Abstract
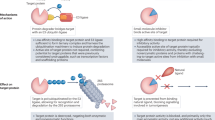
For decades, anticancer targeted therapies have been designed to inhibit kinases or other enzyme classes and have profoundly benefited many patients. However, novel approaches are required to target transcription factors, scaffolding proteins and other proteins central to cancer biology that typically lack catalytic activity and have remained mostly recalcitrant to drug development. The selective degradation of target proteins is an attractive approach to expand the druggable proteome, and the selective oestrogen receptor degrader fulvestrant served as an early example of this concept. Following a long and tragic history in the clinic, the immunomodulatory imide drug (IMiD) thalidomide was discovered to exert its therapeutic activity via a novel and unexpected mechanism of action: targeting proteins to an E3 ubiquitin ligase for subsequent proteasomal degradation. This discovery has paralleled and directly catalysed myriad breakthroughs in drug development, leading to the rapid maturation of generalizable chemical platforms for the targeted degradation of previously undruggable proteins. Decades of clinical experience have established front-line roles for thalidomide analogues, including lenalidomide and pomalidomide, in the treatment of haematological malignancies. With a new generation of ‘degrader’ drugs currently in development, this experience provides crucial insights into class-wide features of degraders, including a unique pharmacology, mechanisms of resistance and emerging therapeutic opportunities. Herein, we review these past experiences and discuss their application in the clinical development of novel degrader therapies.
Key points
-
Thalidomide analogues including lenalidomide and pomalidomide are anticancer agents used in the treatment of multiple haematological malignancies; induction of targeted protein degradation of disease-relevant proteins is the dominant mechanism of action of these agents.
-
‘Degrader’ therapeutics have a catalytic, event-driven pharmacology, and their biological effects are a summation of the polypharmacology of multiple degraded neosubstrate proteins.
-
Cancers acquire resistance to thalidomide analogues by circumventing target protein degradation or by rewiring pathways downstream of target protein degradation.
-
Quantitative clinical proteomic assays can be used for pharmacodynamic monitoring of degrader therapeutics.
-
Molecular switches gated by degrader therapeutics can be used to control genetically engineered adoptive cell products that are being developed for anticancer immunotherapy.
-
Multiple classes of degrader therapeutics are expanding the druggable proteome to include previously undruggable, noncatalytic, cancer-relevant proteins, such as transcription factors, mRNA-splicing factors and cyclins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Stephens, T. & Brynner, R. Dark Remedy: The Impact of Thalidomide and its Revival as a Vital Medicine (Perseus Publishing, 2009).
Lenz, W., Pfeiffer, R. A., Kosenow, W. & Hayman, D. J. Thalidomide and congenital abnormalities. Lancet 279, 45–46 (1962).
Mcbride, W. G. Thalidomide and congenital abnormalities. Lancet 278, 1358 (1961).
Mintz, M. “Heroine” of FDA keeps bad drug off of market. Washington Post (15 Jul 1962).
Kaur, I., Dogra, S., Narang, T. & De, D. Comparative efficacy of thalidomide and prednisolone in the treatment of moderate to severe erythema nodosum leprosum: a randomized study. Australas. J. Dermatol. 50, 181–185 (2009).
Pearson, J. M. H. & Vedagiri, M. Treatment of moderately severe erythema nodosum leprosum with thalidomide - a double-blind controlled trial. Lepr. Rev. 40, 111–116 (1969).
D’Amato, R. J., Loughnan, M. S., Flynn, E. & Folkman, J. Thalidomide is an inhibitor of angiogenesis. Proc. Natl Acad. Sci. USA 91, 4082–4085 (1994).
Singhal, S. et al. Antitumor activity of thalidomide in refractory multiple myeloma. N. Engl. J. Med. 341, 1565–1571 (1999).
Rajkumar, S. V. et al. Combination therapy with thalidomide plus dexamethasone for newly diagnosed myeloma. J. Clin. Oncol. 20, 4319–4323 (2002).
Rajkumar, S. V. et al. Phase III clinical trial of thalidomide plus dexamethasone compared with dexamethasone alone in newly diagnosed multiple myeloma: a clinical trial coordinated by the Eastern Cooperative Oncology Group. J. Clin. Oncol. 24, 431–436 (2006).
Krönke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2013).
Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305–309 (2013).
Gandhi, A. K. et al. Immunomodulatory agents lenalidomide and pomalidomide co-stimulate T cells by inducing degradation of T cell repressors Ikaros and Aiolos via modulation of the E3 ubiquitin ligase complex CRL4 CRBN. Br. J. Haematol. 164, 811–821 (2013).
Ito, T. et al. Identification of a primary target of thalidomide teratogenicity. Science 327, 1345–1350 (2010).
Krönke, J. et al. Lenalidomide induces ubiquitination and degradation of CK1α in del(5q) MDS. Nature 523, 183–188 (2015).
Fink, E. C. & Ebert, B. L. The novel mechanism of lenalidomide activity. Blood 126, 2366–2369 (2015).
Fischer, E. S. et al. Structure of the DDB1–CRBN E3 ubiquitin ligase in complex with thalidomide. Nature 512, 49–53 (2014).
Bartlett, J. B., Dredge, K. & Dalgleish, A. G. The evolution of thalidomide and its IMiD derivatives as anticancer agents. Nat. Rev. Cancer 4, 314–322 (2004).
Ramsay, A. G. et al. Chronic lymphocytic leukemia T cells show impaired immunological synapse formation that can be reversed with an immunomodulating drug. J. Clin. Invest. 118, 2427–2437 (2008).
Hagner, P. R. et al. CC-122, a pleiotropic pathway modifier, mimics an interferon response and has antitumor activity in DLBCL. Blood 126, 779–789 (2015).
Luptakova, K. et al. Lenalidomide enhances anti-myeloma cellular immunity. Cancer Immunol. Immunother. 62, 39–49 (2013).
Muller, G. W. et al. Structural modifications of thalidomide produce analogs with enhanced tumor necrosis factor inhibitory activity. J. Med. Chem. 39, 3238–3240 (1996).
Choueiri, T. K. et al. Phase II study of lenalidomide in patients with metastatic renal cell carcinoma. Cancer 107, 2609–2616 (2006).
Eisen, T. et al. Results of a multicenter, randomized, double-blind phase 2/3 study of lenalidomide in the treatment of pretreated relapsed or refractory metastatic malignant melanoma. Cancer 116, 146–154 (2009).
