Abstract
Evasion of apoptosis is critical for tumorigenesis, and sustained survival of nascent neoplastic cells may depend upon the endogenous levels of pro-survival BCL-2 family members. Indeed, previous studies using gene-targeted mice revealed that BCL-XL, but surprisingly not BCL-2, is critical for the development of c-MYC-induced pre-B/B lymphomas. However, it remains unclear whether another pro-survival BCL-2 relative contributes to their development. MCL-1 is an intriguing candidate, because it is required for cell survival during early B-lymphocyte differentiation. It is expressed abnormally high in several types of human B-cell lymphomas and is implicated in their resistance to chemotherapy. To test the B-cell intrinsic requirement for endogenous MCL-1 in lymphoma development, we conditionally deleted Mcl-1 in B-lymphoid cells of Eμ-Myc transgenic mice. We found that MCL-1 loss in early B-lymphoid progenitors delayed MYC-driven lymphomagenesis. Moreover, the lymphomas that arose when MCL-1 levels were diminished appeared to have been selected for reduced levels of BIM and/or increased levels of BCL-XL. These results underscore the importance of MCL-1 in lymphoma development and show that alterations in the levels of other cell death regulators can compensate for deficiencies in MCL-1 expression.
Similar content being viewed by others
Main
Apoptosis is a genetically programmed process for eliminating unwanted cells and is critical for normal development and tissue homeostasis in multi-cellular organisms.1 Defects in apoptosis are implicated in several disease states, particularly cancer2 and autoimmunity.3 Proteins of the BCL-2 family are major regulators of apoptosis.4, 5 The essential triggers are its BH3-only sub-family members (for example, BIM, PUMA and NOXA), which are activated transcriptionally and/or post-transcriptionally in response to diverse intracellular stresses.6, 7 The pro-apoptotic multi-BCL-2 homology (BH) domain proteins BAX, BAK (and possibly BOK8) have the essential role of permeabilizing the mitochondrial outer membrane, which constitutes the ‘point-of-no-return’ in apoptosis signaling and unleashes the caspase cascade that mediates cell demolition.4, 5, 9 The pro-survival BCL-2 family members, including BCL-2, BCL-XL, MCL-1, BCL-W and A1/BFL1, counter the members of both these pro-apoptotic sub-families; they function in a cell type specific but frequently also overlapping manner. For example, MCL-1 is essential for early embryonic development10 and studies with conditional knockout mice revealed that it is critical for the survival of diverse cell types, including hematopoietic stem cells,11 immature as well as mature B- and T-lymphoid cells12 and certain myeloid cell populations.13
Many cancers display abnormalities in the levels of pro-survival and/or pro-apoptotic BCL-2 family members and evasion of apoptosis is widely thought to be essential to sustain the survival of nascent neoplastic cells and hence critical for tumorigenesis.14, 15 However, the mechanisms that protect cells undergoing neoplastic transformation from apoptosis remain incompletely understood.2, 16 Abnormalities in the BCL-2-governed apoptotic pathway or its regulators have been implicated in B-cell lymphoma development. For example, BCL-2 is overexpressed due to the t[14;18] chromosomal translocation in human follicular center B-cell lymphoma, whereas both alleles of BIM are frequently lost in mantle cell lymphoma.17, 18, 19, 20 Accordingly, transgenic overexpression of BCL-2 (or its relatives BCL-XL or MCL-1), or engineered loss of BIM, PUMA or BAX, can accelerate lymphomagenesis, particularly if cell cycle control is impaired, for example by enforced expression of c-MYC21, 22, 23, 24, 25 or v-Abl.26 Although lymphomas elicited by combined overexpression of c-MYC and BCL-2 are ‘addicted to’ sustained BCL-2 overexpression for continued expansion,27 endogenous BCL-2 is dispensable for c-MYC-induced lymphomagenesis.28 In contrast, BCL-XL proved essential for the survival of both normal and pre-leukemic B cells undergoing neoplastic transformation and its loss greatly impaired lymphoma development in Eμ-Myc transgenic mice.29 Notably, the impaired tumor development could be overcome by concomitant loss of pro-apoptotic BIM.30
However, it is still unclear whether BCL-XL is the sole pro-survival BCL-2 family member required for MYC-induced pre-B/B-lymphoma development. MCL-1 is of particular interest. Increases in MCL-1 gene copy number and concomitantly elevated MCL-1 protein are found in a substantial fraction of diverse cancer types.31 For a few cell lines derived from such cancers, MCL-1 knockdown by RNA interference was shown to cause apoptosis, demonstrating that MCL-1 is critical for their sustained survival.31 Similarly, acute myeloid leukemia (AML) cells driven by enforced expression of c-MYC or the MLL-ENL and MLL-AF9 fusion onco-proteins and c-MYC- or BCR-ABL-driven pre-B/B lymphomas were rapidly killed upon inducible genetic deletion or blockade of MCL-1.32, 33, 34, 35
MCL-1 is critical for the survival of rapidly proliferating hematopoietic progenitors36 and non-transformed pro-B/pre-B cells,12 the cells thought to be the origin of Eμ-Myc lymphoma.37, 38 Therefore, we examined the role of MCL-1 in pre-B/B cell lymphoma development in Eμ-Myc transgenic mice by incorporating CD19-Cre or Rag1-Cre alleles to impose Mcl-1 gene deletion exclusively in the B-lymphoid compartment. We report that there was marked selection against Mcl-1 gene loss during c-MYC-driven lymphoma development and a delay in tumor onset. Moreover, the lymphomas that arose despite successful Mcl-1fl recombination exhibited abnormally low levels of pro-apoptotic BIM and/or increased levels of pro-survival BCL-XL. These results show that MCL-1 is critical for c-MYC-driven pre-B/B-lymphoma development, and suggest that alterations in other core components of the apoptotic machinery can compensate for a reduction in MCL-1 levels.
