: Wolters Kluwer Health/Lippincott Williams & Wilkins; 2008.
Buchowski MS, Schlundt DG, Hargreaves MK, Hankin JH, Signorello LB, Blot WJ. Development of a culturally sensitive food frequency questionnaire for use in the Southern Community Cohort Study. Cell Mol Biol. 2003;49:1295–304.
Kumanyika SK, Mauger D, Mitchell DC, Phillips B, Smiciklas-Wright H, Palmer JR. Relative validity of food frequency questionnaire nutrient estimates in the Black Women’s Health Study. Ann Epidemiol. 2003;13:111–8.
Wu J, Peters BA, Dominianni C, Zhang Y, Pei Z, Yang L, et al. Cigarette smoking and the oral microbiome in a large study of American adults. ISME J. 2016;10:2435–46.
Yu G, Phillips S, Gail MH, Goedert JJ, Humphrys MS, Ravel J, et al. The effect of cigarette smoking on the oral and nasal microbiota. Microbiome. 2017;5:3.
Huang C, Shi G. Smoking and microbiome in oral, airway, gut and some systemic diseases. J Transl Med. 2019;17:225.
McShane BB, Gal D, Gelman A, Robert C, Tackett JL. Abandon statistical significance. Am Statistician. 2019;73:235–45.
Benjamini Y, Hochberg Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J R Stat Soc Ser B. 1995;57:289–300.
Vogtmann E, Han Y, Caporaso JG, Bokulich N, Mohamadkhani A, Moayyedkazemi A, et al. Oral microbial community composition is associated with pancreatic cancer: a case-control study in Iran. Cancer Med. 2020;9:797–806.
Farrell JJ, Zhang L, Zhou H, Chia D, Elashoff D, Akin D, et al. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut. 2012;61:582–8.
Lu H, Ren Z, Li A, Li J, Xu S, Zhang H, et al. Tongue coating microbiome data distinguish patients with pancreatic head cancer from healthy controls. J Oral Microbiol. 2019;11:1563409.
Del Castillo E, Meier R, Chung M, Koestler DC, Chen T, Paster BJ, et al. The microbiomes of pancreatic and duodenum tissue overlap and are highly subject specific but differ between pancreatic cancer and noncancer subjects. Cancer Epidemiol Biomark Prev. 2019;28:370–83.
Half E, Keren N, Reshef L, Dorfman T, Lachter I, Kluger Y, et al. Fecal microbiome signatures of pancreatic cancer patients. Sci Rep. 2019;9:16801.
Michaud DS, Izard J, Wilhelm-Benartzi CS, You DH, Grote VA, Tjonneland A, et al. Plasma antibodies to oral bacteria and risk of pancreatic cancer in a large European prospective cohort study. Gut. 2013;62:1764–70.
Hanioka T, Morita M, Yamamoto T, Inagaki K, Wang PL, Ito H, et al. Smoking and periodontal microorganisms. Jpn Dent Sci Rev. 2019;55:88–94.
Scott DA, Singer DL. Suppression of overt gingival inflammation in tobacco smokers - clinical and mechanistic considerations. Int J Dent Hyg. 2004;2:104–10.
Palmer RM, Wilson RF, Hasan AS, Scott DA. Mechanisms of action of environmental factors–tobacco smoking. J Clin Periodontol. 2005;32:180–95.
Bagaitkar J, Williams LR, Renaud DE, Bemakanakere MR, Martin M, Scott DA, et al. Tobacco-induced alterations to Porphyromonas gingivalis-host interactions. Environ Microbiol. 2009;11:1242–53.
Bagaitkar J, Daep CA, Patel CK, Renaud DE, Demuth DR, Scott DA. Tobacco smoke augments Porphyromonas gingivalis-Streptococcus gordonii biofilm formation. PLoS ONE. 2011;6:e27386.
Gloor GB, Macklaim JM, Pawlowsky-Glahn V, Egozcue JJ. Microbiome datasets are compositional: and this is not optional. Front Microbiol. 2017;8:2224.
Paulson JN, Stine OC, Bravo HC, Pop M. Differential abundance analysis for microbial marker-gene surveys. Nat Methods. 2013;10:1200–2.