Polizzotto, M. N. et al. Pomalidomide for symptomatic kaposi’s sarcoma in people with and without HIV infection: a phase I/II study. J. Clin. Oncol. 34, 4125–4131 (2016).
Weber, D. M. et al. Lenalidomide plus dexamethasone for relapsed multiple myeloma in North America. N. Engl. J. Med. 357, 2133–2142 (2007).
Dimopoulos, M. et al. Lenalidomide plus dexamethasone for relapsed or refractory multiple myeloma. N. Engl. J. Med. 357, 2123–2132 (2007).
Leonard, J. P. et al. AUGMENT: a phase III study of lenalidomide plus rituximab versus placebo plus rituximab in relapsed or refractory indolent lymphoma. J. Clin. Oncol. 19, 1188–1199 (2019).
List, A. et al. Lenalidomide in the myelodysplastic syndrome with chromosome 5q deletion. N. Engl. J. Med. 355, 1456–1465 (2006).
FDA. FDA grants accelerated approval to pomalidomide for Kaposi sarcoma. US Food & Drug Administration https://www.fda.gov/drugs/resources-information-approved-drugs/fda-grants-accelerated-approval-pomalidomide-kaposi-sarcoma (2020).
Richardson, P. G. et al. Immunomodulatory drug CC-5013 overcomes drug resistance and is well tolerated in patients with relapsed multiple myeloma. Blood 100, 3063–3067 (2002).
Rajkumar, S. V. et al. Combination therapy with lenalidomide plus dexamethasone (Rev/Dex) for newly diagnosed myeloma. Blood 106, 4050–4053 (2005).
Richardson, P. G. et al. A randomized phase 2 study of lenalidomide therapy for patients with relapsed or relapsed and refractory multiple myeloma. Blood 108, 3458–3464 (2006).
Benboubker, L. et al. Lenalidomide and dexamethasone in transplant-ineligible patients with myeloma. N. Engl. J. Med. 371, 906–917 (2014).
Richardson, P. G. et al. Lenalidomide, bortezomib, and dexamethasone combination therapy in patients with newly diagnosed multiple myeloma. Blood 116, 679–686 (2010).
Facon, T. et al. Daratumumab plus lenalidomide and dexamethasone for untreated myeloma. N. Engl. J. Med. 380, 2104–2115 (2019).
McCarthy, P. L. et al. Lenalidomide after stem-cell transplantation for multiple myeloma. New Engl. J. Med. 366, 1770–1781 (2012).
Attal, M. et al. Lenalidomide maintenance after stem-cell transplantation for multiple myeloma. N. Engl. J. Med. 366, 1782–1791 (2012).
Barlogie, B. et al. Thalidomide and hematopoietic-cell transplantation for multiple myeloma. N. Engl. J. Med. 354, 1021–1030 (2006).
Palumbo, A. et al. Autologous transplantation and maintenance therapy in multiple myeloma. N. Engl. J. Med. 371, 895–905 (2014).
McCarthy, P. L. et al. Lenalidomide maintenance after autologous stem-cell transplantation in newly diagnosed multiple myeloma: a meta-analysis. J. Clin. Oncol. 35, 3279–3289 (2017).
Goldschmidt, H. et al. Bortezomib before and after high-dose therapy in myeloma: long-term results from the phase III HOVON-65/GMMG-HD4 trial. Leukemia 32, 383–390 (2017).
Palumbo, A. et al. Continuous therapy versus fixed duration of therapy in patients with newly diagnosed multiple myeloma. J. Clin. Oncol. 33, 3459–3466 (2015).
Madan, S. et al. Efficacy of retreatment with immunomodulatory drugs (IMiDs) in patients receiving IMiDs for initial therapy of newly diagnosed multiple myeloma. Blood 118, 1763–1765 (2011).
Wang, M. et al. Lenalidomide plus dexamethasone is more effective than dexamethasone alone in patients with relapsed or refractory multiple myeloma regardless of prior thalidomide exposure. Blood 112, 4445–4451 (2008).
Stewart, A. K. et al. Carfilzomib, lenalidomide, and dexamethasone for relapsed multiple myeloma. N. Engl. J. Med. 372, 142–152 (2015).
Moreau, P. et al. Oral ixazomib, lenalidomide, and dexamethasone for multiple myeloma. N. Engl. J. Med. 374, 1621–1634 (2016).
Dimopoulos, M. A. et al. Daratumumab, lenalidomide, and dexamethasone for multiple myeloma. N. Engl. J. Med. 375, 1319–1331 (2016).
Siegel, D. S. et al. Pomalidomide plus low‐dose dexamethasone in relapsed refractory multiple myeloma after lenalidomide treatment failure. Br. J. Haematol. 188, 501–510 (2019).
Richardson, P. G. et al. Pomalidomide alone or in combination with low-dose dexamethasone in relapsed and refractory multiple myeloma: a randomized phase 2 study. Blood 123, 1826–1832 (2014).
Lacy, M. Q. et al. Pomalidomide plus low-dose dexamethasone in myeloma refractory to both bortezomib and lenalidomide: comparison of 2 dosing strategies in dual-refractory disease. Blood 118, 2970–2975 (2011).
Richardson, P. G. et al. Pomalidomide, bortezomib, and dexamethasone for patients with relapsed or refractory multiple myeloma previously treated with lenalidomide (OPTIMISMM): a randomised, open-label, phase 3 trial. Lancet Oncol. 20, 781–794 (2019).
Chari, A. et al. Daratumumab plus pomalidomide and dexamethasone in relapsed and/or refractory multiple myeloma. Blood 130, 974–981 (2017).
Dimopoulos, M. A. et al. Elotuzumab plus pomalidomide and dexamethasone for multiple myeloma. N. Engl. J. Med. 379, 1811–1822 (2018).
Stewart, A. K., Richardson, P. G. & San-Miguel, J. F. How I treat multiple myeloma in younger patients. Blood 114, 5436–5443 (2009).
Qian, Z. et al. Lenalidomide synergizes with dexamethasone to induce growth arrest and apoptosis of mantle cell lymphoma cells in vitro and in vivo. Leukemia Res. 35, 380–386 (2011).
Rychak, E. et al. Pomalidomide in combination with dexamethasone results in synergistic anti-tumour responses in pre-clinical models of lenalidomide-resistant multiple myeloma. Br. J. Haematol. 172, 889–901 (2016).