Results
Impact of B-cell lineage-restricted deletion of Mcl-1 on MYC-driven lymphomagenesis
To explore the impact of B cell-restricted deletion of one or both allele(s) of Mcl-1 on c-MYC-driven lymphoma development, we generated Eμ-Myc mice with one or both Mcl-1 alleles flanked by loxP sites (hereafter called Mcl-1fl/+ or Mcl-1fl/fl, respectively). Some cohorts also expressed the Cre recombinase selectively either from the common lymphoid progenitor stage (CLP), using a Rag-1-Cre transgene, or from the late pro-B cell stage onwards, using a CD19-Cre transgene.39 In our Mcl-1 gene-targeted mice, recombination of the Mcl-1fl allele subjugates a human CD4 reporter transgene to the Mcl-1 promoter/enhancer elements. Hence, human CD4 (hCD4) expression, which is readily detectable by flow cytometric analysis using fluorochrome-labeled anti-human-CD4 antibodies, serves as a reporter of Mcl-1fl deletion.33, 40, 41
We first compared the incidence and rate of pre-B/B-cell lymphoma development in Eμ-Myc, Eμ-Myc;CD19-Cre, Eμ-Myc;CD19-Cre;Mcl-1fl/+ and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice (Figure 1a). The lymphoma-free survival of the control mice without Mcl-1 deletion (Eμ-Myc and Eμ-Myc;CD19-Cre) was similar: median survivals of 91 days and 117 days, respectively (Mantle–Cox Log-rank test P=0.069, Figure 1a). With one or two floxed Mcl-1 alleles, there was a modest (albeit not statistically significant) delay in lymphomagenesis compared with the Eμ-Myc;CD19-Cre animals: 130 and 123 days, respectively (P=0.16 for both).
Minor impact of CD19-Cre-mediated loss of one or both alleles of Mcl-1 in B-lymphoid cells on pre-B/B-cell lymphoma development in Eμ-Myc mice. (a) Kaplan–Meier survival curves comparing survival of Eμ-Myc (median 91 days), Eμ-Myc;CD19-Cre (117 days), Eμ-Myc;CD19-Cre;Mcl-1fl/+ (130 days) and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice (123 days). Eμ-Myc versus Eμ-Myc;CD19-Cre P=0.069; Eμ-Myc;CD19-Cre versus Eμ-Myc;CD19-Cre;Mcl-1fl/+ P=0.16; Eμ-Myc;CD19-Cre versus Eμ-Myc;CD19-Cre;Mcl-1fl/fl P=0.16). (b) Lymphoma burden in sick Eμ-Myc, Eμ-Myc;CD19-Cre, Eμ-Myc;CD19-Cre;Mcl-1fl/+and Eμ-Myc;CD19-Cre;Mcl-1fl/flmice. No significant differences were observed, compared with sick Eμ-Myc;CD19-Cre mice, in the weights of the spleen or lymph nodes, respectively. (c) Peripheral blood analysis of sick, lymphoma-burdened compound mutant mice using an ADVIA counter. Eμ-Myc;CD19-Cre versus Eμ-Myc;CD19-Cre;Mcl-1fl/+ display no significant changes; Eμ-Myc;CD19-Cre versus Eμ-Myc;CD19-Cre;Mcl-1fl/fl: P*lymphocytes=0.0172)
Autopsy on the sick, lymphoma-burdened mice revealed that the Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice (P*=0.0172) had significantly less lymphoma cells in the blood than Eμ-Myc;CD19-Cre mice, but no such drop was found for the sick Eμ-Myc;CD19-Cre;Mcl-1fl/+mice. No significant differences between the genotypes appeared for spleen and lymph node weights (Figure 1b), or the numbers of erythrocytes and thrombocytes in the blood (Figure 1c).
