Abstract
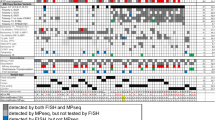
Deletions of the long arm of chromosome 13 (13q−) are observed in patients with multiple myeloma (MM), are rarely observed in the monoclonal gammopathy of undetermined significance (MGUS) and have been associated with a worsened prognosis in MM. However, no minimally deleted region in the13q arm has been defined at 13q, and consequently no tumor suppressor genes have yet been identified that are important for disease pathogenesis. We attempted to characterize these chromosome 13q deletions at the molecular cytogenetic level. We studied 351 newly diagnosed patients, entered into the E9486/E9487 clinical study of the Eastern Cooperative Oncology Group. Fluorescent in situ hybridization (FISH) combined with immune fluorescent detection (cIg-FISH) of clonal plasma cells (PC) and cytomorphology were used to analyze interphase, bone marrow (BM) cell, cytospin slides. We simultaneously used DNA probes for the following locus specific probes (LSI); LSI 13 (Rb) and D13S319, which hybridize to 13q14. We subsequently studied distal deletions using the D13S25 probe (13q14.3) and a subtelomeric probe (13qSTP) for the 13q-arm (D13S327) in 40 cases with documented LSI 13 (Rb)/D13S319 deletion and 40 without deletion of these loci. Of 325 evaluable patients, we found 13q deletions in 176 (54%) using LSI 13 (Rb) and D13S319 probes. Of 40 patients with LSI 13 (Rb)/D13S319 deletions, 34 (85%) had coexistent deletion of both D13S25/13qSTP. These results indicate that chromosome 13 deletions in MM involve loss of most if not all of the 13q arm perhaps even indicating monosomy. In six cases the 13qSTP signal was conserved, but D13S25 was lost indicating large interstitial deletions involving 13q14. In 39 of the 40 cases without LSI 13 (Rb)/D13S319 deletions, the normal pattern of two pairs of signals was observed for D13S25/13qSTP. Deletions involving 13q14 are very common in MM as detected by cIg-FISH. These deletions appear to predominantly involve loss of large segments of the 13q arm or monosomy 13, and only occasionally represent an interstitial deletion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dewald GW, Kyle RA, Hicks GA, Greipp PR . The clinical significance of cytogenetic studies in 100 patients with multiple myeloma, plasma cell leukemia, or amyloidosis Blood 1985 66: 380–390
Sawyer JR, Waldron JA, Jagannath S, Barlogie B . Cytogenetic findings in 200 patients with multiple myeloma Cancer Genet Cytogenet 1995 82: 41–49
Smadja NV, Louvet C, Isnard F et al. Cytogenetic study in multiple myeloma at diagnosis: comparison of two techniques Br J Haematol 1995 90: 619–624
Avet-Loiseau H, Andree-Ashley LE, Moore D 2nd et al. Molecular cytogenetic abnormalities in multiple myeloma and plasma cell leukemia measured using comparative genomic hybridization Genes Chromos Cancer 1997 19: 124–133
Cigudosa JC, Rao PH, Calasanz MJ et al. Characterization of nonrandom chromosomal gains and losses in multiple myeloma by comparative genomic hybridization Blood 1998 91: 3007–3010
Rao PH, Cigudosa JC, Ning Y et al. Multicolor spectral karyotyping identifies new recurring breakpoints and translocations in multiple myeloma Blood 1998 92: 1743–1748
Sawyer JR, Lukacs JL, Munshi N et al. Identification of new nonrandom translocations in multiple myeloma with multicolor spectral karyotyping Blood 1998 92: 4269–4278
Tricot G, Barlogie B, Jagannath S et al. Poor prognosis in multiple myeloma is associated only with partial or complete deletions of chromosome 13 or abnormalities involving 11q and not with other karyotype abnormalities Blood 1995 86: 4250–4256