Das, D. S. et al. Synergistic anti-myeloma activity of the proteasome inhibitor marizomib and the IMiD ® immunomodulatory drug pomalidomide. Br. J. Haematol. 171, 798–812 (2015).
Chauhan, D. et al. Combination of novel proteasome inhibitor NPI-0052 and lenalidomide trigger in vitro and in vivo synergistic cytotoxicity in multiple myeloma. Blood 115, 834–845 (2010).
Ocio, E. M. et al. In vitro and in vivo rationale for the triple combination of panobinostat (LBH589) and dexamethasone with either bortezomib or lenalidomide in multiple myeloma. Haematologica 95, 794–803 (2009).
Siegel, D. S. et al. Vorinostat in combination with lenalidomide and dexamethasone in patients with relapsed or refractory multiple myeloma. Blood Cancer J. 4, e202 (2014).
Shi, C.-X. et al. Proteasome inhibitors block Ikaros degradation by lenalidomide in multiple myeloma. Haematologica 100, e315–e317 (2015).
Palmer, A. C. & Sorger, P. K. Combination cancer therapy can confer benefit via patient-to-patient variability without drug additivity or synergy. Cell 171, 1678–1691.e13 (2017).
Orlowski, R. Z. et al. Phase I trial of the proteasome inhibitor PS-341 in patients with refractory hematologic malignancies. J. Clin. Oncol. 20, 4420–4427 (2002).
Chamberlain, P. P. & Hamann, L. G. Development of targeted protein degradation therapeutics. Nat. Chem. Biol. 15, 937–944 (2019).
Zonder, J. A. et al. A phase 1, multicenter, open-label, dose escalation study of elotuzumab in patients with advanced multiple myeloma. Blood 120, 552–559 (2012).
Gavriatopoulou, M., Terpos, E. & Dimopoulos, M. A. The extended 4-year follow-up results of the ELOQUENT-2 trial. Oncotarget 10, 82–83 (2019).
Lonial, S. et al. Elotuzumab therapy for relapsed or refractory multiple myeloma. N. Engl. J. Med. 373, 621–631 (2015).
Balasa, B. et al. Elotuzumab enhances natural killer cell activation and myeloma cell killing through interleukin-2 and TNF-α pathways. Cancer Immunol. Immunother. 64, 61–73 (2014).
Tai, Y.-T. et al. Anti-CS1 humanized monoclonal antibody HuLuc63 inhibits myeloma cell adhesion and induces antibody-dependent cellular cytotoxicity in the bone marrow milieu. Blood 112, 1329–1337 (2008).
Collins, S. M. et al. Elotuzumab directly enhances NK cell cytotoxicity against myeloma via CS1 ligation: evidence for augmented NK cell function complementing ADCC. Cancer Immunol. Immunother. 62, 1841–1849 (2013).
Veer, M. S. van der et al. Towards effective immunotherapy of myeloma: enhanced elimination of myeloma cells by combination of lenalidomide with the human CD38 monoclonal antibody daratumumab. Haematologica 96, 284–290 (2010).
Palumbo, A. et al. Daratumumab, bortezomib, and dexamethasone for multiple myeloma. N. Engl. J. Med. 375, 754–766 (2016).
Goy, A. et al. Single-agent lenalidomide in patients with mantle-cell lymphoma who relapsed or progressed after or were refractory to bortezomib: phase II MCL-001 (EMERGE) study. J. Clin. Oncol. 31, 3688–3695 (2013).
Trněný, M. et al. Lenalidomide versus investigator’s choice in relapsed or refractory mantle cell lymphoma (MCL-002; SPRINT): a phase 2, randomised, multicentre trial. Lancet Oncol. 17, 319–331 (2016).
Leonard, J. P. et al. Randomized trial of lenalidomide alone versus lenalidomide plus rituximab in patients with recurrent follicular lymphoma: CALGB 50401 (Alliance). J. Clin. Oncol. 33, 3635–3640 (2015).
Andorsky, D. J. et al. MAGNIFY: phase IIIb interim analysis of induction R2 followed by maintenance in relapsed/refractory indolent non-Hodgkin lymphoma [abstract]. J. Clin.Oncol. 37, 7513 (2019).
Morschhauser, F. et al. Rituximab plus lenalidomide in advanced untreated follicular lymphoma. N. Engl. J. Med. 379, 934–947 (2018).
Ruan, J. et al. Lenalidomide plus rituximab as initial treatment for mantle-cell lymphoma. N. Engl. J. Med. 373, 1835–1844 (2015).
Badoux, X. C. et al. Phase II study of lenalidomide and rituximab as salvage therapy for patients with relapsed or refractory chronic lymphocytic leukemia. J. Clin. Oncol. 31, 584–591 (2013).
Chanan-Khan, A. A. et al. Lenalidomide maintenance therapy in previously treated chronic lymphocytic leukaemia (CONTINUUM): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Haematol. 4, e534–e543 (2017).
Morschhauser, F. et al. Obinutuzumab combined with lenalidomide for relapsed or refractory follicular B-cell lymphoma (GALEN): a multicentre, single-arm, phase 2 study. Lancet Haematol. 6, e429–e437 (2019).
Yang, Y. et al. Exploiting synthetic lethality for the therapy of ABC diffuse large B cell lymphoma. Cancer Cell 21, 723–737 (2012).
Nowakowski, G. S. et al. Lenalidomide combined with R-CHOP overcomes negative prognostic impact of non-germinal center B-cell phenotype in newly diagnosed diffuse large B-cell lymphoma: a phase II study. J. Clin. Oncol. 33, 251–257 (2015).
Vitolo, U. et al. ROBUST: first report of phase III randomized study of lenalidomide/R-CHOP (R 2 -CHOP) vs placebo/R-CHOP in previously untreated ABC-type diffuse large B-cell lymphoma. Hematol. Oncol. 37, 36–37 (2019).
Ferreri, A. J. M. et al. Lenalidomide maintenance in patients with relapsed diffuse large B-cell lymphoma who are not eligible for autologous stem cell transplantation: an open label, single-arm, multicentre phase 2 trial. Lancet Haematol. 4, e137–e146 (2017).
Carpio, C. et al. Avadomide monotherapy in relapsed/refractory DLBCL: safety, efficacy, and a predictive gene classifier. Blood 135, 996–1007 (2020).