Selection against MCL-1 loss
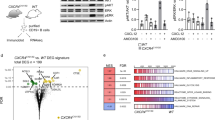
As constitutive or inducible loss of MCL-1 impairs the development as well as sustained expansion of many tumors,33, 34, 41 we tested whether the Mcl-1fl allele(s) had been recombined in the lymphomas that arose in the Eμ-Myc;CD19-Cre;Mcl-1fl/+and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice, or whether selection against Mcl-1 gene loss had occurred during their malignant transformation. Western blot analysis revealed that many lymphomas from Eμ-Myc;CD19-Cre;Mcl-1fl/+(3/3 tested) and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice (2/3 tested) retained MCL-1 expression, but its levels were significantly lower than in lymphomas from Eμ-Myc control mice (Figure 2a). The reduced MCL-1 protein expression appeared to be accompanied by a significant decrease in BIM protein expression in the Eμ-Myc;CD19-Cre;Mcl-1fl/+and Eμ-Myc;CD19-Cre;Mcl-1fl/fl lymphomas tested. Also, BCL-XL appeared to be upregulated in the Eμ-Myc;CD19-Cre;Mcl-1fl/+ lymphomas, in which Mcl-1fl deletion was efficient (Figure 2a). Compared with Eμ-Myc;CD19-Cre control lymphomas, BCL-2 protein expression was comparable in the Eμ-Myc;CD19-Cre;Mcl-1fl/+ lymphomas or, curiously, was lower in the Eμ-Myc;CD19-Cre;Mcl-1fl/fl lymphomas that had retained their Mcl-1fl alleles (Figure 2a). Consistent with the Western blot results, PCR analysis of FACS-sorted primary lymphoma cells confirmed that some of the lymphoma cells arising in Eμ-Myc;CD19-Cre;Mcl-1fl/+ and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice had not excised or only partially excised their Mcl-1fl alleles (Figure 2b).
Expression of BCL-2 family members, p53, MDM2 and p19-ARF in lymphomas from Eμ-Myc mice with CD19-Cre-mediated deletion of Mcl-1. (a) The levels of the indicated proteins were determined by Western blot analysis in three lymphomas from each of the indicated genotypes. The protein from the Mcl-1fl allele is slightly larger than the wt MCL-1 protein indicated by arrows.53 Probing for HSP70 was used as a loading control. (b) Genotype analysis of FACS-sorted B220+ pre-B/B-lymphoma cells from mice of the indicated genotypes, using primers that recognize both the Mcl-1wt and Mcl-1fl alleles, or detect the Cre recombinase transgene or the Eμ-Myc transgene. (c) Flow cytometric analysis of human CD4 reporter expression in lymphomas that arose in Eμ-Myc (negative control), Eμ-Myc;CD19-Cre;Mcl-1fl/+ or Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice
Flow cytometric analysis for the human CD4 reporter confirmed that most tumors arising in Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice had efficiently excised their Mcl-1fl allele. However, only ~50–60% of the two floxed Mcl-1 alleles had been recombined in the lymphomas from the Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice (Figure 2c). This suggests that the remaining wild-type (wt) Mcl-1 allele in Eμ-Myc;CD19-Cre;Mcl-1fl/+ B-lymphoid cells is sufficient to sustain their survival during neoplastic transformation. In contrast, deletion of both Mcl-1fl alleles must impose a stress from which lymphoma-initiating B-lymphoid progenitors are unable to recover. Thus, lymphomas that arise in Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice have potently selected against loss of both Mcl-1fl alleles and the stress caused by loss of one Mcl-1fl allele is partially relieved by adjustments in the levels of the BCL-XL and BIM proteins.
Overexpression of c-MYC causes selection bias against deletion of Mcl-1fl genes in pre-leukemic B-lymphoid cells
c-MYC promotes cell growth and cell proliferation, but under conditions of stress, such as nutrient or growth factor deprivation, high c-MYC levels predispose cells to undergo apoptosis.42, 43, 44 Pre-leukemic Eμ-Myc mice exhibit increased numbers of pre-B cells in their bone marrow, spleen, lymph nodes and blood, but these cells are not transformed and consequently do not form tumors when transplanted into congenic recipient mice.37
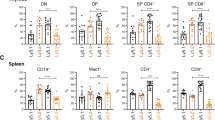
Given that loss of one allele of Mcl-1 suffices to potently induce cell death in malignant Eμ-Myc lymphomas,35 we hypothesized that loss of one Mcl-1fl allele might also reduce the numbers of pre-leukemic pro-B, pre-B and/or sIg+-B cells (at 3–4 weeks of age) in Eμ-Myc;CD19-Cre;Mcl-1fl/+mice compared with Eμ-Myc and Eμ-Myc;CD19-Cre control animals. The total bone marrow and lymph node cellularities of pre-leukemic Eμ-Myc, Eμ-Myc;CD19-Cre and Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice were comparable to each other and to wt mice, but there was a notable increase, although not statistically significant, in the overall leukocyte numbers in the spleens of Eμ-Myc and Eμ-Myc;CD19-Cre mice compared with the Eμ-Myc;CD19-Cre;Mcl-1fl/+ animals and the wt controls (Figure 3a). As reported,37 Eμ-Myc mice had more pre-B cells in their bone marrow than wt controls (Figure 3b). Interestingly, Eμ-Myc;CD19-Cre mice had significantly fewer pre-B cells than Eμ-Myc mice (Figure 3b; P*=0.0452), suggesting that the Cre recombinase imposes a cytotoxic stress on these cells. Eμ-Myc;CD19-Cre;Mcl-1fl/+ animals had even fewer pre-leukemic pre-B cells than the Eμ-Myc;CD19-Cre animals, but this difference was not statistically significant (P=0.10; Figure 3b), although the difference to the Eμ-Myc mice was significant. There were no significant differences in the numbers of pro-B cells or sIg+-B cells in the bone marrow between mice of any of the genotypes examined (Figure 3b).
