Abstract
COVID-19 can lead to life-threatening respiratory failure, with increased inflammatory mediators and viral load. Here, we perform single-cell RNA-sequencing to establish a high-resolution map of blood antigen-presenting cells (APCs) in 15 patients with moderate or severe COVID-19 pneumonia, at day 1 and day 4 post admission to intensive care unit or pulmonology department, as well as in 4 healthy donors. We generated a unique dataset of 81,643 APCs, including monocytes and rare dendritic cell (DC) subsets. We uncovered multi-process defects in antiviral immune defence in specific APCs from patients with severe disease: (1) increased pro-apoptotic pathways in plasmacytoid DCs (pDCs, key effectors of antiviral immunity), (2) a decrease of the innate sensors TLR9 and DHX36 in pDCs and CLEC9a+ DCs, respectively, (3) downregulation of antiviral interferon-stimulated genes in monocyte subsets and (4) a decrease of major histocompatibility complex (MHC) class II-related genes and MHC class II transactivator activity in cDC1c+ DCs, suggesting viral inhibition of antigen presentation. These novel mechanisms may explain patient aggravation and suggest strategies to restore the defective immune defence.
Similar content being viewed by others
Main
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) infection is at the origin of coronavirus disease 2019 (COVID-19), characterized by a first phase of benign flu-like symptoms with an efficient control of the infection in most cases. In a second phase, disease aggravation may lead to acute respiratory failure, sepsis and death1,2,3,4,5,6. This is due to a multiplicity of factors: (1) an exacerbated inflammatory reaction, with systemic and organ-specific manifestations, (2) persistent viral load and (3) defective antiviral defence pathways1,2,3,4,5,6,7. Identifying the underlying cellular and molecular mechanisms is of paramount importance to understand COVID-19 physiopathology and guide the development of appropriate therapies.
Studies have characterized the systemic inflammatory response, revealing an excess production of inflammatory cytokines such as interleukin-6 (IL-6) and IL-1, tumour necrosis factor-α (TNF-α) and interferon-γ (IFN-γ)2,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22, suggesting new therapeutic targets. The endothelium may also contribute to the overt inflammatory reaction through the production of soluble mediators23,24. Anti-IL-6 compounds have given promising results in severe COVID-1925,26,27. However, the cellular mechanisms underlying the excessive inflammatory response remain mostly unknown.
Another unresolved question relates to the inefficiency of the innate and adaptive immune system to control the infection in patients with severe COVID-19. It has been suggested that production of IFN-α, a major antiviral cytokine, is decreased in these patients compared to those with moderate disease6,9,21,28,29,30. However, a recent study argued that increased IFN-α production might contribute to the pathogenic inflammatory response17. Other antiviral mechanisms and their cellular source remain to be studied.
Dendritic cells (DCs) form a family of innate antigen (Ag)-presenting cells (APCs) that contribute to the control of pathogens and subsequent presentation of pathogen-specific Ag to T cells31. Their study is challenging for three main reasons: (1) they are found in very low numbers in the circulation and in tissue, (2) they lack specific lineage-defining markers and (3) they include an ever-increasing number of subsets31,32. All DC subsets may potentially and variably contribute to modulating the inflammatory response following viral sensing, producing antiviral effector molecules and priming an Ag-specific adaptive immune response33. Plasmacytoid pre-DCs (pDCs) are a particular subset specialized in antiviral immunity through the production of large amounts of type I IFN34. Despite their central role in antiviral defence, the contribution of DCs to severe COVID-19 pathogenesis is not yet known.
In this paper we perform a high-resolution single-cell RNA-sequencing (scRNAseq) analysis of all APC subsets from fresh peripheral blood of patients with COVID-19. A pre-enrichment step enables the characterization of even rare DC subsets that were not captured in previous peripheral blood mononuclear cell (PBMC) scRNAseq studies12,17,35. We reveal previously unrecognized multi-process defects in patients with severe COVID-1922,36,37.
Results
APC subset distribution in patients with COVID-19
To characterize the molecular profile of circulating APCs, we performed scRNAseq on freshly sampled APC-enriched PBMCs from five patients with moderate COVID-19 (non-mechanically ventilated, oxygen supply <10 l min−1) and ten patients with severe COVID-19 (mechanically ventilated or oxygen supply ≥10 l min−1), at day 1 and day 4 following hospital and/or intensive care unit (ICU) admission, as well as four elderly healthy controls (HC) (Fig. 1a and Supplementary Tables 1 and 2). To obtain single-cell suspensions and minimize DC–DC and DC–T cell clusters and clumps, EDTA-containing medium was used for the enrichment steps in the first set of samples, which we further define as the ‘discovery set’. This set is composed of a total of 12 samples from two HCs, three patients with moderate COVID-19 and four patients with severe COVID-19 from both day 1 and day 4 time points (results are presented in the figures and Extended Data Figs. 1 and 2). However, EDTA is known to decrease reverse transcription (RT) efficiency through RT deactivation and ion chelation, resulting in reduced amounts of complementary DNA (cDNA) during amplification. We therefore validated the main results derived from the discovery set by using RPMI (EDTA-free) medium for the enrichment steps in a second set of samples, including a total of 15 samples (defined as the ‘validation set’) from two HCs, two patients with moderate COVID-19 and six with severe COVID-19 (Extended Data Figs. 3–7).
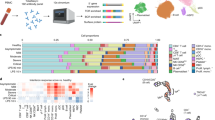
a, Schematic of the experimental workflow. APCs were enriched from fresh PBMCs of healthy donors and patients with COVID-19 with either moderate or severe clinical symptoms at both day 1 and day 4 post hospital admission. The total APCs were sequenced using the 10X Genomics facility. b,c, Cellular maps of APC subsets (n = 42,784 cells) from the discovery set at single-cell resolution level displayed on UMAP dimension reduction based either on identified cell types (b) and severity (c). Proportions of the APC subtypes are displayed on the doughnut plot. d, UMAP plot of detected APC populations split by severity group (healthy controls and patients with moderate and severe COVID-19). The discovery set comprises a total of 12 samples (n = 2 controls, n = 4 moderate and n = 6 severe samples) collected from a total of seven patients and two healthy donors.
For each fresh sample, 25,000 cells (~20,000 monocytes and 5,000 total DCs) were loaded onto the 10X lane (10X Genomics technology) (Fig. 1a). As expected, more cells per sample were effectively sequenced in the EDTA (discovery set) than the RPMI (validation set) dataset (mean: 3,360 versus 2,528 cells, respectively) (Extended Data Fig. 3a), confirming that EDTA optimizes single-cell suspension efficiency for rare DC types. This retrospectively justified the importance of using two complementary experimental protocols, split into two independent datasets, to avoid biasing the results. All main findings were validated in both datasets, indicating the reproducibility and robustness to experimental procedures. Altogether, we analysed a total of 81,643 APCs, split into 42,784 cells in the discovery set and 38,859 cells in the validation set. The two sets were analysed separately after sample integration using Harmony38. Graph-based clustering (SNN-based), community detection and nonlinear dimension reduction, using uniform manifold approximation and projection (UMAP), were independently applied to both sets for cell cluster visualization. Manual annotation of the cell clusters using canonical gene signature markers for each APC subset established a comprehensive map of APCs in HCs and patients with COVID-19 in both sets (Fig. 1b,c and Extended Data Fig. 3b,c). In our discovery set, among the 42,784 APCs, we recovered six subsets: 22,690 CD14+ monocytes; 866 CD16+ monocytes; 13,252 CD1c+ DCs; 1,754 CLEC9a+ DCs; 3,538 pDCs; 684 Axl+Siglec6+ AS-DCs (Extended Data Fig. 3a). The validation set included 29,409 CD14+ and 1,021 CD16+ monocytes, 5,754 CD1c+, 197 CLEC9a+ DCs, 1,602 pDCs and 876 AS-DCs (Extended Data Fig. 3a). In both sets, APC populations were captured across all the collected samples (Supplementary Table 3).
The accurate identification of all six APC populations was confirmed by the expression of canonical markers defining each subset (Extended Data Fig. 3d). All DC populations expressed higher levels of human leukocyte antigen HLA-DR and CD86 compared to monocytes (Extended Data Fig. 3d). None of the cells expressed CD19 (B-cell marker), GNLY (natural killed (NK) marker) or CD3E (T-cell marker), validating the pure APC populations. CD14+ monocytes expressed lineage-defining CD14, whereas CD16+ monocytes expressed FCGR3A. AXL expression distinguished AS-DCs from pDCs, whereas CD1c and CLEC9a characterized the respective cDC subsets39,40. In both sets (discovery and validation), UMAP embeddings coloured by severity revealed the heterogeneity of APC distribution between the three groups (Fig. 1c). This was confirmed by splitting the UMAP embeddings per severity (Fig. 1d). Overall, our enrichment strategy allowed the efficient identification of all APC populations including the rare pDCs, AS-DCs and CLEC9A+ DCs, enabling further molecular and phenotypic characterization.