Tricot G, Sawyer JR, Jagannath S et al. Unique role of cytogenetics in the prognosis of patients with myeloma receiving high-dose therapy and autotransplants J Clin Oncol 1997 15: 2659–2666
Konigsberg R, Zojer N, Ackermann J et al. Predictive role of interphase cytogenetics for survival of patients with multiple myeloma J Clin Oncol 2000 18: 804–812
Zojer N, Konigsberg R, Ackermann J et al. Deletion of 13q14 remains an independent adverse prognostic variable in multiple myeloma despite its frequent detection by interphase fluorescence in situ hybridization Blood 2000 95: 1925–1930
Juneau AL, Kaehler M, Christensen ER et al. Detection of RB1 deletions by fluorescence in situ hybridization in malignant hematologic disorders Cancer Genet Cytogenet 1998 103: 117–123
Avet-Loiseau H, Li JY, Morineau N et al. Monosomy 13 is associated with the transition of monoclonal gammopathy of undetermined significance to multiple myeloma. Intergroupe Francophone du Myelome Blood 1999 94: 2583–2589
Shaughnessy J, Tian E, Sawyer J et al. High incidence of chromosome 13 deletion in multiple myeloma detected by multiprobe interphase FISH Blood 2000 96: 1505–1511
Oken MM, Leong T, Lenhard RE Jr et al. The addition of interferon or high dose cyclophosphamide to standard chemotherapy in the treatment of patients with multiple myeloma: phase III Eastern Cooperative Oncology Group Clinical Trial EST 9486 Cancer 1999 86: 957–968
Ahmann GJ, Jalal SM, Juneau AL et al. A novel three-color, clone-specific fluorescence in situ hybridization procedure for monoclonal gammopathies Cancer Genet Cytogenet 1998 101: 7–11
Kalachikov S, Migliazza A, Cayanis E et al. Cloning and gene mapping of the chromosome 13q14 region deleted in chronic lymphocytic leukemia Genomics 1997 42: 369–377
Avet-Loiseau H, Facon T, Daviet A et al. 14q32 translocations and monosomy 13 observed in monoclonal gammopathy of undetermined significance delineate a multistep process for the oncogenesis of multiple myeloma. Intergroupe Francophone du Myelome Cancer Res 1999 59: 4546–4550
Knudson AG Jr, Strong LC, Anderson DE . Heredity and cancer in man Prog Med Genet 1973 9: 113–158
Bouyge-Moreau I, Rondeau G, Avet-Loiseau H et al. Construction of a 780-kb PAC, BAC, and cosmid contig encompassing the minimal critical deletion involved in B cell chronic lymphocytic leukemia at 13q14.3 Genomics 1997 46: 183–190
Brown AG, Ross FM, Dunne EM, Steel CM, Weir-Thompson EM . Evidence for a new tumour suppressor locus (DBM) in human B-cell neoplasia telomeric to the retinoblastoma gene Nat Genet 1993 3: 67–72
Stilgenbauer S, Nickolenko J, Wilhelm J et al. Expressed sequences as candidates for a novel tumor suppressor gene at band 13q14 in B-cell chronic lymphocytic leukemia and mantle cell lymphoma Oncogene 1998 16: 1891–1897
Corcoran MM, Rasool O, Liu Y et al. Detailed molecular delineation of 13q14.3 loss in B-cell chronic lymphocytic leukemia Blood 1998 91: 1382–1390
Garcia-Marco JA, Caldas C, Price CM, Wiedemann LM, Ashworth A, Catovsky D . Frequent somatic deletion of the 13q12.3 locus encompassing BRCA2 in chronic lymphocytic leukemia Blood 1996 88: 1568–1575
Bullrich F, Veronese ML, Kitada S et al. Minimal region of loss at 13q14 in B-cell chronic lymphocytic leukemia Blood 1996 88: 3109–3115
Dohner H, Stilgenbauer S, Fischer K, Bentz M, Lichter P . Cytogenetic and molecular cytogenetic analysis of B cell chronic lymphocytic leukemia: specific chromosome aberrations identify prognostic subgroups of patients and point to loci of candidate genes Leukemia 1997 11: S19–S24