Impact of c-MYC overexpression on the deletion of Mcl-1fl alleles in pre-leukemic B-lymphoid cells. (a) Total leukocyte numbers in the bone marrow, spleen and lymph nodes of 3–4-week-old pre-leukemic Eμ-Myc, Eμ-Myc;CD19-Cre and Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice and wild-type (control) mice was determined by cell counting (n=3–5). (b) The total numbers of pro-B (B220+c-Kit+sIg-), pre-B (B220+c-Kit-sIg-) and sIg+-B cells (B220+c-Kit-sIg+) in the bone marrow of 3–4-week-old pre-leukemic wild-type (control), Eμ-Myc, Eμ-Myc;CD19-Cre and Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice were determined by flow cytometric analysis. Comparison between Eμ-Myc;CD19-Cre and Eμ-Myc;CD19-Cre;Mcl-1fl/+mice: Ppro-B=0.24; Ppre-B=0.10; PsIg+-B=0.15 (n=3–5). (c) Expression of the human CD4 reporter on B-lymphoid cells (gated as B220+) from wild-type (control) and pre-leukemic Eμ-Myc, Eμ-Myc;CD19-Cre and Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice was determined by flow cytometric analysis
As some lymphomas that arose in Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice had been selected for retention of their Mcl-1fl allele, we hypothesized that there may be potent selection against loss of the Mcl-1fl allele already in the pre-leukemic state. To examine this, we stained spleen cells from pre-leukemic Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice as well as those from Eμ-Myc, CD19-Cre;Mcl-1fl/+ and wt animals with antibodies against B220 (B-cell marker) and hCD4 (reporter for Mcl-1fl deletion; Figure 3c). As expected, the B-lymphoid cells from Eμ-Myc and wt mice did not express hCD4. The B-lymphoid cells from CD19-Cre;Mcl-1fl/+mice were composed of two distinct populations, one negative (~60%) and the other positive (~40%) for hCD4 (Figure 3c). This demonstrates that some B-lymphoid cells in these animals were able to delete the Mcl-1fl allele, although the efficiency was not very high. This is consistent with the previously reported45 relatively poor recombination efficiency of the CD19-Cre deletion strain that we used in our experiments. Interestingly, in the Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice only ~20% of the pre-leukemic B-lymphoid cells were hCD4+ (Figure 3c).
These findings reveal that deregulated c-MYC expression exerts potent selection against loss of one Mcl-1fl allele in B-lymphoid cells, whereas loss of one Mcl-1 allele is more readily tolerated in normal B-lymphoid cells.
Efficient deletion of Mcl-1fl allele(s) in B-lymphoid progenitors using the Rag-1-Cre transgene substantially delays lymphomagenesis in Eμ-Myc mice
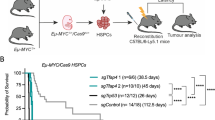
As CD19-Cre-mediated deletion of Mcl-1fl alleles was rather inefficient, we wanted to test whether deleting Mcl-1fl allele(s) more efficiently and at an earlier stage in B-cell development would have a greater impact in our lymphoma model. For this we employed the Rag-1-Cre knockin allele, which was reported to recombine floxed genes with very high efficiency at the CLP stage.46, 47 Lymphoma onset was slightly delayed in the Eμ-Myc;Rag-1-Cre mice compared with the Eμ-Myc control animals. Although this difference was not significant (P=0.06), this indicates that the Rag-1-Cre transgene exerts some toxicity on B-lymphoid cells undergoing neoplastic transformation. Remarkably, the median lymphoma-free survival of Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice (346 days) was far longer than in control Eμ-Myc (91 days) and Eμ-Myc;Rag-1-Cre mice (129 days, P**=0.003, Figure 4a), clearly demonstrating the importance of MCL-1 in c-MYC-induced lymphomagenesis.
Rag1-Cre-mediated deletion of one allele of Mcl-1 greatly delays lymphoma development in Eμ-Myc mice. (a) Kaplan–Meier animal survival curves comparing Eμ-Myc, Eμ-Myc;Rag1-Cre and Eμ-Myc;Rag1-Cre;Mcl-1fl/+ mice. Eμ-Myc;Rag1-Cre versus Eμ-Myc;Rag1-Cre;Mcl-1fl/+; Mantle–Cox Log-rank test P*=0.0030). Median onset of pre-B/B-cell lymphoma: Eμ-Myc: 91 days; Eμ-Myc;Rag1-Cre: 129 days, Eμ-Myc;Rag1-Cre;Mcl-1fl/+=346 days. (b) Lymphoma burden (spleen and lymph node weights) at autopsy in sick mice of indicated genotypes. A significant difference was observed in lymph node weights (P*=0.0307) between sick Eμ-Myc;Rag1-Cre and Eμ-Myc;Rag1-Cre;Mcl-1fl/+mice. (c) The numbers of lymphoid cells, erythrocytes and thrombocytes in the peripheral blood of lymphoma-burdened mice of the indicated genotypes were determined at autopsy using the ADVIA counter. Eμ-MycRag-1-Cre versus Eμ-Myc;Rag-1-Cre;Mcl-1fl/+: P*lymph=0.0464)
The lymphoma-burdened, sick Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice showed significantly lower lymph node weights (*P=0.031) and lymphocyte numbers in the peripheral blood (*P=0.046) than sick Eμ-Myc;Rag-1-Cre mice (Figures 4b and c). No significant differences were found in the spleen weights or in the numbers of erythrocytes and thrombocytes in the blood.