Mullighan, C. G. et al. BCR–ABL1 lymphoblastic leukaemia is characterized by the deletion of Ikaros. Nature 453, 110–114 (2008).
Mullighan, C. G. et al. Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia. N. Engl. J. Med. 360, 470–480 (2009).
Georgopoulos, K. et al. The ikaros gene is required for the development of all lymphoid lineages. Cell 79, 143–156 (1994).
Wang, J.-H. et al. Aiolos regulates B cell activation and maturation to effector state. Immunity 9, 543–553 (1998).
Morschhauser, F. et al. A phase 2, multicentre, single-arm, open-label study to evaluate the safety and efficacy of single-agent lenalidomide (Revlimid®) in subjects with relapsed or refractory peripheral T-cell non-Hodgkin lymphoma: the EXPECT trial. Eur. J. Cancer 49, 2869–2876 (2013).
Ishida, T. et al. Multicenter phase II study of lenalidomide in relapsed or recurrent adult T-cell leukemia/lymphoma: ATLL-002. J. Clin. Oncol. 34, 4086–4093 (2016).
Toumishey, E. et al. Final report of a phase 2 clinical trial of lenalidomide monotherapy for patients with T-cell lymphoma: lenalidomide therapy for T-cell lymphoma. Cancer 121, 716–723 (2014).
Querfeld, C. et al. Results of an open-label multicenter phase 2 trial of lenalidomide monotherapy in refractory mycosis fungoides and Sézary syndrome. Blood 123, 1159–1166 (2014).
List, A. et al. Efficacy of lenalidomide in myelodysplastic syndromes. N. Engl. J. Med. 352, 549–557 (2005).
List, A., Ebert, B. L. & Fenaux, P. A decade of progress in myelodysplastic syndrome with chromosome 5q deletion. Leukemia 32, 1493–1499 (2018).
Schneider, R. K. et al. Role of casein kinase 1A1 in the biology and targeted therapy of del(5q) MDS. Cancer Cell 26, 509–520 (2014).
Martinez-Høyer, S. et al. Loss of lenalidomide-induced megakaryocytic differentiation leads to therapy resistance in del(5q) myelodysplastic syndrome. Nat. Cell Biol. 22, 526–533 (2020).
Jädersten, M. et al. TP53 mutations in low-risk myelodysplastic syndromes with del(5q) predict disease progression. J. Clin. Oncol. 29, 1971–1979 (2011).
Sekeres, M. A. et al. A phase 2 study of lenalidomide monotherapy in patients with deletion 5q acute myeloid leukemia: Southwest Oncology Group Study S0605. Blood 118, 523–528 (2011).
Boutboul, D. et al. Dominant-negative IKZF1 mutations cause a T, B, and myeloid cell combined immunodeficiency. J. Clin. Invest. 128, 3071–3087 (2018).
Fang, J. et al. A calcium- and calpain-dependent pathway determines the response to lenalidomide in myelodysplastic syndromes. Nat. Med. 22, 727–734 (2016).
Fehniger, T. A. et al. A phase 2 study of high-dose lenalidomide as initial therapy for older patients with acute myeloid leukemia. Blood 117, 1828–1833 (2010).
Blum, W. et al. Dose escalation of lenalidomide in relapsed or refractory acute leukemias. J. Clin. Oncol. 28, 4919–4925 (2010).
DeAngelo, D. J. et al. A phase I study of lenalidomide plus chemotherapy with mitoxantrone, etoposide, and cytarabine for the reinduction of patients with acute myeloid leukemia. Am. J. Hematol. 93, 254–261 (2017).
Greenberg, P. L. et al. Mitoxantrone, etoposide, and cytarabine with or without valspodar in patients with relapsed or refractory acute myeloid leukemia and high-risk myelodysplastic syndrome: a phase III trial (E2995). J. Clin. Oncol. 22, 1078–1086 (2004).
Feldman, E. J. et al. Phase III randomized multicenter study of a humanized anti-CD33 monoclonal antibody, lintuzumab, in combination with chemotherapy, versus chemotherapy alone in patients with refractory or first-relapsed acute myeloid leukemia. J. Clin. Oncol. 23, 4110–4116 (2005).
Narayan, R. et al. Sequential azacitidine plus lenalidomide in previously treated elderly patients with acute myeloid leukemia and higher risk myelodysplastic syndrome. Leukemia Lymphoma 57, 609–615 (2015).
Craddock, C. et al. combination lenalidomide and azacitidine: a novel salvage therapy in patients who relapse after allogeneic stem-cell transplantation for acute myeloid leukemia. J. Clin. Oncol. 37, 580–588 (2019).
Chihara, D. et al. Long-term results of a phase II trial of lenalidomide plus prednisone therapy for patients with myelofibrosis. Leukemia Res. 48, 1–5 (2016).
Bhutani, M., Polizzotto, M. N., Uldrick, T. S. & Yarchoan, R. Kaposi sarcoma–associated herpesvirus-associated malignancies: epidemiology, pathogenesis, and advances in treatment. Semin. Oncol. 42, 223–246 (2015).
Shimada, K., Hayakawa, F. & Kiyoi, H. Biology and management of primary effusion lymphoma. Blood 132, 1879–1888 (2018).
Zhang, L. et al. Phase 2 study using oral thalidomide-cyclophosphamide-prednisone for idiopathic multicentric Castleman disease. Blood 133, 1720–1728 (2019).
Sperling, A. S. et al. Patterns of substrate affinity, competition, and degradation kinetics underlie biological activity of thalidomide analogs. Blood 134, 160–170 (2019).
Rasco, D. W. et al. A first-in-human study of novel cereblon modulator avadomide (CC-122) in advanced malignancies. Clin. Cancer Res. 25, 90–98 (2018).
Matyskiela, M. E. et al. A cereblon modulator (CC-220) with improved degradation of ikaros and aiolos. J. Med. Chem. 61, 535–542 (2017).
Bjorklund, C. C. et al. Iberdomide (CC-220) is a potent cereblon E3 ligase modulator with antitumor and immunostimulatory activities in lenalidomide- and pomalidomide-resistant multiple myeloma cells with dysregulated CRBN. Leukemia 34, 1197–1201 (2019).
Sievers, Q. L. et al. Defining the human C2H2 zinc finger degrome targeted by thalidomide analogs through CRBN. Science 362, eaat0572 (2018).