Inflammation-related pathways are hallmarks of COVID-19 APCs
We performed differential expression and pathway enrichment analyses among APC severity groups, revealing 368 differentially expressed genes (DEGs) among the three groups (absolute fold change > 1.4). Among them, 101 genes were upregulated in HCs (as compared to patients with moderate and severe COVID-19), 109 in patients with moderate COVID-19 and 134 in patients with severe COVID-19 as compared to the two other groups, respectively (Fig. 2a). The top 50 DEGs upregulated in severe APCs as compared to HCs and patients with moderate COVID-19 included pro-inflammatory molecules (IL1B, CXCR4), surface markers (CD36, CD83, AREG, ITGAM), enzymes (CTSD, CTSB) and secreted molecules (RETN, EREG, ANXA2) (Fig. 2b). Next, we sought to identify enriched pathways discriminating each severity group from HCs. We found enriched IFN-γ and IFN-α response pathways in APCs from patients with moderate COVID-19, whereas hypoxia and TNF-α signalling were enriched in patients with severe COVID-19 (Fig. 2c).
a, Barplot of the number of differentially expressed genes (DEGs) for each severity group (healthy versus patients with moderate and severe COVID-19; moderate versus healthy and severe; severe versus healthy and moderate). Upregulated (log fold change (FC) > 0.25) genes are shown in black, downregulated (log FC < −0.25) genes are shown in grey. b, Heatmap representation of the top upregulated genes in severe APCs, as compared to moderate and healthy groups. The z-score values of average expression levels of cells per severity group are colour-coded. c,d, Comparative analysis of enriched pathways from the upregulated genes in moderate or severe APCs as compared to healthy cells (c), as well as pairwise comparison of upregulated genes in moderate compared to severe (shown in pink) and upregulated genes in severe compared to moderate (shown in yellow) (d). Horizontal axes display the adjusted P values (−log10). e, Representation of ranked genes in descending order according to their absolute log FC, upregulated in moderate as compared to severe (red plot) and upregulated in severe as compared to moderate (blue plot). Top genes, with an absolute value of log FC above 0.5, are shown. In a–e, comparative analyses were performed on the discovery set (n = 42,784 cells), composed of n = 2 HC, n = 4 moderate and n = 6 severe samples. The two-sided Wilcoxon rank-sum test was used for comparison, P values were adjusted to multiple testing using ‘Bonferroni’ correction, and only genes with adjusted P < 0.05 were considered.
We next compared the enriched pathways upregulated in severe versus moderate COVID-19 and in moderate versus severe, respectively. We found that IFN-γ and IFN-α pathways could be used to discriminate moderate from severe APCs at the global level (Fig. 2d).
To allow for an accurate comparison between the two transcriptional signatures, we ranked the DEGs of the pairwise comparison according to decreasing fold change. Severe APCs significantly upregulated AREG (amphiregulin), IL1R2 (IL-1 receptor), NRGN (calmodulin binding protein) and pro-inflammatory molecules (S100A12) (Fig. 2e). However, moderate APCs overexpressed interferon-stimulated genes (ISGs; IFITM2, ISG15 and IFI27) and HLAII molecules (HLA-DRB5 and HLA-DQA2), suggesting decreased Ag presentation and antiviral programs in severe as compared to moderate APCs (Fig. 2e). Similar observations were recovered from our validation set (Extended Data Fig. 4a–d). Additional upregulated genes in severe as compared to moderate APCs were found in the validation set, including CXCL8, NAMPT and G0S2 (Extended Data Fig. 4e).
Defective IFN responses in COVID-19 APCs
Increases in inflammatory cytokines have been reported in COVID-19. We addressed the global contribution of APCs to the expression of inflammatory cytokines and their receptors. As compared to APCs derived from HCs, IL1B, CXCL2, CXCL8 and CCL3 were significantly increased, whereas IL18 was decreased in both severity groups (Fig. 3a and Extended Data Fig. 1a). TGFB1 and IL10RA expression decreased in severe, but not in moderate subsets, as compared to HCs (Fig. 3a and Extended Data Fig. 1a), whereas IL6 was not detected in our discovery set (Extended Data Fig. 1a). Despite the low expression levels of most cytokines, we explored downstream biological pathways associated with inflammatory cytokine signalling (mainly IL1B, IL6 and TNF-α). In comparison to APCs from HCs, both moderate and severe APCs showed higher score levels for hallmark inflammatory pathways, including ‘IL6_JAK_STAT3’, ‘TGF-β’, ‘P53’, ‘TNFa_SIGNALLING_VIA_NFKB’ and ‘KRAS_SIGNALLING’ (Fig. 3b). The IFN family of cytokines is one of the most important for innate and adaptive antiviral responses. We showed the expression levels of IFNL1, IFNL1R, IFNAR1, IFNAR2, IFNA1, IFNGR1 and IFNGR2 and further explored their distribution in the three severity groups via a scaled heatmap (Fig. 3c). Both IFN receptor types (IFNAR1, IFNAR2, IFNGR1 and IFNGR2) were broadly expressed in the APC subsets, whereas detection of IFNL1 and IFNLR1 was patchy in our discovery dataset (Extended Data Fig. 3b). The heatmap representation indicated that severe APCs expressed lower levels of IFN molecules, suggesting a potential defect in IFN signalling (Fig. 3c). To further validate this hypothesis, we investigated the expression levels of ISGs. We observed higher expression levels of ISGs (MX2, ISG15, IRF7, BST2, IFITM2 and ADAR) in moderate APCs, but lower levels in severe APCs, supporting the hypothesis of defective antiviral programs contributing to the severity of COVID-19 (Fig. 3c). We further stratified a more exhaustive ISG signature according to their respective functions related to ‘antiviral’ and ‘regulators of IFN signalling’. Moderate APCs displayed higher levels of these two ISG families compared to both severe and HC groups (Fig. 3d,e). These results suggest a global perturbation of IFN downstream functions in severe COVID-19 APCs.
a, Violin plot representation of cytokine (IL1B, TGFB1 and IL-18) receptor (IL6R, TNFRSF1A) and chemokine (CXCL8) gene expression levels detected by scRNAseq and comparison between severity groups. Each dot represents a cell and horizontal lines display the mean expression value. b, Dot plot of enrichment scores of pathways downstream of IL-1B, IL-6 and TGFB1 inflammatory cytokines; score levels are colour-coded, and the percentage of cells expressing the pathway score is size-coded. c, Heatmap representation of expression levels of IFN genes (ligands and receptors) and ISG expression levels in healthy, moderate and severe APCs. Expression levels are colour-coded. d, Dot plots of regulators of IFN signalling and antiviral ISG genes in HCs and patients with moderate and severe COVID-19. Expression levels are colour-coded, and the percentage of cells expressing the respective gene is size-coded. e, Violin plot representation of antiviral ISGs among the severity groups; the small horizontal line indicates the mean expression value for each plotted gene expression. In a and e, the violin plots were designed using the total APC subsets from the discovery set (n = 42,784 cells), composed of n = 2 HC, n = 4 moderate and n = 6 severe samples. Comparative analysis was performed using the two-sided Wilcoxon rank-sum test; P values were adjusted to multiple testings using ‘Bonferroni’ correction. Asterisks above severe indicate P values for severe versus control; asterisks above moderate indicate significance of moderate versus control. *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant.
Multi-process effector defects in severe COVID-19 pDCs
After having analysed COVID-19 APCs at the global level, we sought to decipher alterations occurring in specific APC subsets. To depict the alterations occurring in pDC subsets, we isolated and sub-clustered pDCs (Fig. 4a) and performed pairwise differential expression among the three severity groups. Pathway enrichment analysis using MsigDB hallmark signatures was conducted on the upregulated genes in each subset. Compared to both moderate and HC pDCs, severe pDCs were enriched for the ‘TNFa_SIGNALING’, ‘IL2_STAT5’ and ‘HYPOXIA’ signalling pathways. In parallel, compared to pDCs from patients with moderate COVID-19, pDCs from patients with severe COVID-19 were enriched in the ‘IL6_JAK_STAT3’, ‘P53’ and ‘MTORC’ signalling pathways (Fig. 4b). When comparing pDCs between patients with moderate and severe COVID-19, the most notably enriched pathways were related to IFN signalling (IFNG and IFNA response), along with MYC targets signalling pathways (Fig. 4b). We asked whether apoptosis and pro-inflammatory signalling signatures would be associated with changes in pDC innate sensing receptors, including TLR9, DHX36, IFNAR1 and IFNAR2. We imputed the expression values to recover the signal from dropped-out features using Nebulosa (https://github.com/powellgenomicslab/Nebulosa), and plotted the density estimation values on UMAP embeddings (Fig. 4c). We observed zero-value density levels for TLR9, along with decreased density levels for DHX36, IFNAR1 and IFNAR2, in pDCs from patients with severe COVID-19 (Fig. 4c). To explore whether these modulations may impact pDC functions, we defined four original functional modules using a literature-driven manual curation: ‘immune cell attraction’ (hereafter ‘attraction’) (18 genes), ‘innate sensing’ (12 genes), ‘antiviral effector molecules’ (23 genes) and ‘cytotoxicity’ (12 genes) (Fig. 4d). Each of these modules was crossed with the pDC expression matrix, and detected genes were depicted for each patient group (Fig. 4d). No major differences between groups were detected within the ‘attraction’ module. On the contrary, many genes in the ‘innate sensing’, ‘antiviral effector molecules’ and ‘cytotoxicity’ modules were detected in the three groups, and followed the same pattern: baseline in HCs, increased in patients with moderate COVID-19 and decreased in patients with severe COVID-19 (Fig. 4d,e and Extended Data Fig. 5). This was particularly striking for the viral sensors TLR7, DHX9 and DHX36, the cytotoxic molecule TNFSF10 and the antiviral effector IRF7. These results were supported by the downregulation of antiviral ISGs and innate sensors in pDCs from patients with severe COVID-19, including BST2 and PYCARD (Fig. 4e), in both experimental datasets (Extended Data Fig. 5).