The marked delay in lymphoma development seen in the Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice suggested that Rag-1-Cre was considerably more efficient in Mcl-1fl deletion than CD19-Cre. To test this hypothesis, we analyzed lymphoma cells from Eμ-Myc;Rag-1-Cre and Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice for hCD4 expression (Figure 5a). Strikingly, the selection against cells expressing the hCD4 reporter (i.e. selection against cells that had deleted the Mcl-1fl allele) was clearly more potent in Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ lymphoma cells than in those from the Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice (compare data in Figures 2c and 5a). In the absence of oncogenic stress, Rag-1-Cre;Mcl-1fl/+mice efficiently deleted one Mcl-1 allele in B-lymphoid cells, but interestingly, there was potent selection against loss of both Mcl-1 alleles even without c-MYC overexpression (Figure 5b). These results reveal that non-transformed B-lymphoid cells can tolerate loss of one but not loss of both Mcl-1 alleles, whereas cells with deregulated c-MYC expression (both pre-leukemic cells undergoing transformation as well as malignant lymphomas) cannot tolerate even loss of a single allele. Unfortunately we were unable to generate Eμ-Myc;Rag-1-Cre;Mcl-1fl/fl mice due to issues with infertility.
Lymphomas from Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice are selected against loss of their Mcl-1fl allele. (a) Flow cytometric analysis of hCD4 reporter expression (reflecting Mcl-1fl recombination) on lymphoma cells from Eμ-Myc (control), Eμ-Myc;Rag-1-Cre and Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice. (b) Flow cytometric analysis of hCD4 reporter expression on normal B-lymphoid cells (gated as B220+) from 3–4-week-old Rag-1-Cre; Rag1-Cre;Mcl-1fl/+ and Rag1-Cre;Mcl-1fl/fl mice
Collectively, these data show that Mcl-1 is essential for the survival of MYC overexpressing pre-leukemic B-lymphoid cells undergoing neoplastic transformation. Therefore, B-lymphoid-restricted loss of one allele of Mcl-1 can substantially delay pre-B/B-lymphoma development in Eμ-Myc mice.
Discussion
Evasion of cell death is considered an essential requirement for the development of cancer.2 Impaired apoptosis in cancer cells (particularly in hematological malignancies) often results from deregulated expression of pro-survival or pro-apoptotic members of the BCL-2 protein family.48 In cells undergoing neoplastic transformation, apoptosis can be triggered by stress conditions induced by newly acquired oncogenic mutations (e.g. deregulated c-MYC expression) or by limiting availability of nutrients or growth factors from the tumor micro-environment. Regardless of the trigger that activates apoptosis signaling, evasion of cell death is essential for a population of nascent neoplastic cells to expand and sub-clones to acquire additional oncogenic lesions that cooperate with the initiating oncogenic mutation(s) to promote emergence of malignant cells.16
Although BCL-2 overexpression greatly accelerates lymphomagenesis in Eμ-Myc transgenic mice,25 endogenous BCL-2 is dispensable for MYC-driven lymphoma development.28 In contrast, BCL-XL was found to be essential for the survival of both normal as well as c-MYC overexpressing B-cell progenitors and its loss therefore inhibited lymphoma development in Eμ-Myc mice.29 Here we show that MCL-1 is also critical for c-MYC-driven lymphoma development.
We employed two Cre transgenic strains to delete Mcl-1 either at the late pro-B cell (CD19-Cre39) or the CLP stage (Rag-1-Cre46, 47). Surprisingly, we found that lymphoma development in the Eμ-Myc;CD19-Cre;Mcl-1fl/+ and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice was only slightly slower than in the control Eμ-Myc and Eμ-Myc;CD19-Cre mice. The difference to the Eμ-Myc mice was statistically significant but the difference to the Eμ-Myc;CD19-Cre mice was not, probably because constitutive Cre activity imposes a slight toxicity in B-lymphoid cells, as previously observed in other cell types.49 Interestingly, in young, pre-leukemic Eμ-Myc;CD19-Cre;Mcl-1fl/+ mice considerably fewer B-lymphoid cells had deleted their Mcl-1fl allele (detected as human CD4+) than in the CD19-Cre;Mcl-1fl/+ animals. This demonstrates that deregulated c-MYC expression renders nascent neoplastic cells exquisitely dependent on an adequate MCL-1 protein level (i.e., provided by both Mcl-1 alleles) for their survival. This selection against pre-leukemic B-lymphoid cells that had deleted their Mcl-1fl allele(s) explains why some pre-B/B lymphomas arising in Eμ-Myc;CD19-Cre;Mcl-1fl/+ and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice had been selected against loss of their Mcl-1fl allele(s). Thus, cells retaining their full MCL-1 complement had an advantage in progressing through further steps of neoplastic transformation. Moreover, lymphomas that arose in Eμ-Myc;CD19-Cre;Mcl-1fl/+ and Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice despite loss of one Mcl-1 allele appeared to have undergone selection for upregulation of BCL-XL and/or a reduction in pro-apoptotic BIM. This in turn suggests that keeping BIM in check constitutes a major function for MCL-1 in B-lymphoid cells undergoing transformation.