Quinn TP, Erb I, Richardson MF, Crowley TM. Understanding sequencing data as compositions: an outlook and review. Bioinformatics. 2018;34:2870–8.
Quinn TP, Crowley TM, Richardson MF. Benchmarking differential expression analysis tools for RNA-Seq: normalization-based vs. log-ratio transformation-based methods. BMC Bioinform. 2018;19:274.
Calle ML. Statistical analysis of metagenomics data. Genomics Inf. 2019;17:e6.
Hawinkel S, Mattiello F, Bijnens L, Thas O. A broken promise: microbiome differential abundance methods do not control the false discovery rate. Brief Bioinform. 2019;20:210–21.
Calgaro M, Romualdi C, Waldron L, Risso D, Vitulo N. Assessment of statistical methods from single cell, bulk RNA-seq, and metagenomics applied to microbiome data. Genome Biol. 2020;21:191.
Mallick H, Rahnavard A, McIver LJ, Ma S, Zhang Y, Nguyen LH, et al. Multivariable association discovery in population-scale meta-omics studies. bioRxiv:2021.2001.2020.427420 [Preprint] 2021. Available from: https://doi.org/10.1101/2021.01.20.427420.
Jovel J, Patterson J, Wang W, Hotte N, O’Keefe S, Mitchel T, et al. Characterization of the gut microbiome using 16S or shotgun metagenomics. Front Microbiol. 2016;7:459.
Franzosa EA, Hsu T, Sirota-Madi A, Shafquat A, Abu-Ali G, Morgan XC, et al. Sequencing and beyond: integrating molecular ‘omics’ for microbial community profiling. Nat Rev Microbiol. 2015;13:360–72.
Li S, Fuhler GM, Bn N, Jose T, Bruno MJ, Peppelenbosch MP, et al. Pancreatic cyst fluid harbors a unique microbiome. Microbiome. 2017;5:147.
Coker OO, Nakatsu G, Dai RZ, Wu WKK, Wong SH, Ng SC, et al. Enteric fungal microbiota dysbiosis and ecological alterations in colorectal cancer. Gut. 2019;68:654–62.
Klimesova K, Jiraskova Zakostelska Z, Tlaskalova-Hogenova H. Oral bacterial and fungal microbiome impacts colorectal carcinogenesis. Front Microbiol. 2018;9:774.
Swidsinski A, Schlien P, Pernthaler A, Gottschalk U, Barlehner E, Decker G, et al. Bacterial biofilm within diseased pancreatic and biliary tracts. Gut. 2005;54:388–95.
Pasolli E, Truong DT, Malik F, Waldron L, Segata N. Machine learning meta-analysis of large metagenomic datasets: tools and biological insights. PLoS Comput Biol. 2016;12:e1004977.
Lloyd-Price J, Arze C, Ananthakrishnan AN, Schirmer M, Avila-Pacheco J, Poon TW, et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature. 2019;569:655–62.
Crasta K, Daly CG, Mitchell D, Curtis B, Stewart D, Heitz-Mayfield LJ. Bacteraemia due to dental flossing. J Clin Periodontol. 2009;36:323–32.
Lockhart PB, Brennan MT, Sasser HC, Fox PC, Paster BJ, Bahrani-Mougeot FK. Bacteremia associated with toothbrushing and dental extraction. Circulation. 2008;117:3118–25.
Garlet GP. Destructive and protective roles of cytokines in periodontitis: a re-appraisal from host defense and tissue destruction viewpoints. J Dent Res. 2010;89:1349–63.
Cardoso EM, Reis C, Manzanares-Cespedes MC. Chronic periodontitis, inflammatory cytokines, and interrelationship with other chronic diseases. Postgrad Med. 2018;130:98–104.
Salaspuro M. Acetaldehyde as a common denominator and cumulative carcinogen in digestive tract cancers. Scand J Gastroenterol. 2009;44:912–25.
Kurkivuori J, Salaspuro V, Kaihovaara P, Kari K, Rautemaa R, Gronroos L, et al. Acetaldehyde production from ethanol by oral streptococci. Oral Oncol. 2007;43:181–6.
Shapiro KB, Hotchkiss JH, Roe DA. Quantitative relationship between oral nitrate-reducing activity and the endogenous formation of N-nitrosoamino acids in humans. Food Chem Toxicol. 1991;29:751–5.