a, pDCs displayed on UMAP (n = 3,538 cells) coloured by severity group of origin. b, Violin plot distribution of enrichment scores for enriched hallmark pathways from upregulated genes from pairwise comparison between the three severity groups; severity groups are colour-coded, each dot represents a cell and the horizontal line displays the mean value of the enrichment score of each given pathway c, UMAP representation of density scores corresponding to IFN-a receptors (IFNAR1 and IFNAR2) and pDC sensors (TLR9 and DHX36). Density levels were computed using Nebulosa and are colour-coded. d, Dot plot of in-house constructed pDC-related functional modules (cytotoxicity, antiviral effector molecules, innate sensing and attraction) and comparison among severity groups. Expression levels are colour-coded, and the percentage of cells expressing the respective gene is size-coded. e, Violin plot representation of genes involved in pDC defined biological functions between severity groups; each dot represents a cell, and horizontal lines display the mean expression value. In b and e, the violin plots were designed using the total pDC subsets from the discovery set, including n = 2 HC, n = 4 moderate and n = 6 severe samples. Comparative analysis was performed using the two-sided Wilcoxon rank-sum test, P values were adjusted to multiple testings using ‘Bonferroni’ correction. Asterisks above severe indicate P values for severe versus control; asterisks above moderate indicate significance of moderate versus control. *P < 0.05, **P < 0.01, ***P < 0.001.
Coordinated transcriptional adaptation in monocyte subsets
Monocytes have been implicated in the physiopathology of severe sepsis and COVID-19. We performed dimensionality reduction through independent component analysis (ICA) and highlighted cells according to their severity group. We observed that IC1 clearly separated moderate from severe and HC CD14+ monocytes, whereas IC2 distinguished HC from COVID-19 CD14+ monocytes (Fig. 5a). The top 50 genes contributing to either IC1 or IC2 revealed distinct transcriptional signatures for the CD14+ monocyte subsets identified in each severity group: the severe subset expressed higher levels of complement (C1GC and C1GB), B7 family (VSIG4) and CD163, which may function as an innate immune sensor and inducer of local inflammation. The moderate monocyte subset expressed increased levels of antiviral ISGs (IFITM1, IFITM3, IFI27, MZB1 and IFI6) and the HLA-II gene (HLA-DRB5), suggesting an efficient antiviral program (Fig. 5b). Compared to HCs, several transcription factors (TFs) were downregulated in both moderate and severe groups, including the AP-1 superfamily (FOS, JUNB and ZFP36) and DUSP1, involved in MAPK dephosphorylation (Fig. 5b). Pathway enrichment analysis on the top 50 genes contributing to IC1 and IC2 identified key pathways that segregated COVID-19 CD14+ monocytes from HCs (Fig. 5c). The ‘complement’, ‘TNF-α’, ‘KRAS’ and ‘hypoxia’ signalling pathways were upregulated in COVID-19 monocytes, whereas ‘IFN-α’ and ‘IFN-γ’ response signalling were decreased in the severe subset, as compared to the HC and moderate subsets (Fig. 5c). To estimate antiviral effector functions, we used our manually curated gene functional module across patient groups (Fig. 5d). We observed a decrease of almost all antiviral effector molecules in patients with severe COVID-19, as compared to either HCs or patients with moderate COVID-19, in both experimental datasets (Fig. 5d and Extended Data Fig. 6). In parallel, we subclustered CD16+ monocytes and reduced the data dimension using UMAP projection to depict the corresponding clusters for each severity group (Fig. 5e). Differential expression between the three severity groups of this subset indicated similar trends as described in CD14+ monocytes (Fig. 5b,f). This included overexpression of ‘complement’-related genes (C1QA, C1QB and C1GC) by the severe subset, upregulation of antiviral ISGs (ISG15, IFI6 and IFI44L) in the moderate subset, as compared to the HC subset (Fig. 5f). Overall, these disease-associated changes in CD16+ paralleled those observed in CD14+ monocytes, suggesting common adaptation mechanisms.
a, ICA representation of CD14+ monocytes derived from all severity groups (n = 22,690 cells). IC1 and IC2 components allowed the separation of CD14+ monocytes according to severity. b, Heatmap representation of the top 50 unique genes contributing to either IC1 or IC2 in the CD14+ monocytes subset; z-scores of average expression levels are colour-coded. c, Violin plot distribution of enrichment score values of enriched pathways in the top 50 genes contributing to either IC1 or IC2; severity groups are colour-coded, each dot represents a cell and the horizontal line displays the mean value of the enrichment score of each given pathway. The violin plots were designed using the total CD14+ monocyte subsets from the discovery set, obtained from n = 2 HC, n = 4 moderate and n = 6 severe samples. Comparative analysis was performed using the two-sided Wilcoxon rank-sum test. P values were adjusted to multiple testings using ‘Bonferroni’ correction. Asterisks above severe indicate P values for severe versus control; asterisks above moderate indicate significance of moderate versus control. *P < 0.05, **P < 0.01, ***P < 0.001. d, Dot plot of in-house constructed antiviral effector molecule modules across CD14+ monocytes and severity groups. Expression levels are colour-coded, and the percentage of cells expressing the respective gene is size-coded. e, UMAP representation of CD16+ monocytes (n = 866 cells) labelled according to severity group. f, Heatmap representation of top 50 DEGs among the three groups from the CD16+ monocytes subset; z-scores of average expression levels are colour-coded.
CLEC9A+ DC- and AS-DC-specific transcriptional alterations
Thanks to our APC enrichment protocol, we could recover rare CLEC9a+ DC and AS-DC subsets. Differential expression of AS-DC severity groups revealed significant upregulated genes in severe AS-DCs (SEPT7 and AREG), compared to the moderate and HC subsets. We could also observe a significant downregulation of the HLA-DQA2 gene and antiviral IFI27 gene in severe, compared to moderate AS-DCs (Fig. 6a). In the search for upstream regulatory mechanisms, we inferred TF activity using the Dorothea algorithm41 and scored the activity of each regulon using the Viper inference tool42. This identified a large number of highly variant TF activity scores (Fig. 6b). In moderate AS-DCs, we observed a higher activity scored for IRF1, IRF9 and STAT2, reported to be involved in the ISG transcription cycle (Fig. 6b). In AS-DCs from patients with severe COVID-19, we found increased TF activities for RELA, NFKB1, STAT5 and STAT3, indicative of a higher activation of NFKB/STAT signalling, potentially induced by the pro-inflammatory cytokines described in the ‘APC subset distribution in patients with COVID-19’ section, along with hypoxia activation, indicated by a higher activity of HIF1A (Fig. 6b).
a, Violin plot representation of the top upregulated (AREG, SEPT7)/downregulated (IFI27, HLA-DQA2) genes in severe as compared to moderate AS-DCs. The violin plots were designed using the total AS-DC subsets from the discovery set (n = 684 cells), obtained from n = 2 HC, n = 4 moderate and n = 6 severe samples. Comparative analysis was performed using the two-sided Wilcoxon rank-sum test; P values were adjusted to multiple testings using ‘Bonferroni’ correction. Asterisks above severe indicate P values for severe versus control; asterisks above moderate indicate significance of moderate versus control. *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. b, Heatmap of top 50 highly variable TF activities among the three severity groups; the z-scores of TF activities are colour-coded. c, Heatmap representation of top 50 DEGs among the three severity groups isolated from the CLEC9a+ DC subset (a total of 1,754 cells); z-scores of expression level values are colour-coded; d, Violin plot distribution of IFNGR1 and HLA-DQA2 genes among the three CLEC9a+ DC severity groups.
DEGs among the CLEC9a+ DC subclusters included specific transcriptional signatures segregating patients with moderate and severe COVID-19 from HCs (Fig. 6c). We remarkably observed a downregulation of HLA-II genes, including HLA-DQB1 and HLA-DPB1, in severe as compared to HCs, along with a significant upregulation of a larger subset of ISGs, including IRF1, IFI44L, IFI6, IFI27, IFITM2, IFITM3, IFI44L, ISG15 and ISG20, in moderate as compared to both HC and severe subsets (Fig. 6c). Expression values representation indicated a significant increase of AREG and SEPT7 genes, which were also upregulated by severe AS-DCs (Fig. 6a,d). Most importantly, we noted a significant decrease of the IFNGR1 CLEC9a+ DC subset in patients with moderate and severe COVID-19 as compared to HCs (Fig. 6d), supporting a defective antiviral program.
