Abstract
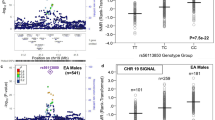
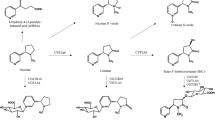
A common haplotype of the flavin-containing monooxygenase gene FMO3 is associated with aberrant mRNA splicing, a twofold reduction in in vivo nicotine N-oxidation and reduced nicotine dependence. Tobacco remains the largest cause of preventable mortality worldwide. CYP2A6, the primary hepatic nicotine metabolism gene, is robustly associated with cigarette consumption but other enzymes contribute to nicotine metabolism. We determined the effects of common variants in FMO3 on plasma levels of nicotine-N-oxide in 170 European Americans administered deuterated nicotine. The polymorphism rs2266780 (E308G) was associated with N-oxidation of both orally administered and ad libitum smoked nicotine (P⩽3.3 × 10−5 controlling for CYP2A6 genotype). In vitro, the FMO3 G308 variant was not associated with reduced activity, but rs2266780 was strongly associated with aberrant FMO3 mRNA splicing in both liver and brain (P⩽6.5 × 10−9). Surprisingly, in treatment-seeking European American smokers (n=1558) this allele was associated with reduced nicotine dependence, specifically with a longer time to first cigarette (P=9.0 × 10−4), but not with reduced cigarette consumption. As N-oxidation accounts for only a small percentage of hepatic nicotine metabolism we hypothesized that FMO3 genotype affects nicotine metabolism in the brain (unlike CYP2A6, FMO3 is expressed in human brain) or that nicotine-N-oxide itself has pharmacological activity. We demonstrate for the first time nicotine N-oxidation in human brain, mediated by FMO3 and FMO1, and show that nicotine-N-oxide modulates human α4β2 nicotinic receptor activity in vitro. These results indicate possible mechanisms for associations between FMO3 genotype and smoking behaviors, and suggest nicotine N-oxidation as a novel target to enhance smoking cessation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mokdad AH, Marks JS, Stroup DF, Gerberding JL . Actual causes of death in the United States, 2000. JAMA 2004; 291: 1238–1245.
WHO. World Health Statistics Report—2008 2008.
Prevention CfDCa. Quitting Smoking Among Adults—United States, 2001–2010 2010.
Cahill K, Stevens S, Perera R, Lancaster T . Pharmacological interventions for smoking cessation: an overview and network meta-analysis. Cochrane Database Syst Rev 2013; 5: CD009329.
Hukkanen J, Jacob P 3rd, Benowitz NL . Metabolism and disposition kinetics of nicotine. Pharmacol Rev 2005; 57: 79–115.
von Weymarn LB, Brown KM, Murphy SE . Inactivation of CYP2A6 and CYP2A13 during nicotine metabolism. J Pharmacol Exp Ther 2006; 316: 295–303.
Ho MK, Tyndale RF . Overview of the pharmacogenomics of cigarette smoking. Pharmacogenomics J 2007; 7: 81–98.
Lerman C, Jepson C, Wileyto EP, Patterson F, Schnoll R, Mroziewicz M et al. Genetic variation in nicotine metabolism predicts the efficacy of extended-duration transdermal nicotine therapy. Clin Pharmacol Ther 2010; 87: 553–557.
Schnoll RA, Patterson F, Wileyto EP, Tyndale RF, Benowitz N, Lerman C . Nicotine metabolic rate predicts successful smoking cessation with transdermal nicotine: a validation study. Pharmacol Biochem Behav 2009; 92: 6–11.
Thorgeirsson TE, Gudbjartsson DF, Surakka I, Vink JM, Amin N, Geller F et al. Sequence variants at CHRNB3-CHRNA6 and CYP2A6 affect smoking behavior. Nat Genet 2010; 42: 448–453.
TAG-Consortium. Genome-wide meta-analyses identify multiple loci associated with smoking behavior. Nat Genet 2010; 42: 441–447.
Cho MH, Castaldi PJ, Wan ES, Siedlinski M, Hersh CP, Demeo DL et al. A genome-wide association study of COPD identifies a susceptibility locus on chromosome 19q13. Hum Mol Genet 2012; 21: 947–957.
Bloom AJ, Harari O, Martinez M, Madden PA, Martin NG, Montgomery GW et al. Use of a predictive model derived from in vivo endophenotype measurements to demonstrate associations with a complex locus, CYP2A6. Hum Mol Genet 2012; 21: 3050–3062.
Saccone NL, Wang JC, Breslau N, Johnson EO, Hatsukami D, Saccone SF et al. The CHRNA5-CHRNA3-CHRNB4 nicotinic receptor subunit gene cluster affects risk for nicotine dependence in African-Americans and in European-Americans. Cancer Res 2009; 69: 6848–6856.
Thorgeirsson TE, Geller F, Sulem P, Rafnar T, Wiste A, Magnusson KP et al. A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature 2008; 452: 638–642.