Lymphoma-free survival was extended to a much greater extent in Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice compared with the Eμ-Myc;CD19-Cre;Mcl-1fl/+ and Eμ-Myc;CD19-Cre;Mcl-1fl/fl animals. This may indicate that loss of one Mcl-1 allele at the earlier CLP stage of lymphoid cell development (i.e., when Rag1-Cre but not CD19-Cre is expressed) is more efficient in killing incipient neoplastic cells and therefore more efficient in delaying lymphoma development compared with Mcl-1fl deletion at the later pro-B-cell stage (when CD19-Cre expression commences). Alternatively, the Rag1-Cre transgene may simply be more efficient than the CD19-Cre transgene; the latter would therefore more readily allow escape of B-lymphoid cells that had failed to excise Mcl-1fl.
In conclusion, our findings demonstrate that MCL-1 is critical for the survival of c-MYC overexpressing lymphoma-initiating cells and hence for development of lymphoma. MCL-1 appears to be more important than BCL-XL because loss of one Mcl-1 allele substantially delayed lymphoma development in Eμ-Myc;Rag-1-Cre;Mcl-1fl/+ mice, whereas loss of one Bclx allele had only minor impact.29, 30 Loss of BIM-restored lymphoma development in mice with an Eμ-Myc;Bclx−/− lymphoid system and many pre-B/B lymphomas that arose in Eμ-Myc;CD19-Cre;Mcl-1fl/+ or Eμ-Myc;CD19-Cre;Mcl-1fl/fl mice despite loss of one Mcl-1 allele appeared to have undergone selection for low levels of BIM. This suggests that BIM is the critical pro-apoptotic BH3-only protein activated in response to oncogenic stress to kill Eμ-Myc pre-leukemic B-lymphoid cells to suppress progression to malignant lymphoma. These results and the observation that loss of even a single allele of Mcl-1 efficiently kills malignant Eμ-Myc lymphoma cells41 provide further impetus to develop MCL-1 specific inhibitors (e.g. BH3 mimetics) for cancer therapy.50, 51
Materials and methods
Experimental mice
All experiments with mice were conducted according to the guidelines of The Walter and Eliza Hall Institute of Medical Research Animal Ethics Committee. Eμ-Myc transgenic mice (generated on a mixed C57BL/6xSJL background and then backcrossed for >30 generations onto a C57BL/6 background) expressing the c-Myc transgene under control of the immunoglobulin heavy chain gene enhancer Eμ have been previously reported.52 The Mcl-1fl mice were generated on a C57BL/6 background using C57BL/6-derived ES cells.40 The Rag-1-CreKi/+46 and CD19-CreKi/+39 mice were generated on a mixed C57BL/6x129SV genetic background using 129SV-derived ES cells and then backcrossed onto a C57BL/6 background for >20 generations before commencement of our studies.
Genotyping
Genotyping was performed as previously reported.36 Oligonucleotide sequences for genotyping of these alleles will be provided on request.
Analysis of lymphoma-burdened mice
Eμ-Myc transgenic mice were examined daily by animal technicians for signs of malignant disease. Mice were sacrificed when declared unwell by the animal technicians. Signs of disease included splenomegaly, lymphadenopathy, hind-limb paralysis, hunched stature, weight loss and labored breathing (indicative of lymphoma growth in the thymus). Sick mice were euthanized, tissues removed, weighed and then used for flow cytometric as well as histological analyses and tissue culture.
Western blot analysis
Cells were lysed in RIPA buffer supplemented with a protease inhibitor cocktail (Roche, Basel, Switzerland). Protein lysates (30 μg protein) mixed with 4x Laemmli buffer were loaded onto a 10% Bis/Tris gel (Life Technologies, Scoresby, VIC, Australia) and electrophoresis was conducted according to the manufacturer’s instructions. Proteins were transferred onto nitrocellulose membranes using the iBlot system (Life Technologies, Scoresby, VIC, Australia). Nitrocellulose membranes were blocked for 2 h using 5% skim milk powder dissolved in phosphate-buffered saline supplemented with 0.5% Tween-20. Western blots were probed with the following monoclonal or polyclonal antibodies: rabbit anti-mouse MCL-1 (19C4-15), hamster anti-mouse BCL-2 (3F11), mouse anti-mouse BCL-XL (BD Pharmingen, BD BioSciences, San Jose, CA, USA; 2F12), rabbit anti-mouse BIM (Stressgen, 9292), rabbit anti-mouse PUMA (Ab-27669, Abcam, Melbourne, Victoria, Australia), mouse anti-HSP70 (R Anderson, Peter McCallum Cancer Centre; loading control), rabbit anti-mouse p53 (Leica Biosystems, Mount Waverley, Victoria, Australia; CM5) and rat anti-mouse p19-ARF (Rockland Immunochemicals, Pottstown, PA, USA; 5.C3.1), overnight at 4 °C. Blots were washed three times in phosphate-buffered saline supplemented with 0.5% Tween-20. The blots were then incubated for 1 h at room temperature with secondary HRP-conjugated antibodies against mouse, rat, hamster or rabbit IgG and again washed before exposure to the Amersham ECL reaction and developing on an autoradiograph Hyperfilm (GE Healthcare, Parramatta, NSW, Australia).