Downregulation of MHC-II and CIITA activity in CD1C+ DCs
We next focused on disease-induced alterations in CD1c+ DCs. We first explored the gene expression levels of MHC-II-related genes (Fig. 7a). We noted a global decrease of HLA-II genes (mainly HLA-DQA2 and HLA-DRB5) in patients with severe COVID-19 (Fig. 7a). Similar findings were reported for the validation set (Extended Data Fig. 7). We then grouped the expression values of these MHC-II genes (HLA-DRB1, HLA-DMA, HLA-DQA2, HLA-DRB5, HLA-DPB1, HLA-DQB1 and HLA-DMB), constructed a signature that we named the ‘HLAII’ module, and scored CD1c+ DCs using the ‘AddModuleScore’ Seurat function. The ‘HLAII’ signature was significantly reduced in severe as compared to HC CD1c+ DCs (Fig. 7b). This was associated with decreased expression of upstream MHC-II regulators (including RFX5, RFXANK and CIITA) in the severe group (Fig. 7b). Comparison of the scaled values revealed a reduction of IRF1 and RFX5 TF activities, mainly described to be involved in MHC-II gene synthesis, whereas C/EBP family member (CEBPB and CEBPD) TF activities, known to be involved in myeloid fate differentiation, were increased in severe subsets (Fig. 7c). We also noted higher TF activities for RELA (NFKB superfamily) and the AP-1 family in patients with severe COVID-19, including FOSL1, FOSL2 and JUN, which regulate a large range of cellular processes, including cell survival, death and proliferation (Fig. 7c).
a, Dot plot distribution of HLA-II-related genes at the patient level within the CD1c+ DC subset; expression levels are colour-coded, and the percentage of cells expressing the respective gene is size-coded. b, Violin plot distribution of HLA-II and the upstream regulatorsʼ (HLAII_Regulators) module scores among the three severity groups within the CD1c+ DC subset; severity groups are colour-coded, each dot represents a cell and the horizontal line displays the mean value of the enrichment score of each given pathway. The violin plots were designed using the total CD1c+ DC subsets from the discovery set obtained from n = 2 HC, n = 4 moderate and n = 6 severe samples. Comparative analysis was performed using the two-sided Wilcoxon rank-sum test. P values were adjusted to multiple testings using ‘Bonferroni’ correction. Asterisks above severe indicate P values for severe versus control; asterisks above moderate indicate significance of moderate versus control. *P < 0.05, **P < 0.01, ***P < 0.001; NS, not significant. c, Heatmap representation of top 50 highly variable TF activities between CD1c+ DC severity groups; the z-score of activity scores is colour-coded. d, Pseudotime inference tree on UMAP embeddings (left) of the CD1c+ DC subset using Monocle3; pseudotime values are colour-coded (right). e, UMAP representation of density scores for the top genes contributing to the pseudotime tree initially inferred in d. Density scores were computed using Nebulosa and are colour-coded. All statistical tests displayed in this figure were performed using the discovery set, comprising a total of 12 samples (n = 2 controls, n = 4 moderate and n = 6 samples) collected from seven patients and two healthy donors.
To further decipher the transcriptional changes occurring in DCs when transitioning from healthy to moderate and severe conditions, we conducted pseudo-temporal inference using Monocle3, using the UMAP embedding two-dimensional space of the DC subsets (Fig. 7d). The pseudotime tree revealed a continuous trajectory from healthy to moderate, and a marked transition to the severe subsets. This trajectory was correlated to pseudotime values (Fig. 7d, right). To recover the genes contributing to this transition tree, we conducted a graph-based test to assess the most significant genes. The top genes were associated with Ag presentation, including B2M and HLA-DPA1, along with genes related to ISG expression (GABARAP and IFITM3; Fig. 7e).
Given that MHC-II genes are involved in the DC–T cell interaction, we hypothesized that a more global dysfunction of DC–T cell communication may occur in COVID-19 APCs. To test this hypothesis, we applied our cell communication inference computational framework ICELLNET43. Using our ‘reference partner cell’ methodology, we inferred potential communication between each of the APC subsets and CD4+ T cells in each of the disease groups (Fig. 8a). Cell connectivity networks revealed a global decrease in APC–T cell communication in patients with severe, as compared to moderate COVID-19 and HCs, predominantly in CD1c+ DCs, CLEC9a+ DCs and CD14+ monocytes (Fig. 8a,b). We then explored the various molecular families that may explain this decrease. This revealed a dominant contribution of immune checkpoint molecules and cytokines for CD1c+ DC–T cell communication (Fig. 8b), in particular decreased JAG-NOTCH, CD80-CD28 and CD48-CD2 interactions (Fig. 8c). Cytokines were mostly underlying the decrease in CD14+ monocytes–T cell communication (Fig. 8b). As expected, signalling through HLA-II-related genes (HLA-II/LAG3 pairs) was significantly decreased in moderate and severe subsets as compared to HCs. Among the cytokines, IL10-, CCL5- and TGFb-mediated interactions were predominantly damped in patients with severe COVID-19, which may contribute to immunopathology through excessive Th1 responses (Fig. 8c).
a, Connectivity maps describing outward communication from APCs from the single-cell dataset at day 1 to T lymphocytes (n = 39), according to patient severity (healthy, moderate and severe). The T lymphocyte transcriptomic profiles are from the Human Primary Cell Atlas, included in the ICELLNET R package. For APCs, average cluster gene expression profiles were considered. Only DC–T cell interactions are taken into account to compute the communication score (manually curated, n = 144 interactions). b, Barplot of each communication score with contribution by families of communication molecules for outward communication from APCs to T lymphocytes. c, Focus on CD1c+ DC outward communication to T lymphocytes, representing specific individual interaction scores that differ by at least 10 between patients with moderate and severe COVID-19 (cutoff chosen for the purpose of clarity).
Persistent defects in severe COVID-19 APCs across samples
We also asked whether functional pathway alterations observed across APC subtypes were sustained over time. In parallel, we wanted to ensure that our main findings were not driven by a single patient and/or time point. We compared scRNAseq datasets generated at day 1 versus day 4 post hospital admission for each patient, in all severity groups. We used a focused approach, by selecting genes involved in previously identified altered functions, in APCs from patients with severe COVID-19, and systematically compared day 1 and day 4 expression levels. Most of the day 1 defects were sustained at day 4, in particular the low score of the HLA-II module in CD1c+ DCs (Extended Data Fig. 2a) together with the decreased expression of the antiviral effector molecules in CD14+ monocytes (Extended Data Fig. 2b) and increased ‘apoptosis p53 pathway’ in pDCs (Extended Data Fig. 2c). At a patient level, we observed that HC samples displayed similar score (or expression) levels for these three biological processes, whereas both moderate and severe samples displayed slight differences due to inter-individual heterogeneity. Overall, we confirmed that our findings were not associated with either a specific time point or a dominant single patient effect. However, this does not exclude changes in APC molecular profiles at later times in the course of moderate and severe COVID-19.
Discussion
Severe COVID-19 harbours a complex physiopathology stemming from host–pathogen interactions evolving over time, and involves a large number of underlying cellular and molecular mechanisms. Hence, detailed studies on various immune cell compartments are required to obtain a global view of the process. DCs are central to immune responses by linking innate and adaptive immunity, in particular during infection31. DCs are rare cell types composed of multiple subsets32, justifying dedicated studies to uncover putative dysfunctions. So far, very little is known about the role of DC subsets in COVID-1944,45,46. scRNAseq atlas studies of total PBMCs in patients with severe and moderate COVID-19 identified inflammatory monocytes defective for MHC-II molecules12, as was previously shown in severe sepsis patients47, and increased apoptosis pathways in both NK cells and monocytes27,35,48,49,50. So far, none of these studies were tailored to provide sufficient resolution into the DC compartments. The challenge is even greater knowing that some DC subsets, such as pDCs and CD141 (CLEC9A)+ DCs, are depleted from the blood in severe COVID-1945,51. A recent study analysed PBMCs by scRNAseq, after DC enrichment in EDTA-containing medium, but focused only on the IFN pathway and ISGs30. Most of these studies utilized frozen/thawed PBMCs as a starting biological material, potentially inducing loss in some rare DC subsets. Through dedicated enrichment steps performed immediately after blood sampling (fresh samples), we were able to capture sufficient cell numbers to define molecular profiles and identify specific defects in all known DC subsets.
As with most immune cells, DCs are not limited to a single function31. They play a key role in the first line of immune defence by sensing microbial pathogens, and also contribute to direct pathogen control through the production of antimicrobial peptides and antiviral effector molecules52. Other effector functions include the secretion of pro- and anti-inflammatory cytokines, and cytotoxic molecules31. Finally, they function as APCs to T cells, with which they communicate through secreted and surface molecules expressed within the immune synapse53. By using scRNAseq, and a combination of supervised and unsupervised bioinformatics methods, we were able to uncover defects in almost all of these processes, in specific APC subsets, associated with COVID-19 severity. This provides the first detailed molecular map of DC subsets and underlying molecular pathways in COVID-19.
Several studies have shown an increase of inflammatory cytokines in severe COVID-19, which may contribute to the severity of the disease44. Increased circulating levels of IL-1β and IL-6 were detected in patients with severe COVID-199,10,11,12,13,14,15,16,17,18,19,20,21,22,44. However, the cellular source does not seem to be from circulating cells, but rather from inflammatory monocytes attracted to the lung54, as well as endothelial cells23,24. Our study corroborates these findings for IL-6, with no significant expression detected across APC subsets. However, we did find increased expression of IL-1β, CXCL8 and CXCL2 in APCs at the global level, and this may contribute to systemic inflammation. In parallel, we observed increased TNF signalling in pDCs, but decreased in monocytes, suggesting that distinct APCs may respond differently to circulating inflammatory mediators.