Bierut LJ, Madden PA, Breslau N, Johnson EO, Hatsukami D, Pomerleau OF et al. Novel genes identified in a high-density genome wide association study for nicotine dependence. Hum Mol Genet 2007; 16: 24–35.
Berg JZ, Mason J, Boettcher AJ, Hatsukami DK, Murphy SE . Nicotine metabolism in African Americans and European Americans: variation in glucuronidation by ethnicity and UGT2B10 haplotype. J Pharmacol Exp Ther 2010; 332: 202–209.
Benowitz NL, Perez-Stable EJ, Fong I, Modin G, Herrera B, Jacob P 3rd . Ethnic differences in N-glucuronidation of nicotine and cotinine. J Pharmacol Exp Ther 1999; 291: 1196–1203.
Murphy SE, Park SS, Thompson EF, Wilkens LR, Patel Y, Stram DO et al. Nicotine N-glucuronidation relative to N-oxidation and C-oxidation and UGT2B10 genotype in five ethnic/racial groups. Carcinogenesis 2014; 35: 2526–2533.
Zhang J, Cashman JR . Quantitative analysis of FMO gene mRNA levels in human tissues. Drug Metab Dispos 2006; 34: 19–26.
Bhagwat SV, Bhamre S, Boyd MR, Ravindranath V . Cerebral metabolism of imipramine and a purified flavin-containing monooxygenase from human brain. Neuropsychopharmacology 1996; 15: 133–142.
Ferguson CS, Tyndale RF . Cytochrome P450 enzymes in the brain: emerging evidence of biological significance. Trends Pharmacol Sci 2011; 32: 708–714.
Shimizu M, Yano H, Nagashima S, Murayama N, Zhang J, Cashman JR et al. Effect of genetic variants of the human flavin-containing monooxygenase 3 on N- and S-oxygenation activities. Drug Metab Dispos 2007; 35: 328–330.
Bloom J, Hinrichs AL, Wang JC, von Weymarn LB, Kharasch ED, Bierut LJ et al. The contribution of common CYP2A6 alleles to variation in nicotine metabolism among European-Americans. Pharmacogenet Genomics 2011; 21: 403–416.
McCarthy DE, Piasecki TM, Lawrence DL, Jorenby DE, Shiffman S, Fiore MC et al. A randomized controlled clinical trial of bupropion SR and individual smoking cessation counseling. Nicotine Tob Res 2008; 10: 717–729.
Piper ME, Federman EB, McCarthy DE, Bolt DM, Smith SS, Fiore MC et al. Efficacy of bupropion alone and in combination with nicotine gum. Nicotine Tob Res 2007; 9: 947–954.
Piper ME, Smith SS, Schlam TR, Fiore MC, Jorenby DE, Fraser D et al. A randomized placebo-controlled clinical trial of 5 smoking cessation pharmacotherapies. Arch Gen Psychiatry 2009; 66: 1253–1262.
Bloom AJ, von Weymarn LB, Martinez M, Bierut LJ, Goate A, Murphy SE . The contribution of common UGT2B10 and CYP2A6 alleles to variation in nicotine glucuronidation among European Americans. Pharmacogenet Genomics 2013; 23: 706–716.
Bloom AJ, Hartz SM, Baker TB, Chen LS, Piper ME, Fox L et al. Beyond cigarettes per day. A genome-wide association study of the biomarker carbon monoxide. Ann Am Thorac Soc 2014; 11: 1003–1010.
Stephens M, Donnelly P . A comparison of bayesian methods for haplotype reconstruction from population genotype data. Am J Hum Genet 2003; 73: 1162–1169.
Bloom AJ, Harari O, Martinez M, Zhang X, McDonald SA, Murphy SE et al. A compensatory effect upon splicing results in normal function of the CYP2A6*14 allele. Pharmacogenet Genomics 2013; 23: 107–116.
Wang JC, Cruchaga C, Saccone NL, Bertelsen S, Liu P, Budde JP et al. Risk for nicotine dependence and lung cancer is conferred by mRNA expression levels and amino acid change in CHRNA5. Hum Mol Genet 2009; 18: 3125–3135.
Gadel S, Friedel C, Kharasch ED . Differences in methadone metabolism by CYP2B6 variants. Drug Metab Dispos 2015; 43: 994–1001.
Aliverti A, Curti B, Vanoni MA . Identifying and quantitating FAD and FMN in simple and in iron-sulfur-containing flavoproteins. Methods Mol Biol 1999; 131: 9–23.
Li P, Ann J, Akk G . Activation and modulation of human alpha4beta2 nicotinic acetylcholine receptors by the neonicotinoids clothianidin and imidacloprid. J Neurosci Res 2011; 89: 1295–1301.
Kang J, Chung W, Lee K, Park C, Kang J, Shin I et al. Phenotypes of flavin-containing monooxygenase activity determined by ranitidine N-oxidation are positively correlated with genotypes of linked FM03 gene mutations in a Korean population. Pharmacogenetics 2000; 10: 67–78.