Risch HA. Pancreatic cancer: Helicobacter pylori colonization, N-nitrosamine exposures, and ABO blood group. Mol Carcinogen. 2012;51:109–18.
Duell EJ. Epidemiology and potential mechanisms of tobacco smoking and heavy alcohol consumption in pancreatic cancer. Mol Carcinogen. 2012;51:40–52.
Keeley TS, Yang S, Lau E. The diverse contributions of fucose linkages in cancer. Cancers. 2019;11:1241, https://doi.org/10.3390/cancers11091241.
Rothman KJ. No adjustments are needed for multiple comparisons. Epidemiology. 1990;1:43–6.
Silva-Aycaguer LC, Suarez-Gil P, Fernandez-Somoano A. The null hypothesis significance test in health sciences research (1995–2006): statistical analysis and interpretation. BMC Med Res Methodol. 2010;10:44.
Mallick H, Rahnavard A, McIver L. Maaslin2: Maaslin2. R package version 1.3.0. 2020. http://huttenhower.sph.harvard.edu/maaslin. Accessed October 15, 2020.
Vogtmann E, Hua X, Zhou L, Wan Y, Suman S, Zhu B, et al. Temporal variability of oral microbiota over 10 months and the implications for future epidemiologic studies. Cancer Epidemiol Biomark Prev. 2018;27:594–600.
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R. Bacterial community variation in human body habitats across space and time. Science. 2009;326:1694–7.
David LA, Materna AC, Friedman J, Campos-Baptista MI, Blackburn MC, Perrotta A, et al. Host lifestyle affects human microbiota on daily timescales. Genome Biol. 2014;15:R89.
Stahringer SS, Clemente JC, Corley RP, Hewitt J, Knights D, Walters WA, et al. Nurture trumps nature in a longitudinal survey of salivary bacterial communities in twins from early adolescence to early adulthood. Genome Res. 2012;22:2146–52.
Human Microbiome Project, C. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207–14.
Acknowledgements
Pathology data were obtained from the following state cancer registries (AZ, CA, CO, CT, DE, DC, FL, GA, IL, IN, KY, LA, MD, MA, MI, NJ, NY, NC, OK, PA, SC, TN, TX, VA), and results reported do not necessarily represent their views. We thank participants and staff of the BWHS and SCCS for their contributions.
Funding
This work was supported by National Institutes of Health grants U01 CA164974, U01 CA187508 and R01 CA058420; the Karin Grunebaum Cancer Research Foundation and Boston University Peter Paul Career Development Professorship.
Author information
Authors and Affiliations
Contributions
Statistical analysis, interpretation of data, drafting of the paper and critical revision of the paper for important intellectual content: JLP. Statistical analysis, interpretation of data and critical revision of the paper for important intellectual content: JEW and HG. Study concept and design, analysis and interpretation of data, and critical revision of the paper for important intellectual content: DSM and CH. Interpretation of data and critical revision of the paper for important intellectual content: QC, BMW, EAR-N, JL, YY, and WEJ. Study concept and design, acquisition of data, analysis and interpretation of data and critical revision of the paper for important intellectual content: LBS and X-OS. Study concept and design, acquisition of data, analysis and interpretation of data, drafting of the paper and critical revision of the paper for important intellectual content: JRP. All authors approved the final draft submitted.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All study participants provided written informed consent. The Institutional Review Boards of Boston University (Boston, MA) and Vanderbilt University Medical Center and Meharry Medical College (Nashville, TN) approved the BWHS and SCCS, respectively, and reviewed the studies annually.
Competing interests
BMW has received research funding from Celgene and Eli Lily and also serves as a consultant for BioLineRx, Celgene and Grail. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Petrick, J.L., Wilkinson, J.E., Michaud, D.S. et al. The oral microbiome in relation to pancreatic cancer risk in African Americans. Br J Cancer 126, 287–296 (2022). https://doi.org/10.1038/s41416-021-01578-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01578-5
This article is cited by
-
Microbiome as a biomarker and therapeutic target in pancreatic cancer
BMC Microbiology (2024)
-
Connecting the Human Microbiome and Pancreatic Cancer
Cancer and Metastasis Reviews (2022)