Dierlamm J, Michaux L, Criel A, Wlodarska I, Van den Berghe H, Hossfeld DK . Genetic abnormalities in chronic lymphocytic leukemia and their clinical and prognostic implications Cancer Genet Cytogenet 1997 94: 27–35
Merup M, Jansson M, Corcoran M et al. A FISH cosmid ‘cocktail’ for detection of 13q deletions in chronic lymphocytic leukaemia – comparison with cytogenetics and Southern hybridization Leukemia 1998 12: 705–709
Dominici M, Luppi M, Campioni D et al. PCR with degenerate primers for highly conserved DNA polymerase gene of the herpesvirus family shows neither human herpesvirus 8 nor a related variant in bone marrow stromal cells from multiple myeloma patients Int J Cancer 2000 86: 76–82
Rajkumar SV, Fonseca R, Dewald GW et al. Cytogenetic abnormalities correlate with the plasma cell labeling index and extent of bone marrow involvement in myeloma Cancer Genet Cytogenet 1999 113: 73–77
Perez-Simon JA, Garcia-Sanz R, Tabernero MD et al. Prognostic value of numerical chromosome aberrations in multiple myeloma: a FISH analysis of 15 different chromosomes Blood 1998 91: 3366–3371
Fonseca R, Harrington D, Oken MM et al. Prognostic significance of 13q− deletions and the t(11;14)(q13;q32) in myeloma (MM); an interphase FISH study of 351 patients entered into the Eastern Cooperative Oncology Group E9487 Clinical Trial Blood 2000 96: 547a
Duesberg P, Rasnick D, Li R, Winters L, Rausch C, Hehlmann R . How aneuploidy may cause cancer and genetic instability Anticancer Res 1999 19: 4887–4906
Duesberg P, Rausch C, Rasnick D, Hehlmann R . Genetic instability of cancer cells is proportional to their degree of aneuploidy Proc Natl Acad Sci USA 1998 95: 13692–13697
Drach J, Ackermann J, Fritz E et al. Presence of a p53 gene deletion in patients with multiple myeloma predicts for short survival after conventional-dose chemotherapy Blood 1998 92: 802–809
Acknowledgements
This work was supported in part by Public Health Service grant R01 CA83724–01 from the National Cancer Institute. RF is a Leukemia and Lymphoma Society Translational Research Awardee. RF and PRG are also supported by the Mayo Foundation and the CI-5 Cancer Research Fund-Lilly Clinical Investigator Award of the Damon Runyon–Walter Winchell Foundation. PRG is supported in part by research grant P01 CA62242 from the National Cancer Institute. PRG and NEK are supported by the ECOG grant CA21115–25C from the National Cancer Institute.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fonseca, R., Oken, M., Harrington, D. et al. Deletions of chromosome 13 in multiple myeloma identified by interphase FISH usually denote large deletions of the q arm or monosomy. Leukemia 15, 981–986 (2001). https://doi.org/10.1038/sj.leu.2402125
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2402125
Keywords
This article is cited by
-
Characterization and application of a lactate and branched chain amino acid metabolism related gene signature in a prognosis risk model for multiple myeloma
Cancer Cell International (2023)
-
Increasing genomic discovery in newly diagnosed multiple myeloma: defining disease biology and its correlation to risk
Annals of Hematology (2022)
-
Genetic pathogenesis of immunoglobulin light chain amyloidosis: basic characteristics and clinical applications
Experimental Hematology & Oncology (2021)
-
Risk Stratification in Multiple Myeloma in Indian Settings
Indian Journal of Hematology and Blood Transfusion (2020)
-
Applications of fluorescence in situ hybridization in detection of disease biomarkers and personalized medicine
Comparative Clinical Pathology (2019)