Park SB, Jacob P 3rd, Benowitz NL, Cashman JR . Stereoselective metabolism of (S)-(-)-nicotine in humans: formation of trans-(S)-(-)-nicotine N-1'-oxide. Chem Res Toxicol 1993; 6: 880–888.
Lattard V, Zhang J, Cashman JR . Alternative processing events in human FMO genes. Mol Pharmacol 2004; 65: 1517–1525.
Katchamart S, Stresser DM, Dehal SS, Kupfer D, Williams DE . Concurrent flavin-containing monooxygenase down-regulation and cytochrome P-450 induction by dietary indoles in rat: implications for drug-drug interaction. Drug Metab Dispos 2000; 28: 930–936.
Hinrichs AL, Murphy SE, Wang JC, Saccone S, Saccone N, Steinbach JH et al. Common polymorphisms in FMO1 are associated with nicotine dependence. Pharmacogenet Genomics 2011; 21: 397–402.
Bloom AJ, Murphy SE, Martinez M, von Weymarn LB, Bierut LJ, Goate A . Effects upon in vivo nicotine metabolism reveal functional variation in FMO3 associated with cigarette consumption. Pharmacogenet Genomics 2013; 23: 62–68.
Chenoweth MJ, Zhu AZ, Sanderson Cox L, Ahluwalia JS, Benowitz NL, Tyndale RF . Variation in P450 oxidoreductase (POR) A503V and flavin-containing monooxygenase (FMO)-3 E158K is associated with minor alterations in nicotine metabolism, but does not alter cigarette consumption. Pharmacogenet Genomics 2014; 24: 172–176.
Haberstick BC, Timberlake D, Ehringer MA, Lessem JM, Hopfer CJ, Smolen A et al. Genes, time to first cigarette and nicotine dependence in a general population sample of young adults. Addiction 2007; 102: 655–665.
Borland R, Yong HH, O'Connor RJ, Hyland A, Thompson ME . The reliability and predictive validity of the Heaviness of Smoking Index and its two components: findings from the International Tobacco Control Four Country study. Nicotine Tob Res 2010; 12: S45–S50.
Branstetter SA, Mercincavage M, Muscat JE . Predictors of the nicotine dependence behavior time to the first cigarette in a multiracial cohort. Nicotine Tob Res 2014.
Piasecki TM, Piper ME, Baker TB . Tobacco dependence: insights from investigations of self-reported smoking motives. Curr Dir Psychol Sci 2010; 19: 395–401.
Piasecki TM, Piper ME, Baker TB . Refining the tobacco dependence phenotype using the Wisconsin Inventory of Smoking Dependence Motives: II. Evidence from a laboratory self-administration assay. J Abnorm Psychol 2010; 119: 513–523.
Bhamre S, Bhagwat SV, Shankar SK, Boyd MR, Ravindranath V . Flavin-containing monooxygenase mediated metabolism of psychoactive drugs by human brain microsomes. Brain Res 1995; 672: 276–280.
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O'Keeffe S et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci 2014; 34: 11929–11947.
Acknowledgements
We thank Dr Jeffery Milbrandt and his lab for their advice and assistance. We acknowledge the use of tissues procured by the National Disease Research Interchange (NDRI) with support from NIH grant 2 U42 OD011158. The Collaborative Genetic Study of Nicotine Dependence was supported by National Cancer Institute grant P01 CA089392. The University of Wisconsin Transdisciplinary Tobacco Use Research Center was supported by National Institute on Drug Abuse grant P50 DA019706 and National Cancer Institute grant P50 CA084724 (TBB). This work was also supported by the National Institute on Drug Abuse grants T32 DA007261 (AMT & AJB), R01 DA036583 (LJB), R01 DA14211 (EDK), R01 DA25931 (EDK) and K01 DA034035 (AJB), National Cancer Institute grant K05 CA139871 (TBB), National Institute of Heart, Lung, and Blood Disease grant R01 HL109031 (TBB), and the Taylor Family Institute for Innovative Psychiatric Research (GA). Genotyping services for the UW-TTURC sample were provided by the Center for Inherited Disease Research (CIDR) and funding support for CIDR was provided by NIH grant U01 HG004438 and NIH contract HHSN268200782096C to The Johns Hopkins University. Assistance with genotype cleaning was provided by the Gene Environment Association Studies (GENEVA) Coordinating Center (U01 HG004446). LC/MS/MS analysis carried out in the Analytical Biochemistry Core of the University of Minnesota Cancer Center, was supported in part by National Cancer Institute grant CA-77598.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
LJB and AG are listed as an inventors on Issued US Patent 8,080,371, ‘Markers for Addiction’ covering the use of certain SNPs in determining the diagnosis, prognosis and treatment of addiction.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Rights and permissions
About this article
Cite this article
Teitelbaum, A., Murphy, S., Akk, G. et al. Nicotine dependence is associated with functional variation in FMO3, an enzyme that metabolizes nicotine in the brain. Pharmacogenomics J 18, 136–143 (2018). https://doi.org/10.1038/tpj.2016.92
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2016.92