Lymphoma and pre-leukemic analysis by flow cytometry
Lymphoid organs were harvested from lymphoma-burdened mice and single-cell suspensions prepared using forceps. Cells (5 × 106) were resuspended in buffered saline supplemented with 10% FCS and 2% normal rat serum and stained for 30 min at 4 °C with rat monoclonal antibodies to B220 (RA3-6B2, The Walter and Eliza Hall Institute (WEHI)), cKIT (ACK4, WEHI), IgM (5.1, WEHI) and IgD (11-26, WEHI; all produced and conjugated with fluorochromes in our laboratory) and mouse monoclonal antibody to human CD4 (BD Pharmingen #555347, RPA-T4).
Statistical analysis
Kaplan–Meier mouse survival curves were generated and analyzed with GraphPad Prism (GraphPad Software Inc, La Jolla, CA, USA). Mouse cohorts were compared using the log-rank Mantel–Cox test. P-values of <0.05 were considered significant. In vitro cell survival, blood cell counts, organ weights and RNA levels were plotted and analyzed with GraphPad Prism using two-tailed student’s t-test comparing two groups with each other. Error bars are presented as standard error of mean (±s.e.m.).
Abbreviations
- BH3:
-
BCL-2 Homology domain 3
- BIM:
-
BCL-2-interacting mediator of cell death
- PUMA:
-
p53 upregulated modulator of apoptosis
- Noxa:
-
phorbol-12-myr- istate-13-acetate-induced protein 1
- BAK:
-
BCL-2 antagonist/killer-1
- BAX:
-
BCL-2-associated X protein
- BOK:
-
BCL-2 related ovarian killer
- BCL-2:
-
B-cell lymphoma 2
- BCL-XL:
-
BCL-2-like 1 extra long
- MCL-1:
-
myeloid cell leukemia-1
- BCL-W:
-
BCL-2-like 2
- A1/BFL-1:
-
BCL-2-related protein A1
Bcl-2 related gene in the fetal liver
- MLL-ENL:
-
mixed lineage leukemia-eleven-nineteen leukemia
- MLL-AF9:
-
ALL1 fused gene from chromosome 9
- Rag-1:
-
recombination activating gene 1
- CD4:
-
cluster of differentiation 4
- Cre:
-
cyclization recombination
- CD10:
-
cluster of differentiation 19
- PCR:
-
polymerase chain reaction
- FACS:
-
fluorescence-activated cell sorting
References
Hotchkiss RS, Strasser A, McDunn JE, Swanson PE . Cell death. New Engl J Med 2009; 361: 1570–1583.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Strasser A, Harris AW, Cory S . Bcl-2 transgene inhibits T cell death and perturbs thymic self-censorship. Cell 1991; 67: 889–899.
Youle RJ, Strasser A . The BCL-2 protein family: opposing activities that mediate cell death. Nat Rev Mol Cell Biol 2008; 9: 47–59.
Czabotar PE, Lessene G, Strasser A, Adams JM . Control of apoptosis by the BCL-2 protein family: implications for physiology and therapy. Nat Rev Mol Cell Biol 2014; 15: 49–63.
Huang DCS, Strasser A . BH3-only proteins – essential initiators of apoptotic cell death. Cell 2000; 103: 839–842.
Puthalakath H, Strasser A . Keeping killers on a tight leash: transcriptional and post-translational control of the pro-apoptotic activity of BH3-only proteins. Cell Death Differ 2002; 9: 505–512.
Ke F, Voss A, Kerr JB, O'Reilly LA, Tai L, Echeverry N et al. BCL-2 family member BOK is widely expressed but its loss has only minimal impact in mice. Cell Death Differ 2012; 19: 915–925.
Chipuk JE, Green DR . How do BCL-2 proteins induce mitochondrial outer membrane permeabilization? Trends Cell Biol 2008; 18: 157–164.
Rinkenberger JL, Horning S, Klocke B, Roth K, Korsmeyer SJ . Mcl-1 deficiency results in peri-implantation embryonic lethality. Genes Dev 2000; 14: 23–27.
Opferman J, Iwasaki H, Ong CC, Suh H, Mizuno S, Akashi K et al. Obligate role of anti-apoptotic MCL-1 in the survival of hematopoietic stem cells. Science 2005; 307: 1101–1104.
Opferman JT, Letai A, Beard C, Sorcinelli MD, Ong CC, Korsmeyer SJ . Development and maintenance of B and T lymphocytes requires antiapoptotic MCL-1. Nature 2003; 426: 671–676.
Dzhagalov I St, John A, He YW . The antiapoptotic protein Mcl-1 is essential for the survival of neutrophils but not macrophages. Blood 2007; 109: 1620–1626.
Letai AG . Diagnosing and exploiting cancer's addiction to blocks in apoptosis. Nat Rev Cancer 2008; 8: 121–132.
Adams JM, Cory S . The Bcl-2 apoptotic switch in cancer development and therapy. Oncogene 2007; 26: 1324–1337.
Adams JM, Cory S . Bcl-2-regulated apoptosis: mechanism and therapeutic potential. Curr Opin Immunol 2007; 19: 488–496.
Strasser A, Harris AW, Vaux DL, Webb E, Bath ML, Adams JM et al. Abnormalities of the immune system induced by dysregulated bcl-2 expression in transgenic mice. Curr Top Microbiol Immunol 1990; 166: 175–181.