Type I and III IFNs are critical antiviral cytokines55. APCs are a central source of IFN following viral sensing. Studies have shown that type I IFN responses are impaired in severe COVID-196,9,17,21,28,29,30,46,48,56, which may contribute to persistent viral load. Our data support these findings, as we did not detect any expression of IFN-α and IFN-λ1 across all APC subsets. However, we were also able to detect critical defects in the response to type I IFN. First, expression of IFNAR1 and 2 was globally decreased in APC subsets from patients with severe COVID-19. Second, most downstream ISGs (both antiviral and regulators of IFN signalling) were expressed at lower levels in patients with severe COVID-19 compared to HCs, which themselves are expected to express low levels of ISGs given the absence of innate stimulation. Overall, the IFN pathway was defective in severe COVID-19 APCs at several levels: IFN production, receptor expression and downstream ISG responses.
pDCs are a cell type that is highly specialized in antiviral immunity, producing large amounts of all type I IFN57. Circulating pDCs have been shown to be diminished in COVID-1951, but the underlying mechanisms remain unknown. We identified increased expression of pro-apoptotic molecules in pDCs from patients with severe COVID-19. This suggests that pDCs could be globally altered through increased cell death. In a separate study, we have shown that in vitro SARS-CoV-2 stimulation of pDCs from healthy donors leads to improved pDC survival58, suggesting that the increased apoptosis we observed in pDCs from patients with severe COVID-19 was not due to direct virus-induced killing. In parallel, we detected several defects in various pDC functions: decreased innate sensing, through loss of TLR7 and DHX36, which are key viral sensors59, decreased antiviral effector functions and cytotoxicity. Hence, we report multi-process defects affecting key aspects of pDC antiviral functions. Interestingly, a recent study performed ex vivo stimulation of PBMCs of a patient with COVID-19 with TLR7/9 ligands, and showed decreased type I IFN production30. This provides an independent functional validation, while our study provides molecular mechanisms, in particular the increased pDC apoptosis and the decrease in TLR7 expression.
Transcriptomic data, including scRNAseq, allow for the application of methods to infer TF activity, as a way to provide potential upstream mechanisms. We found that several important TF activities were decreased in CD1+ DCs, suggesting defective immune effector functions in patients with severe COVID-19. STAT2 activity downregulation may indicate a deficiency to cross-present to CD8+ T cells and license their cytotoxic function60. Subversion of DC immunogenicity by targeting STAT2 was observed in other viral infections. ZIKV evades type I IFN responses by antagonizing STAT2 phosphorylation61. The low estimated activity of ESR1, CIITA, USF1 and RFX5 in CD1+ DCs may explain the decrease in MHC-II molecules we observed in patients with severe COVID-19, through decreased trans-activation of the MHC-II promoter62,63. Finally, the low activity of EGR1 and RUNX1 TF in CD1+ DCs of patients with severe COVID-19 may contribute to an impaired function in CD8 T-cell activation and induction of IFN-γ64,65. Collectively, our results suggest that several aspects of CD1+ DC effector functions may be altered through decreased activity of key TFs controlling MHC-II expression and T-cell stimulation.
Our study provides a unique insight into the physiopathology of APCs in severe COVID-19, uncovering previously unknown defects in multiple functional pathways, related to both innate and adaptive immunity. We were able to map molecular pathways in rare DC subsets, many of them previously unexplored in the context of COVID-19. Combined with studies in other anatomical sites44, in particular the lung54, and other disease severity stages, our results should contribute to a better understanding of COVID-19 immunopathology. They also open interesting perspectives for clinical applications. Simple molecular markers of defective APC subsets may be explored as prognostic and stratification biomarkers. This hypothesis echoes the immune pathology of bacterial sepsis, for which multiple defects in APCs have already been described47,66. A persistent decrease in circulating DCs, as well as monocyte deactivation as assessed by decreased HLA-DR expression or decreased CD74 messenger RNA (mRNA) expression, are already known to be predictive of ICU-acquired superinfections in patients with bacterial sepsis67,68. It would be interesting to explore whether such markers, for example pDC apoptosis or CD1c+ DC MHC-II downregulation, appear earlier in the course of COVID-19 and may predict aggravation. From a therapeutic standpoint, many innate adjuvants have been developed to target DC subsets69,70, and could be considered as personalized immunotherapies depending on patient-specific DC dysfunction69. Finally, DCs are being considered in preventive vaccine development (ClinicalTrials.gov: NCT04386252). Ultimately, our study may form the ground for novel therapies to restore defective APC functions in patients with COVID-19.
Methods
Patient characteristics and recruitment into the study
Our study is compliant with all relevant ethical regulations regarding research involving human participants.This study was part of the DENDRISEPSIS project, aimed at investigating the functional profiles of APCs in patients with sepsis. The full study protocol can be accessed at https://clinicaltrials.gov/ct2/show/NCT03788772?term=dendrisepsis&cond=sepsis&draw=2&rank=1. The study was approved by the appropriate institutional review board and independent ethics committee (Comité de Protection des Personnes I (CPP), Rouen, France, ref: #2018-A01934-51). We included adult HCs, and patients with PCR-proven COVID-19 pneumonia, within 48 h of admission to ICU or to the pulmonology department from an urban tertiary care centre. Exclusion criteria were the following: haematological malignancy or significant history of bone marrow disease, HIV infection, any immunosuppressive drugs, bone marrow or solid organ transplant recipients, leucopenia (<1,000 mm−3) except if due to COVID-19 or pregnancy. With respect to HCs, exclusion criteria were the following: history of inflammatory disease, corticosteroid treatment at any dose and infection symptoms within the previous month. Informed consent was obtained from patients or next of kin. Patients were classified into moderate pneumonia if requiring oxygen supply of <10 l.min−1 and severe pneumonia if requiring invasive mechanical ventilation or oxygen supply of ≥10 l.min−1. Patients were sampled at admission (day 1) and at day 4. HCs were sampled once. Detailed patient characteristics are provided in Supplementary Tables 1 and 2.
Cell purification
Blood samples (20 ml) were collected from each patient at days 1 and 4 post hospital admission, and from HCs. PBMCs were isolated by centrifugation on a density gradient (Lymphoprep, Proteogenix). After FICOLL (GE Healthcare and Lymphoprep StemCell) gradient centrifugation, total PBMCs were enriched in CD14+ monocytes using human CD14 microbeads (Miltenyi Biotec) for positive magnetic selection according to the manufacturer’s instructions. The negative fraction remaining after the positive selection of CD14+ cells was used for pan-DC enrichment employing the EasySep human pan-DC enrichment kit (StemCell Technologies). Total pan-DCs were resuspended with 20,000 CD14+ cells and sent for sequencing. Monocyte and pan-DC enrichments were performed immediately after sampling. To avoid DC–T cell clusters, which often form in DC-enriched preparations, EDTA-containing medium (DPBS 1×, 0.5% EDTA, 1% human serum) was used for sample enrichment in the first set, the ‘discovery set’. The latter was composed of a total of 12 samples from two healthy donors, three patients with moderate COVID-19 and four with severe COVID-19 from both day 1 and day 4 time points. The reported results in the main figures (Figs. 1–8) along with Supplementary Figs. 1 and 2 were generated based on the discovery set. Because EDTA can decrease reverse transcription efficiency, we validated the findings derived from the discovery set by using RPMI for all enrichment steps (1640 + Glutamax, 2% BSA, 1% penicillin/streptomycin, 1% sodium puryvate, 1% minimum essential medium–non-essential amino acids) in a second set of samples, including a total of 15 samples, defined as the ‘validation set’. The latter included two healthy donors, two patients with moderate COVID-19 and six with severe COVID-19. The results of the validation set are presented in Supplementary Figs. 3–7. All main findings were validated in both datasets.
Preparation and isolation of single-cell suspensions
Cell suspensions were subjected to gel bead emulsion using the Chromium 10X Genomics controller according to the manufacturer’s guidelines. To perform scRNAseq after cDNA amplification, the concentration of each sample was measured using a Tapestation 2200 system (Agilent). To prepare the cDNA libraries for the 10X Genomics Chromium controller, we used the single-cell 3′ v3.1 kit. Quality control libraries were performed using the Tapestation 2200 (Agilent). An Illumina Novaseq6000 system (100-cycle cartridge) with a sequencing depth of at least 50,000 reads per cell was used for sequencing. The input number of cells was estimated at 20,000 cells per sample.
Quality control and pre-processing of expression matrices
The raw scRNAseq fastq files were processed using Cell Ranger 3.1.0 from 10X Genomics Technology and aligned to the Grch38 reference genome. Bam files and filtered expression matrices were generated using ‘cellranger_count’. All expression matrices were loaded into R 4.0.0 using the ‘Read10X’ function from the Seurat library (https://github.com/satijalab/seurat) version 3.1.5. The latter library was used to perform the analysis.
Pre-processing steps were applied to remove genes expressed in fewer than 20 cells, and to remove cells with fewer than 50 genes or displaying more than 50% mitochondrial transcripts. To minimize technical confounding factors related to the sequencing steps, we evaluated the violin plot distribution of the number of unique molecular identifiers (nUMI), along with the total number of detected genes (nFeatures) per cell for all samples. Two upper cutoffs of 6,000 and 50,000 were manually set for the nUMi and nFeatures, respectively, for each sample. These quality control metrics filtered out low-quality cells. Normalization to 10,000 reads, centering and scaling were sequentially applied on the expression matrices to correct for the sequencing depth variability. To reduce the computational time for sample integration, we filtered out cells from cell types other than APCs. Cell type annotation is detailed in the section ‘Manual annotation of cell types’). To decipher specific alterations occurring in each specific APC subset, we separately subclustered each cell subtype, scaled the data and applied graph-based clustering to obtain cell clusters. Genes encoding for immunoglobulins were removed before performing the subclustering step for each cell type to get rid of ambient RNA.