Lonial, S. et al. First clinical (phase 1b/2a) study of iberdomide (CC-220; IBER), a CELMoD, in combination with dexamethasone (DEX) in patients (pts) with relapsed/refractory multiple myeloma (RRMM). J. Clin. Oncol. 37, 8006–8006 (2019).
Schafer, P. H. et al. Cereblon modulator iberdomide induces degradation of the transcription factors Ikaros and Aiolos: immunomodulation in healthy volunteers and relevance to systemic lupus erythematosus. Ann. Rheum. Dis. 77, 1516–1523 (2018).
Lopez-Girona, A. et al. CC-92480 is a novel cereblon E3 ligase modulator with enhanced tumoricidal and immunomodulatory activity against sensitive and resistant multiple myeloma cells. Blood 134, 1812–1812 (2019).
Matyskiela, M. E. et al. A novel cereblon modulator recruits GSPT1 to the CRL4CRBN ubiquitin ligase. Nature 535, 252–257 (2016).
Uy, G. L. et al. Clinical activity of CC-90009, a cereblon E3 ligase modulator and first-in-class GSPT1 degrader, as a single agent in patients with relapsed or refractory acute myeloid leukemia (R/R AML): first results from a phase I dose-finding study. Blood 134, 232–232 (2019).
Kumar, S. et al. Impact of lenalidomide therapy on stem cell mobilization and engraftment post-peripheral blood stem cell transplantation in patients with newly diagnosed myeloma. Leukemia 21, 2035–2042 (2007).
Mazumder, A. et al. Effect of lenalidomide therapy on mobilization of peripheral blood stem cells in previously untreated multiple myeloma patients. Leukemia 22, 1280–1281 (2007).
Micallef, I. N. M. et al. Plerixafor (Mozobil) for stem cell mobilization in patients with multiple myeloma previously treated with lenalidomide. Bone Marrow Transpl. 46, 350–355 (2011).
Musto, P. et al. Second primary malignancies in multiple myeloma: an overview and IMWG consensus. Ann. Oncol. 28, 228–245 (2017).
Jones, J. R. et al. Second malignancies in the context of lenalidomide treatment: an analysis of 2732 myeloma patients enrolled to the Myeloma XI trial. Blood Cancer J. 6, e506–e506 (2016).
Carrier, M., Gal, G. L., Tay, J., Wu, C. & Lee, A. Y. Rates of venous thromboembolism in multiple myeloma patients undergoing immunomodulatory therapy with thalidomide or lenalidomide: a systematic review and meta-analysis: venous thromboembolism in multiple myeloma. J. Thromb. Haemost. 9, 653–663 (2011).
Chen, C. et al. Expanded safety experience with lenalidomide plus dexamethasone in relapsed or refractory multiple myeloma. Br. J. Haematol. 146, 164–170 (2009).
Knight, R., DeLap, R. J. & Zeldis, J. B. Lenalidomide and venous thrombosis in multiple myeloma. N. Engl. J. Med. 354, 2079–2080 (2006).
Scarpace, S. et al. Arterial thrombosis in four patients treated with thalidomide. Leukemia Lymphoma 46, 239–242 (2005).
Palumbo, A. et al. Prevention of thalidomide- and lenalidomide-associated thrombosis in myeloma. Leukemia 22, 414–423 (2008).
Kaushal, V., Kaushal, G. P., Melkaveri, S. N. & Mehta, P. Thalidomide protects endothelial cells from doxorubicin-induced apoptosis but alters cell morphology. J. Thromb. Haemost. 2, 327–334 (2004).
Nardone, B. et al. Risk of rash associated with lenalidomide in cancer patients: a systematic review of the literature and meta-analysis. Clin. Lymphoma Myeloma Leuk. 13, 424–429 (2013).
Tinsley, S. M., Kurtin, S. E. & Ridgeway, J. A. Practical management of lenalidomide-related rash. Clin. Lymphoma Myeloma Leuk. 15, S64–S69 (2015).
Lee, M. J. et al. Lenalidomide desensitization for delayed hypersensitivity reactions in 5 patients with multiple myeloma. Br. J. Haematol. 167, 127–131 (2014).
FDA. Drugs @ FDA: FDA-approved drugs. US Food & Drug Administration https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&varApplNo=021880 (2019).
Pawlyn, C. et al. Lenalidomide-induced diarrhea in patients with myeloma is caused by bile acid malabsorption that responds to treatment. Blood 124, 2467–2468 (2014).
Fisher, S. L. & Phillips, A. J. Targeted protein degradation and the enzymology of degraders. Curr. Opin. Chem. Biol. 44, 47–55 (2018).
Lai, A. C. & Crews, C. M. Induced protein degradation: an emerging drug discovery paradigm. Nat. Rev. Drug Discov. 16, 101–114 (2016).
Burslem, G. M. & Crews, C. M. Proteolysis-targeting chimeras as therapeutics and tools for biological discovery. Cell 181, 102–114 (2020).
Bondeson, D. P. et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat. Chem. Biol. 11, 611–617 (2015).
Zhu, Y. X. et al. Cereblon expression is required for the antimyeloma activity of lenalidomide and pomalidomide. Blood 118, 4771–4779 (2011).
Sievers, Q. L., Gasser, J. A., Cowley, G. S., Fischer, E. S. & Ebert, B. L. Genome-wide screen identifies cullin-RING ligase machinery required for lenalidomide-dependent CRL4CRBN activity. Blood 132, 1293–1303 (2018).
Patil, A., Manzano, M. & Gottwein, E. Genome-wide CRISPR screens reveal genetic mediators of cereblon modulator toxicity in primary effusion lymphoma. Blood Adv. 3, 2105–2117 (2019).
Tateno, S. et al. Genome-wide screening reveals a role for subcellular localization of CRBN in the anti-myeloma activity of pomalidomide. Sci. Rep. 10, 4012 (2020).
Kortüm, K. M. et al. Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes. Blood 128, 1226–1233 (2016).
Thakurta, A. et al. Absence of mutations in cereblon (CRBN) and DNA damage-binding protein 1 (DDB1) genes and significance for IMiD therapy. Leukemia 28, 1129–1131 (2013).
Gooding, S. et al. Multiple cereblon genetic changes associate with acquired resistance to lenalidomide or pomalidomide in multiple myeloma. Blood https://doi.org/10.1182/blood.2020007081 (2020).