Integration of individual cell matrices into a merged expression matrix from all the samples
To allow comparison across severity states, we integrated the whole expression matrices from all the samples using the Harmony algorithm. Integration anchors, retrieved from the first 50 principal components using the ‘FindIntegrationAnchors’ Seurat function, were then used to integrate the datasets using the ‘IntegrateData’ function. This crucial step added an ‘integrated’ assay to the Seurat object. Scaling and principal component analysis dimension reduction were performed on the integrated assay with 50 principal components. High-resolution (resolution = 0.8) graph-based clustering and UMAP dimension reduction were conducted to retrieve and visualize cell clusters. ICA dimension reduction was specifically performed for CD14+ monocytes, using 30 dimensions.
Manual annotation of cell types
Cells were manually annotated based on their expressing levels of their respective set of cell-type markers, defined as ‘cell-type signatures’. For each cell-type signature, enrichment scores were computed using the ‘AddModuleScore()’ function per cell with 100 randomly selected control genes, split on 25 bins. Each cell cluster was annotated with a particular cell type if its signature score median value was >0. Cell-type signatures included the following: pDCs, expression of (‘TCF4’, ‘CLEC4C’, ‘IRF7’, ‘IRF8’, ‘LILRA4’, ‘IL3RA’, ‘TLR9’, ‘SPIB’), cDCs (‘ANPEP’, ‘CD1C’, ‘ITGAX’, ‘CST3’, ‘FCER1A’), monocytes (‘CD14’, ‘FCGR1A’, ‘S100A12’, ‘FCGR3A’, ‘MS4A7’, ‘LYZ’, ‘CXCR3’), AS-DCs (‘AXL’, ‘SIGLEC6’, ‘CD22’), NK cells (‘NCAM1’, ‘FCGR3A’, ‘GNLY’, ‘XCL1’, ‘XCL2’, ‘NCR1’, ‘NKG7’), T cells (‘CD3D’, ‘CD3E’, ‘CD3G’), B cells (CD19’, ‘MS4A1’, ‘CD79A’, ‘CD79B’), plasma cells (‘IGHG2’, ‘IGHG1’, ‘IGLC2’, ‘IGHA1’, ‘IGHA2’, ‘IGHA3’, ‘JCHAIN’, ‘IGHM’, ‘XBP1’, ‘MZB1’, ‘CD38’, ‘IGLL5’), erythrocytes (‘HBB’, ‘HBA1’) and platelets (PPBP). Cells that were annotated as non-APC were discarded for each sample, before integration, to avoid high computational load during the integration step. For monocytes and cDCs, a subsequent classification of cells was performed according to their expression levels of monocytes and cDC subset markers (CD14 and FCGR3A for monocytes, CD1C and CLEC9A for cDCs).
Statistical analysis
Differential expression analysis between severity groups was performed using the ‘FindAllMarkers’ Seurat function, using the MAST test and a cutoff set to log FC > 0.3 to filter out false-positive DEGs. We regressed out the ‘gender’ confounding factor by using the ‘MAST’ test for comparative analysis and precising ‘gender’ as a latent variable. This ‘gender’ variable was added in the metadata slot for each cell from the discovery and validation sets: a cell is annotated as from a ‘female’ sample if the expression level of the XIST gene is higher than 0.1, otherwise the gender is annotated as ‘male’. P values were corrected using the Bonferroni correction method. We only tested genes that were detected in a minimum fraction of 10% of each severity group. Median values of violin plot distributions of either gene expression levels or pathway-enrichment scores were compared using a Mann–Whitney–Wilcoxon ranked test, taking as a reference the HC. Note that the statistical calculations for the violin plot distributions are derived from the cell count in expression values/enrichment scores comparisons.
Pathway enrichment analysis
Pathway enrichment analysis was performed to seek for the perturbed or enriched pathways in severity groups, as compared to the HCs. Human MsigDB hallmark signatures (https://www.gsea-msigdb.org/gsea/msigdb/index.jsp) were loaded into the R session using the ‘msigdbr’ library version 7.0.1, and the category was set to ‘H’ for ‘human’. The enrichment test was performed using the ‘enricher’ function from ‘ClusterProfiler’ version 3.16.0. Msig Database hallmark signatures were given as input to the ‘enricher’ function. The P values were corrected using the Bonferroni correction method. Encoding genes for each enriched pathway were extracted and used as the module to construct a ‘pathway-score’ signature using ‘AddModuleScore’ from the Seurat library.
TF activity inference
We sought to decipher the variation of TF activity between severity groups within particular cell types to avoid capturing differentially active TFs related to lineage markers. The Dorothea (https://saezlab.github.io/dorothea/) resource was used to infer TF activity. In this context of single-cell-level resolution, we constructed regulons based on the mRNA expression levels of each TF from a manually curated database, along with the expression level of its direct targets. In this context, TF activity is considered as a proxy of the transcriptional state of its direct targets. We created TF regulons using the ‘dorothea_regulon_human’ wrapper function from ‘dorothea’ library version 0.99.10, and chose ‘A’ and ‘B’ high-confidence TF selection. Viper scores were computed on the dorothea regulons, scaled and added as the ‘Dorothea’ slot on the integrated Seurat object. To allow comparison of TF score activities, mean and standard deviation values of the scaled viper scores were computed per severity group. TFs were ranked according to the variance of their corresponding viper scores. The top 50 highly variable scores per severity group (n = 150 TFs in total) were kept for visualization of their corresponding scores.
Manual construction of functional signatures
To evaluate the dysregulations occurring at the functional level for each APC subset from patients with COVID-19, we established a manually curated list of effector genes involved in specific APC functions: ‘attraction’, ‘antiviral effector molecules’ and ‘cytotoxicity’. The signature construction relied on a thorough mining of existing literature, using a combination of MeSH terms and keywords on the PubMed search tool. Each selected molecule was considered an ‘effector’ of the related function if there was at least one experimental proof in a human setting. Overall, we outlined 12 ‘cytotoxicity’ effector molecules, 29 ‘antiviral’ effector molecules and 18 ‘attraction’ effector molecules. ‘Innate sensing’ effectors included 13 genes (DDX58, DHX58, CGAS, IFI16, AIM2, IRF3, TMEM173, NLRP3, PYCARD, TLR7, TLR9, DHX9 and DHX36), and were from refs. 71,72. Both ‘regulators of interferon signalling’ and ‘antiviral ISG’ were implemented by literature mining from ref. 73.
Drop-out correction
To allow drop-out correction and imputation of missing values, we used Nebulosa (https://github.com/powellgenomicslab/Nebulosa) to represent density-based values on UMAP embeddings. This R package is designed to visualize features from single cells, using a kernel density estimation. It recovers the signal by incorporating the similarity between cells, allowing a convolution of the cell features. For pDCs from the discovery set, we specifically added a ‘MAGIC_RNA’ slot to the Seurat object using MAGIC74 and specifically plotted the violin distribution of imputed values in Fig. 4e.
Pseudotime inference
For the CD1c+ DC subset, we specifically computed pseudotime inference using Monocle3 (https://cole-trapnell-lab.github.io/), directly available using the Seurat Wrappers R package75.
Analysis of intercellular communication networks
Communication scores were generated using the ICELLNET R package (https://github.com/soumelis-lab/ICELLNET/master). This library allows computation of cell–cell communication scores between cell subsets, given their corresponding transcriptomic profiles from the same or different datasets. Considering severity groups separately, only clusters including more than 15 cells were considered for the analysis. The average gene expression profiles of APC subset clusters were provided as input to the ICELLNET package, to compute communication scores between APC subsets and T lymphocytes for each severity group. As our datasets did not include T cells for the analysis, we used as reference the T-lymphocyte transcriptomic profile from the Human Primary Cell Atlas included in the ICELLNET package (n = 39). From the ICELLNET ligand–receptor interaction database, we only selected the 144 interactions known to be involved in DC–T communication76. Barplot and dot plot representations were generated to compare the proportions of communication type scores (checkpoint, cytokines, chemokines) among severity groups.
Statistics and reproducibility
Statistical analysis was performed using R (version 4.0.0). A two-sided Wilcoxon ranked-sum test was used to perform pairwise comparisons. To ensure the reproducibility of our main findings, we split our data analysis cohort into a discovery and a validation set. We reported our main findings from the discovery set and conducted similar analyses on the validation set.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this Article.
Data availability
scRNAseq data that support the findings of this study have been deposited in the Gene Expression Omnibus under accession code GSE169346. Further information and requests for resources and reagents should be directed to and will be fulfilled by the V.S. This study did not generate new unique reagents.
Code availability
The R codes are publicly available on GitHub at https://github.com/MelissaSaichi/Covid_scRNAseq. All of the R packages that were used are available online.
References
Wang, F. et al. The laboratory tests and host immunity of COVID-19 patients with different severity of illness. JCI Insight 5, e137799 (2020).
Pedersen, S. F. & Ho, Y.-C. SARS-CoV-2: a storm is raging. J. Clin. Invest. 130, 2202–2205 (2020).
Yan, R. et al. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367, 1444–1448 (2020).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Olagnier, D. et al. SARS-CoV2-mediated suppression of NRF2-signaling reveals potent antiviral and anti-inflammatory activity of 4-octyl-itaconate and dimethyl fumarate. Nat. Commun. 11, 4938 (2020).