Huang, S.-Y. et al. Expression of cereblon protein assessed by immunohistochemicalstaining in myeloma cells is associated with superior response of thalidomide- and lenalidomide-based treatment, but not bortezomib-based treatment, in patients with multiple myeloma. Ann. Hematol. 93, 1371–1380 (2014).
Franssen, L. E. et al. Cereblon loss and up-regulation of c-Myc are associated with lenalidomide resistance in multiple myeloma patients. Haematologica 103, e368–e371 (2018).
Heintel, D. et al. High expression of cereblon (CRBN) is associated with improved clinical response in patients with multiple myeloma treated with lenalidomide and dexamethasone. Br. J. Haematol. 161, 695–700 (2013).
Jonasova, A. et al. High level of full-length cereblon mRNA in lower risk myelodysplastic syndrome with isolated 5q deletion is implicated in the efficacy of lenalidomide. Eur. J. Haematol. 95, 27–34 (2014).
Schuster, S. R. et al. The clinical significance of cereblon expression in multiple myeloma. Leukemia Res. 38, 23–28 (2013).
Pourabdollah, M. et al. High IKZF1/3 protein expression is a favorable prognostic factor for survival of relapsed/refractory multiple myeloma patients treated with lenalidomide. J. Hematol. Oncol. 9, 123 (2016).
Dimopoulos, K. et al. Expression of CRBN, IKZF1, and IKZF3 does not predict lenalidomide sensitivity and mutations in the cereblon pathway are infrequent in multiple myeloma. Leukemia Lymphoma 60, 1–9 (2018).
Gandhi, A. K. et al. Measuring cereblon as a biomarker of response or resistance to lenalidomide and pomalidomide requires use of standardized reagents and understanding of gene complexity. Br. J. Haematol. 164, 233–244 (2013).
Zhu, Y. X., Kortuem, K. M. & Stewart, A. K. Molecular mechanism of action of immune-modulatory drugs thalidomide, lenalidomide and pomalidomide in multiple myeloma. Leukemia Lymphoma 54, 683–687 (2012).
Krönke, J. et al. IKZF1 expression is a prognostic marker in newly diagnosed standard-risk multiple myeloma treated with lenalidomide and intensive chemotherapy: a study of the German Myeloma Study Group (DSMM). Leukemia 31, 1363–1367 (2017).
Reichermeier, K. M. et al. PIKES analysis reveals response to degraders and key regulatory mechanisms of the CRL4 network. Mol. Cell 77, 1092–1106.e9 (2020).
Järås, M. et al. Csnk1a1 inhibition has p53-dependent therapeutic efficacy in acute myeloid leukemia. J. Exp. Med. 211, 605–612 (2014).
Scharenberg, C. et al. Progression in patients with low- and intermediate-1-risk del(5q) myelodysplastic syndromes is predicted by a limited subset of mutations. Haematologica 102, 498–508 (2016).
Lodé, L. et al. Emergence and evolution of TP53 mutations are key features of disease progression in myelodysplastic patients with lower-risk del(5q) treated with lenalidomide. Haematologica 103, e143–e146 (2017).
Hoofnagle, A. N., Becker, J. O., Wener, M. H. & Heinecke, J. W. Quantification of thyroglobulin, a low-abundance serum protein, by immunoaffinity peptide enrichment and tandem mass spectrometry. Clin. Chem. 54, 1796–1804 (2008).
Zhang, B. et al. Clinical potential of mass spectrometry-based proteogenomics. Nat. Rev. Clin. Oncol. 16, 256–268 (2019).
Donovan, K. A. et al. Thalidomide promotes degradation of SALL4, a transcription factor implicated in duane radial ray syndrome. eLife 7, e38430 (2018).
An, J. et al. pSILAC mass spectrometry reveals ZFP91 as IMiD-dependent substrate of the CRL4CRBN ubiquitin ligase. Nat. Commun. 8, 15398 (2017).
Petzold, G., Fischer, E. S. & Thomä, N. H. Structural basis of lenalidomide-induced CK1α degradation by the CRL4CRBN ubiquitin ligase. Nature 532, 127–130 (2016).
Najafabadi, H. S. et al. C2H2 zinc finger proteins greatly expand the human regulatory lexicon. Nat. Biotechnol. 33, 555–562 (2015).
Higgins, J. J., Pucilowska, J., Lombardi, R. Q. & Rooney, J. P. A mutation in a novel ATP-dependent Lon protease gene in a kindred with mild mental retardation. Neurology 63, 1927–1931 (2004).
Eichner, R. et al. Immunomodulatory drugs disrupt the cereblon–CD147–MCT1 axis to exert antitumor activity and teratogenicity. Nat. Med. 22, 735–743 (2016).
Lee, K. M. et al. Disruption of the cereblon gene enhances hepatic AMPK activity and prevents high-fat diet-induced obesity and insulin resistance in mice. Diabetes 62, 1855–1864 (2013).
Nguyen, T. V. et al. Glutamine triggers acetylation-dependent degradation of glutamine synthetase via the thalidomide receptor cereblon. Mol. Cell 61, 809–820 (2016).
Mita, M. et al. Phase I study of E7820, an oral inhibitor of integrin α-2 expression with antiangiogenic properties, in patients with advanced malignancies. Clin. Cancer Res. 17, 193–200 (2011).
Simon, G. R. et al. A phase I study of tasisulam sodium (LY573636 sodium), a novel anticancer compound in patients with refractory solid tumors. Cancer Chemother. Pharmacol. 68, 1233–1241 (2011).
Rigas, J. R. et al. Phase I clinical and pharmacological study of chloroquinoxaline sulfonamide. Cancer Res. 52, 6619–6623 (1992).
Punt, C. J. A. et al. Phase I and pharmacokinetic study of E7070, a novel sulfonamide, given at a daily times five schedule in patients with solid tumors. A study by the EORTC-Early Clinical Studies Group (ECSG). Ann. Oncol. 12, 1289–1293 (2001).
Han, T. et al. Anticancer sulfonamides target splicing by inducing RBM39 degradation via recruitment to DCAF15. Science 356, eaal3755 (2017).
Uehara, T. et al. Selective degradation of splicing factor CAPERα by anticancer sulfonamides. Nat. Chem. Biol. 13, 675–680 (2017).
Ting, T. C. et al. Aryl sulfonamides degrade RBM39 and RBM23 by recruitment to CRL4-DCAF15. Cell Rep. 29, 1499–1510.e6 (2019).
Faust, T. B. et al. Structural complementarity facilitates E7820-mediated degradation of RBM39 by DCAF15. Nat. Chem. Biol. 16, 7–14 (2020).
Du, X. et al. Structural basis and kinetic pathway of RBM39 recruitment to DCAF15 by a sulfonamide molecular glue E7820. Structure 27, 1625–1633.e3 (2019).
Słabicki, M. et al. The CDK inhibitor CR8 acts as a molecular glue degrader that depletes cyclin K. Nature 585, 293–297 (2020).
Mayor-Ruiz, C. et al. Rational discovery of molecular glue degraders via scalable chemical profiling. Nat. Chem. Biol. 16, 1199–1207 (2020).
Lv, L. et al. Discovery of a molecular glue promoting CDK12-DDB1 interaction to trigger cyclin K degradation. eLife 9, e59994 (2020).
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Simonetta, K. R. et al. Prospective discovery of small molecule enhancers of an E3 ligase-substrate interaction. Nat. Commun. 10, 1402 (2019).
Kerres, N. et al. Chemically induced degradation of the oncogenic transcription factor BCL6. Cell Rep. 20, 2860–2875 (2017).
Słabicki, M. et al. Small-molecule-induced polymerization triggers degradation of BCL6. Nature 588, 164–168 (2020).
Weber, E. W., Maus, M. V. & Mackall, C. L. The emerging landscape of immune cell therapies. Cell 181, 46–62 (2020).
Neelapu, S. S. et al. Chimeric antigen receptor T-cell therapy - assessment and management of toxicities. Nat. Rev. Clin. Oncol. 15, 47–62 (2017).
Roybal, K. T. & Lim, W. A. Synthetic immunology: hacking immune cells to expand their therapeutic capabilities. Annu. Rev. Immunol. 35, 229–253 (2017).
Stasi, A. D. et al. Inducible apoptosis as a safety switch for adoptive cell therapy. N. Engl. J. Med. 365, 1673–1683 (2011).
Chiocca, E. A. et al. Regulatable interleukin-12 gene therapy in patients with recurrent high-grade glioma: results of a phase 1 trial. Sci. Transl Med. 11, eaaw5680 (2019).
Verma, R., Mohl, D. & Deshaies, R. J. Harnessing the power of proteolysis for targeted protein inactivation. Mol. Cell 77, 446–460 (2020).
Banaszynski, L. A., Chen, L., Maynard-Smith, L. A., Ooi, A. G. L. & Wandless, T. J. A rapid, reversible, and tunable method to regulate protein function in living cells using synthetic small molecules. Cell 126, 995–1004 (2006).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Chung, H. K. et al. Tunable and reversible drug control of protein production via a self-excising degron. Nat. Chem. Biol. 11, 713–720 (2015).
Neklesa, T. K. et al. Small-molecule hydrophobic tagging–induced degradation of HaloTag fusion proteins. Nat. Chem. Biol. 7, 538–543 (2011).
Buckley, D. L. et al. HaloPROTACS: use of small molecule protacs to induce degradation of halotag fusion proteins. ACS Chem. Biol. 10, 1831–1837 (2015).
Nabet, B. et al. Rapid and direct control of target protein levels with VHL-recruiting dTAG molecules. Preprint at bioRxiv https://doi.org/10.1101/2020.03.13.980946 (2020).
Nabet, B. et al. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 14, 431–441 (2018).
Clackson, T. et al. Redesigning an FKBP-ligand interface to generate chemical dimerizers with novel specificity. Proc. Natl Acad. Sci. USA 95, 10437–10442 (1998).
Lee, S. M. et al. A chemical switch system to modulate chimeric antigen receptor T cell activity through proteolysis-targeting chimaera technology. ACS Synth. Biol. 9, 987–992 (2020).
Weber, E. W. et al. Transient “rest” induces functional reinvigoration and epigenetic remodeling in exhausted CAR-T cells. Preprint at bioRxiv https://doi.org/10.1101/2020.01.26.920496 (2020).
Koduri, V. et al. Peptidic degron for IMiD-induced degradation of heterologous proteins. Proc. Natl Acad. Sci. USA 116, 2539–2544 (2019).
Carbonneau, S. et al. An IMiD-inducible degron provides reversible regulation for chimeric antigen receptor expression and activity. Cell Chem. Biol. https://doi.org/10.1016/j.chembiol.2020.11.012 (2020).
Jan, M. et al. Reversible ON- and OFF-switch chimeric antigen receptors controlled by lenalidomide. Sci. Transl Med. 13, eabb6295 (2021).
Colombo, M. P. & Trinchieri, G. Interleukin-12 in anti-tumor immunity and immunotherapy. Cytokine Growth Factor Rev. 13, 155–168 (2002).
Moreau, P. et al. Bortezomib, thalidomide, and dexamethasone with or without daratumumab before and after autologous stem-cell transplantation for newly diagnosed multiple myeloma (CASSIOPEIA): a randomised, open-label, phase 3 study. Lancet 394, 29–38 (2019).
Attal, M. et al. Isatuximab plus pomalidomide and low-dose dexamethasone versus pomalidomide and low-dose dexamethasone in patients with relapsed and refractory multiple myeloma (ICARIA-MM): a randomised, multicentre, open-label, phase 3 study. Lancet 394, 2096–2107 (2019).
Varshavsky, A. The ubiquitin system, an immense realm. Biochemistry 81, 167–176 (2012).
Zheng, N. & Shabek, N. Ubiquitin ligases: structure, function, and regulation. Annu. Rev. Biochem. 86, 129–157 (2017).
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Martinez-Høyer, S. & Karsan, A. Mechanisms of lenalidomide sensitivity and resistance. Exp. Hematol. 91, 22–31 (2020).
Cortes, M., Wong, E., Koipally, J. & Georgopoulos, K. Control of lymphocyte development by the Ikaros gene family. Curr. Opin. Immunol. 11, 167–171 (1999).
Cortés, M. & Georgopoulos, K. Aiolos is required for the generation of high affinity bone marrow plasma cells responsible for long-term immunity. J. Exp. Med. 199, 209–219 (2004).
Shaffer, A. L. et al. IRF4 addiction in multiple myeloma. Nature 454, 226–231 (2008).