Galani, I.-E. et al. Untuned antiviral immunity in COVID-19 revealed by temporal type I/III interferon patterns and flu comparison. Nat. Immunol. 22, 32–40 (2021).
Channappanavar, R. & Perlman, S. Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Semin. Immunopathol. 39, 529–539 (2017).
Ong, E. Z. et al. A dynamic immune response shapes COVID-19 progression. Cell Host Microbe 27, 879–882 (2020).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Sadanandam, A. et al. A blood transcriptome-based analysis of disease progression, immune regulation and symptoms in coronavirus-infected patients. Cell Death Discov. 6, 141 (2020).
Laing, A. G. et al. A dynamic COVID-19 immune signature includes associations with poor prognosis. Nat. Med. 26, 1623–1635 (2020).
Wilk, A. J. et al. A single-cell atlas of the peripheral immune response in patients with severe COVID-19. Nat. Med. 26, 1070–1076 (2020).
Del Valle, D. M. et al. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat. Med. 26, 1636–1643 (2020).
Giamarellos-Bourboulis, E. J. et al. Complex immune dysregulation in COVID-19 patients with severe respiratory failure. Cell Host Microbe 27, 992–1000 (2020).
Chua, R. L. et al. COVID-19 severity correlates with airway epithelium–immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 38, 970–979 (2020).
Mangalmurti, N. & Hunter, C. A. Cytokine storms: understanding COVID-19. Immunity 53, 19–25 (2020).
Lee, J. S. et al. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci. Immunol. 5, eabd1554 (2020).
Jøntvedt Jørgensen, M. et al. Increased interleukin-6 and macrophage chemoattractant protein-1 are associated with respiratory failure in COVID-19. Sci. Rep. 10, 21697 (2020).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Mann, E. R. et al. Longitudinal immune profiling reveals key myeloid signatures associated with COVID-19. Sci. Immunol. 5, eabd6197 (2020).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 (2020).
Zhu, L. et al. Single-cell sequencing of peripheral mononuclear cells reveals distinct immune response landscapes of COVID-19 and influenza patients. Immunity 53, 685–696 (2020).
O’Sullivan, J. M., Gonagle, D. M., Ward, S. E., Preston, R. J. S. & O’Donnell, J. S. Endothelial cells orchestrate COVID-19 coagulopathy. Lancet Haematol. https://doi.org/10.1016/S2352-3026(20)30215-5 (2020).
Pons, S., Fodil, S., Azoulay, E. & Zafrani, L. The vascular endothelium: the cornerstone of organ dysfunction in severe SARS-CoV-2 infection. Crit. Care 24, 353 (2020).
Schied, A., Trovillion, E. & Moodley, A. Sars-CoV-2 infection in a neutropenic pediatric patient with leukemia: addressing the need for universal guidelines for treatment of Sars-CoV-2 positive, immunocompromised patients. Pedriat. Blood Cancer https://doi.org/10.1002/pbc.28546(2020).
Somers, E. C. et al. Tocilizumab for treatment of mechanically ventilated patients with COVID-19. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa954 (2020).
Guo, C. et al. Single-cell analysis of two severe COVID-19 patients reveals a monocyte-associated and tocilizumab-responding cytokine storm. Nat. Commun. 11, 3924 (2020).
Lei, C. et al. Neutralization of SARS-CoV-2 spike pseudotyped virus by recombinant ACE2-Ig. Nat. Commun. 11, 2070 (2020).
Zhang, Q. et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 370, eabd4570 (2020).
Arunachalam, P. S. et al. Systems biological assessment of immunity to mild versus severe COVID-19 infection in humans. Science https://doi.org/10.1126/science.abc6261 (2020).
Banchereau, J. et al. Immunobiology of dendritic cells. Annu. Rev. Immunol. 18, 767–811 (2000).
Collin, M. & Bigley, V. Human dendritic cell subsets: an update. Immunology 154, 3–20 (2018).
Coquerelle, C. & Moser, M. DC subsets in positive and negative regulation of immunity. Immunol. Rev. 234, 317–334 (2010).
Liu, Y.-J. IPC: professional type 1 interferon-producing cells and plasmacytoid dendritic cell precursors. Annu. Rev. Immunol. 23, 275–306 (2005).
Zhang, J.-Y. et al. Single-cell landscape of immunological responses in patients with COVID-19. Nat. Immunol. 21, 1107–1118 (2020).
Yao, C. et al. Cell-type-specific immune dysregulation in severely ill COVID-19 patients. Cell Rep. 34, 108590 (2021).
Xu, G. et al. The differential immune responses to COVID-19 in peripheral and lung revealed by single-cell RNA sequencing. Cell Discov. 6, 73 (2020).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Villani, A.-C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes and progenitors. Science 356, eaah4573 (2017).
See, P. et al. Mapping the human DC lineage through the integration of high-dimensional techniques. Science 356, eaag3009 (2017).
Holland, C. H. et al. Robustness and applicability of transcription factor and pathway analysis tools on single-cell RNA-seq data. Genome Biol. 21, 36 (2020).
Alvarez, M. J. et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat. Genet. 48, 838–847 (2016).
Noël, F. et al. Dissection of intercellular communication using the transcriptome-based framework ICELLNET. Nat. Commun. 12, 1089 (2021).
Vabret, N. et al. Immunology of COVID-19: current state of the science. Immunity 52, 910–941 (2020).
Zhou, R. et al. Acute SARS-CoV-2 infection impairs dendritic cell and T cell responses. SSRN Electronic Journal https://doi.org/10.2139/ssrn.3614132 (2020).
Zheng, C. et al. Risk-adapted treatment strategy for COVID-19 patients. Int. J. Infect. Dis. 94, 74–77 (2020).
Delano, M. J. & Ward, P. A. Sepsis-induced immune dysfunction: can immune therapies reduce mortality? J. Clin. Invest. 126, 23–31 (2016).
Yao, H. et al. Molecular architecture of the SARS-CoV-2 virus. Cell 183, 730–738 (2020).
Schulte-Schrepping, J. et al. Severe COVID-19 is marked by a dysregulated myeloid cell compartment. Cell 182, 1419–1440 (2020).
Liao, M. et al. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat. Med. 26, 842–844 (2020).
Sanchez-Cerrillo, I. et al. Differential redistribution of activated monocyte and dendritic cell subsets to the lung associates with severity of COVID-19. Preprint at bioRxiv https://doi.org/10.1101/2020.05.13.20100925 (2020).
Bedoui, S. & Greyer, M. The role of dendritic cells in immunity against primary herpes simplex virus infections. Front. Microbiol. 5, 533 (2014).
Guermonprez, P., Valladeau, J., Zitvogel, L., Théry, C. & Amigorena, S. Antigen presentation and T cell stimulation by dendritic cells. Annu. Rev. Immunol. 20, 621–667 (2002).
Zhou, Y. et al. Pathogenic T-cells and inflammatory monocytes incite inflammatory storms in severe COVID-19 patients. Natl Sci. Rev. 7, 998–1002 (2020).
Broggi, A., Granucci, F. & Zanoni, I. Type III interferons: balancing tissue tolerance and resistance to pathogen invasion. J. Exp. Med. 217, e20190295 (2020).
Stanifer, M. L. et al. Critical role of type III interferon in controlling SARS-CoV-2 infection in human intestinal epithelial cells. Cell Rep. 32, 107863 (2020).
Vremec, D. et al. Production of interferons by dendritic cells, plasmacytoid cells, natural killer cells and interferon-producing killer dendritic cells. Blood 109, 1165–1173 (2007).
Onodi, F. et al. SARS-CoV-2 induces human plasmacytoid predendritic cell diversification via UNC93B and IRAK4. J. Exp. Med. 218, e20201387 (2021).
Zhang, Z. et al. DDX1, DDX21 and DHX36 helicases form a complex with the adaptor molecule TRIF to sense dsRNA in dendritic cells. Immunity 34, 866–878 (2011).
Xu, J. et al. STAT2 is required for TLR-induced murine dendritic cell activation and cross-presentation. J. Immunol. 197, 326–336 (2016).
Bowen, J. R. et al. Zika Virus antagonizes type I interferon responses during infection of human dendritic cells. PLoS Pathog. 13, e1006164 (2017).
Mach, B., Steimle, V., Martinez-Soria, E. & Reith, W. Regulation of MHC class II genes: lessons from a disease. Annu. Rev. Immunol. 14, 301–331 (1996).
Muhlethaler-Mottet, A., Di Berardino, W., Otten, L. A. & Mach, B. Activation of the MHC class II transactivator CIITA by interferon-γ requires cooperative interaction between Stat1 and USF-1. Immunity 8, 157–166 (1998).
Rodríguez-Ubreva, J. et al. Prostaglandin E2 leads to the acquisition of DNMT3A-dependent tolerogenic functions in human myeloid-derived suppressor cells. Cell Rep. 21, 154–167 (2017).
Singh, A., Svaren, J., Grayson, J. & Suresh, M. CD8 T cell responses to lymphocytic choriomeningitis virus in early growth response gene 1-deficient mice. J. Immunol. 173, 3855–3862 (2004).
Boomer, J. S. et al. Immunosuppression in patients who die of sepsis and multiple organ failure. JAMA 306, 2594–2605 (2011).
Grimaldi, D. et al. Profound and persistent decrease of circulating dendritic cells is associated with ICU-acquired infection in patients with septic shock. Intensive Care Med. 37, 1438–1446 (2011).
Peronnet, E. et al. Association between mRNA expression of CD74 and IL10 and risk of ICU-acquired infections: a multicenter cohort study. Intensive Care Med. 43, 1013–1020 (2017).
Bryant, C. E. et al. Dendritic cells as cancer therapeutics. Semin. Cell Dev. Biol. 86, 77–88 (2019).
Saxena, M. & Bhardwaj, N. Turbocharging vaccines: emerging adjuvants for dendritic cell based therapeutic cancer vaccines. Curr. Opin. Immunol. 47, 35–43 (2017).
Zhang, Z., Yuan, B., Lu, N., Facchinetti, V. & Liu, Y.-J. DHX9 pairs with IPS-1 to sense double-stranded RNA in myeloid dendritic cells. J. Immunol. 187, 4501–4508 (2011).
Gaidt, M. M. et al. The DNA inflammasome in human myeloid cells is initiated by a STING-cell death program upstream of NLRP3. Cell 171, 1110–1124.e18 (2017).
Schneider, W. M., Chevillotte, M. D. & Rice, C. M. Interferon-stimulated genes: a complex web of host defenses. Annu. Rev. Immunol. 32, 513–545 (2014).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729.e27 (2018).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Grandclaudon, M. et al. A quantitative multivariate model of human dendritic cell-T helper cell communication. Cell 179, 432–447.e21 (2019).
Acknowledgements
We thank our patients for participating in this study. We also acknowledge N. Marin, M. Cojocaru, N. Carlier, J. Marey, D. Monnet and T.-A. Szwebel (all from Cochin Hospital) for their help with the inclusion of patients and controls. We thank A. Mohammed for performing the sequencing runs on Cell Ranger at the sequencing platform of the Institut du Cerveau et de la Moelle épinière (ICM). We thank M. Blot, F. Niedergang and E. Lauret from the Institut Cochin for their help during the setting up of the study. The COVID-19 emergency plan from Université de Paris supported this study regarding technical aspects for the efficient processing of sample sequencing. This study was supported by the ANR DENDRISEPSIS (ANR-17-CE15-0003), ANR APCOD (ANR-17-CE15-0003-01), Fast Grant for COVID-19 from the Mercatus Center, Université de Paris PLAN D’URGENCE COVID19 and Fund 101 grants. L.M. was supported by a PhD fellowship from La Ligue Contre le Cancer and E.A. by a PhD fellowship from Servier. We thank Fast Grant for COVID-19 from the Mercus Center for supporting F.N. and la Fondation pour la Recherche Médicale for supporting J.M.
Author information
Authors and Affiliations
Contributions
V.S., J.M. and F.P. designed the study and defined the patient selection criteria. F.P. and Z.A.H. recruited the patients and healthy donors. M.Z.L., S.K. and C.R. performed the wet lab experiments. M.S. designed a workflow for the scRNAseq, analysed the data and generated the results and figures. L.M. performed the cell–cell communication analysis using ICELLNET. E.A. constructed the manually curated effector molecule signatures. F.N. helped with the bioinformatics analysis. D.B. and Y.M. performed and supervised the library construction and sequencing. V.S., M.S., M.Z.L., S.K. and F.P. wrote the manuscript. S.K. updated the bibliography. All authors provided feedback for the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Bryan Williams and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Pro-inflammatory defects in the discovery set.
a, Umap representation of IFN subtypes expression values in the discovery set, expression levels are color coded; b, Violin representation of other pro-inflammatory cytokines in the discovery set which included: n=2 HC, n=4 moderate and n=6 severe samples; each dot represents a cell, horizontal lines display the mean expression value; Comparative analysis was performed using the two-sided Wilcoxon Rank-Sum test, P-values were adjusted to multiple testings using ‘Bonferroni’ correction. Asterisks above severe indicate P values for severe versus control; asterisks above moderate indicate significance of moderate versus control. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 2 Maintenance of dysregulated patterns at time and patient levels.
Violin plot representation of a. HLA-II Module Score in CD1c+DC, and b. P53 pathway Module Score in pDC, both from the discovery set (composed of: n=2 HC, n=4 moderate and n=6 severe samples); asterisks above moderate indicate significance of moderate versus control and asterisks above severe indicate significance of severe versus control. Comparative analysis was performed using the two-sided Wilcoxon Rank-Sum test, P-values were adjusted to multiple testings using ‘Bonferroni’ correction. *P < 0.05, **P < 0.01, ***P < 0.001. c, Dot Plot representation of antiviral effector molecules in CD14+ Monocytes across patients, Percentage of cells expressing the respective gene is size-coded.
Extended Data Fig. 3 Cellular map of APC subsets from the validation set.
Cellular map of APC subsets at the single-cell resolution level from the validation set based on either APC subsets (a) or severity (b), c Proportions of APC subsets within the discovery and validation sets; d. Stuck Violin plot representation of canonical APC and non-APC markers for both discovery and validation sets. Validation set included: n=2 HC, n= 4 moderate and n=9 severe samples from a total of 2 healthy donors, 2 moderate and 6 severe patients.
Extended Data Fig. 4 Increased inflammatory pathways in APC validation set.
a. Barplot of the number of Differentially Expressed Genes (DEG) for each severity group (Healthy versus moderate and severe patients; moderate versus healthy and severe; severe versus healthy and moderate). Up-regulated (logFoldChange > 0.25) genes are shown in black, down-regulated (logFoldChange < −0.25) genes are shown in grey; b. Heatmap representation of top up-regulated genes in severe APC from the validation set, as compared to moderate and healthy groups, z-score values of average expression levels of cells per severity group is color coded; Comparative analysis of enriched pathways from the upregulated genes in moderate or severe (c) APC as compared to healthy cells, as well as pairwise comparison of upregulated genes in moderate compared to severe (shown in pink), up-regulated genes in severe compared to moderate (shown in yellow) (d); horizontal axis displays the adjusted p-values (-log10), e. Representation of ranked genes by descendant order according to their absolute log Fold Change (log FC), upregulated in moderate as compared to severe (plot in red), upregulated in severe APC as compared to moderate (plot in blue).Top genes, with an absolute value of logFC above 0.5 are shown. Validation set included: n=2 HC, n= 4 moderate and n=9 severe samples from a total of 2 healthy donors, 2 moderate and 6 severe patients. The two-sided Wilcoxon Rank-Sum test was used for comparison, P-values were adjusted to multiple testing using ‘Bonferroni’ correction; and only genes with adjusted-P Values < 0.05 were considered.
Extended Data Fig. 5 Global defects in pDC-related functions in the validation set.
a. Dot plots of pDC-related functions ‘Attraction’,’Innate sensing’, ‘Anti-viral effector molecules’, ‘Cytotoxicity’ in pDC from HC, moderate and severe patients in the validation set. Expression levels are color-coded; Percentage of cells expressing the respective gene is size-coded, b. Comparative analysis of enriched pathways from the upregulated genes in moderate versus severe pDC (in pink),up-regulated genes in severe compared to moderate (shown in yellow); c. Violin plot representation of gene expression for IFN receptors (IFNAR1 and 2), IRF7, and anti-viral effector molecules. Asterisks above severe indicate P values for severe versus control; asterisks above moderate indicate significance of moderate versus control. Statistical tests were performed using the validation set, including: n=2 HC, n= 4 moderate and n=9 severe samples; Comparative analysis was performed using the two-sided Wilcoxon Rank-Sum test, P-values were adjusted to multiple testings using ‘Bonferroni’ correction. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 6 Defective anti-viral properties in CD14+ monocytes and CD1c+DC.
Dot plots of ‘antiviral effector molecules’ in CD14+ monocytes from HC, moderate and severe patients in the validation set. Expression levels are color-coded; Percentage of cells expressing the respective gene is size coded.
Extended Data Fig. 7 MHC-II antigen presentation defects in CD1c+DC.
Heatmap representation of top 10 DEG (upregulated) for each severity group in CD1C+DC from the validation set.
Supplementary information
Supplementary Tables 1–3
Supplementary Table 1: Clinical characteristics of the included patients. Supplementary Table 2: Detailed description of the clinical characteristics of the included patients (n = 15) and healthy controls (n = 4). Supplementary Table 3: Repartition of APC subsets within the recruited samples from both the Discovery and Validation sets
Rights and permissions
About this article
Cite this article
Saichi, M., Ladjemi, M.Z., Korniotis, S. et al. Single-cell RNA sequencing of blood antigen-presenting cells in severe COVID-19 reveals multi-process defects in antiviral immunity. Nat Cell Biol 23, 538–551 (2021). https://doi.org/10.1038/s41556-021-00681-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00681-2
This article is cited by
-
Impaired balance between neutrophil extracellular trap formation and degradation by DNases in COVID-19 disease
Journal of Translational Medicine (2024)
-
Single cell RNA-seq: a novel tool to unravel virus-host interplay
VirusDisease (2024)
-
Longitudinal single cell atlas identifies complex temporal relationship between type I interferon response and COVID-19 severity
Nature Communications (2024)
-
Longitudinal whole blood transcriptomic analysis characterizes neutrophil activation and interferon signaling in moderate and severe COVID-19
Scientific Reports (2023)
-
Multimodal single-cell analyses of peripheral blood mononuclear cells of COVID-19 patients in Japan
Scientific Reports (2023)