Gandhi, R. et al. Activation of the aryl hydrocarbon receptor induces human type 1 regulatory T cell–like and Foxp3+ regulatory T cells. Nat. Immunol. 11, 846–853 (2010).
Quintana, F. J. et al. Aiolos promotes TH17 differentiation by directly silencing Il2 expression. Nat. Immunol. 13, 770–777 (2012).
Elyada, E. et al. CKIα ablation highlights a critical role for p53 in invasiveness control. Nature 470, 409–413 (2011).
Nijhawan, D. et al. Cancer vulnerabilities unveiled by genomic loss. Cell 150, 842–854 (2012).
Hoshino, S. et al. A human homologue of the yeast GST1 gene codes for a GTP-binding protein and is expressed in a proliferation-dependent manner in mammalian cells. EMBO J. 8, 3807–3814 (1989).
Zhouravleva, G. et al. Termination of translation in eukaryotes is governed by two interacting polypeptide chain release factors, eRF1 and eRF3. EMBO J. 14, 4065–4072 (1995).
Matyskiela, M. E. et al. SALL4 mediates teratogenicity as a thalidomide-dependent cereblon substrate. Nat. Chem. Biol. 14, 981–987 (2018).
Knobloch, J. & Rüther, U. Shedding light on an old mystery: Thalidomide suppresses survival pathways to induce limb defects. Cell Cycle 7, 1121–1127 (2008).
Coleman, K. G. & Crews, C. M. Proteolysis-targeting chimeras: harnessing the ubiquitin-proteasome system to induce degradation of specific target proteins. Annu. Rev. Cancer Biol. 2, 41–58 (2018).
Gerry, C. J. & Schreiber, S. L. Unifying principles of bifunctional, proximity-inducing small molecules. Nat. Chem. Biol. 16, 369–378 (2020).
Stanton, B. Z., Chory, E. J. & Crabtree, G. R. Chemically induced proximity in biology and medicine. Science 359, eaao5902 (2018).
Nowak, R. P. et al. Plasticity in binding confers selectivity in ligand-induced protein degradation. Nat. Chem. Biol. 14, 706–714 (2018).
Huang, H.-T. et al. A chemoproteomic approach to query the degradable kinome using a multi-kinase degrader. Cell Chem. Biol. 25, 88–99.e6 (2018).
Bondeson, D. P. et al. Lessons in PROTAC design from selective degradation with a promiscuous warhead. Cell Chem. Biol. 25, 78–87.e5 (2018).
Dobrovolsky, D. et al. Bruton tyrosine kinase degradation as a therapeutic strategy for cancer. Blood 133, 952–961 (2019).
Buhimschi, A. D. et al. Targeting the C481S ibrutinib-resistance mutation in Bruton’s tyrosine kinase using PROTAC-mediated degradation. Biochemistry 57, 3564–3575 (2018).
Schapira, M., Calabrese, M. F., Bullock, A. N. & Crews, C. M. Targeted protein degradation: expanding the toolbox. Nat. Rev. Drug Discov. 18, 949–963 (2019).
Yamazoe, S. et al. Heterobifunctional molecules induce dephosphorylation of kinases–a proof of concept study. J. Med. Chem. 63, 2807–2813 (2019).
Banik, S. M. et al. Lysosome-targeting chimaeras for degradation of extracellular proteins. Nature 584, 291–297 (2020).
Neklesa, T. K. & Crews, C. M. Greasy tags for protein removal. Nature 487, 308–309 (2012).
Ma, A. et al. Discovery of a first-in-class EZH2 selective degrader. Nat. Chem. Biol. 16, 214–222 (2019).
Acknowledgements
The work of M.J. is supported by a NIH T32 grant to the Massachusetts General Hospital Department of Pathology (NIH-5T32CA009216-39). The work of A.S.S. is supported by NIH grant K08CA252174. The work of B.L.E is supported by NIH grants R01HL082945 and P01CA108631, the Howard Hughes Medical Institute, the Edward P. Evans Foundation, the Leukemia and Lymphoma Society and the Adelson Medical Research Foundation.
Author information
Authors and Affiliations
Contributions
M.J. and A.S.S. researched data for article. All authors contributed substantially to discussions of content and wrote, reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
B.L.E. has received research funding from Celgene, Deerfield and Novartis, and consulting fees from GRAIL. He serves on the scientific advisory boards and holds equity in Exo Therapeutics, Neomorph Therapeutics and Skyhawk Therapeutics. M.J. and A.S.S. declare no competing interests.
Additional information
Peer review information
Nature Reviews Clinical Oncology thanks Rajesh Chopra, Ze’ev Ronai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Molecular glue
-
A small molecule that directly bridges protein–protein interactions.
- Neosubstrate
-
A protein that is conditionally targeted to a ubiquitin ligase in the presence of a small-molecule drug.
- High-risk cytogenetic features
-
Chromosomal alterations that are associated with aggressive disease and unfavourable survival outcomes; in myeloma, these features include t(4;14), t(14;16) and del(17p) alterations.
- Antibody-dependent cellular cytotoxicity
-
(ADCC). A mechanism of cell killing that requires recognition of the antibody-bound cell by immune effector cells, such as natural killer cells.
- Activated B cell-like DLBCL
-
A subtype of diffuse large B cell lymphoma (DLBCL) that is defined by a specific transcriptional signature and is associated with high risk of relapse following standard rituximab, cyclophosphamide, doxorubicin, vincristine and prednisone (R-CHOP) chemoimmunotherapy.
Rights and permissions
About this article
Cite this article
Jan, M., Sperling, A.S. & Ebert, B.L. Cancer therapies based on targeted protein degradation — lessons learned with lenalidomide. Nat Rev Clin Oncol 18, 401–417 (2021). https://doi.org/10.1038/s41571-021-00479-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-021-00479-z
This article is cited by
-
NF-κB in biology and targeted therapy: new insights and translational implications
Signal Transduction and Targeted Therapy (2024)
-
Chemical genetic control of cytokine signaling in CAR-T cells using lenalidomide-controlled membrane-bound degradable IL-7
Leukemia (2024)
-
Unraveling the diversity of molecular glue degraders
Nature Chemical Biology (2024)
-
The progress of novel strategies on immune-based therapy in relapsed or refractory diffuse large B-cell lymphoma
Experimental Hematology & Oncology (2023)
-
Protein degradation: expanding the toolbox to restrain cancer drug resistance
Journal of Hematology & Oncology (2023)