Tsujimoto Y, Cossman J, Jaffe E, Croce CM . Involvement of the bcl-2 gene in human follicular lymphoma. Science 1985; 228: 1440–1443.
Tsujimoto Y, Gorham J, Cossman J, Jaffe E, Croce CM . The t(14;18) chromosome translocations involved in B-cell neoplasms result from mistakes in VDJ joining. Science 1985; 229: 1390–1393.
Tagawa H, Karnan S, Suzuki R, Matsuo K, Zhang X, Ota A et al. Genome-wide array-based CGH for mantle cell lymphoma: identification of homozygous deletions of the proapoptotic gene BIM. Oncogene 2005; 24: 1348–1358.
Egle A, Harris AW, Bath ML, O’Reilly L, Cory S . VavP-Bcl2 transgenic mice develop follicular lymphoma preceded by germinal center hyperplasia. Blood 2004; 103: 2276–2283.
Egle A, Harris AW, Bouillet P, Cory S . Bim is a suppressor of Myc-induced mouse B cell leukemia. Proc Natl Acad Sci USA 2004; 101: 6164–6169.
Hemann MT, Zilfou JT, Zhao Z, Burgess DJ, Hannon GJ, Lowe SW . Suppression of tumorigenesis by the p53 target PUMA. Proc Natl Acad Sci USA 2004; 101: 9333–9338.
Michalak EM, Jansen ES, Happo L, Cragg MS, Tai L, Smyth GK et al. Puma and to a lesser extent Noxa are suppressors of Myc-induced lymphomagenesis. Cell Death Differ 2009; 16: 684–696.
Strasser A, Harris AW, Bath ML, Cory S . Novel primitive lymphoid tumours induced in transgenic mice by cooperation between myc and bcl-2. Nature 1990; 348: 331–333.
Vandenberg CJ, Cory S . ABT-199, a new Bcl-2-specific BH3 mimetic, has in vivo efficacy against aggressive Myc-driven mouse lymphomas without provoking thrombocytopenia. Blood 2013; 121: 2285–2288.
Letai A, Sorcinelli MD, Beard C, Korsmeyer SJ . Antiapoptotic BCL-2 is required for maintenance of a model leukemia. Cancer Cell 2004; 6: 241–249.
Kelly PN, Puthalakath H, Adams JM, Strasser A . Endogenous bcl-2 is not required for the development of Eμ-myc-induced B-cell lymphoma. Blood 2007; 109: 4907–4913.
Kelly PN, Grabow S, Delbridge ARD, Strasser A, Adams JM . Endogenous Bcl-xL is essential for Myc-driven lymphomagenesis in mice. Blood 2011; 118: 6380–6386.
Delbridge AR, Grabow S, Bouillet P, Adams JM, Strasser A . Functional antagonism between pro-apoptotic BIM and anti-apoptotic BCL-XL in MYC-induced lymphomagenesis. Oncogene 2015; 34: 1872–1876.
Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J et al. The landscape of somatic copy-number alteration across human cancers. Nature 2010; 463: 899–905.
Xiang Z, Luo H, Payton JE, Cain J, Ley TJ, Opferman JT et al. Mcl1 haploinsufficiency protects mice from Myc-induced acute myeloid leukemia. J Clin Invest 2010; 120: 2109–2118.
Glaser SP, Lee EF, Trounson E, Bouillet P, Wei A, Fairlie WD et al. Anti-apoptotic Mcl-1 is essential for the development and sustained growth of acute myeloid leukemia. Genes Dev 2012; 26: 120–125.
Koss B, Morrison J, Perciavalle RM, Singh H, Rehg JE, Williams RT et al. Requirement for antiapoptotic MCL-1 in the survival of BCR-ABL B-lineage acute lymphoblastic leukemia. Blood 2013; 122: 1587–1598.
Kelly GL, Grabow S, Glaser SP, Fitzsimmons L, Aubrey BJ, Okamoto T et al. Targeting of MCL-1 kills MYC-driven mouse and human lymphomas even when they bear mutations in p53. Genes Dev 2014; 28: 58–70.
Delbridge A, Opferman JT, Grabow S, Strasser A . Pro-survival MCL-1 and pro-apoptotic PUMA govern stem/progenitor cell survival during emergency hematopoiesis. Blood 2015; 125: 3273–3280.
Langdon WY, Harris AW, Cory S, Adams JM . The c-myc oncogene perturbs B lymphocyte development in Eμ-myc transgenic mice. Cell 1986; 47: 11–18.
Harris AW, Pinkert CA, Crawford M, Langdon WY, Brinster RL, Adams JM . The Eμ-myc transgenic mouse: a model for high-incidence spontaneous lymphoma and leukemia of early B cells. J Exp Med 1988; 167: 353–371.
Rickert RC, Roes J, Rajewsky K . B lymphocyte-specific, Cre-mediated mutagenesis in mice. Nucleic Acids Res 1997; 25: 1317–1318.
Vikstrom I, Carotta S, Lüthje K, Peperzak V, Jost PJ, Glaser S et al. Mcl-1 is essential for germinal center formation and B cell memory. Science 2010; 330: 1095–1099.
Kelly GL, Grabow S, Glaser SP, Fitzsimmons L, Aubrey BJ, Okamoto T et al. Targeting of MCL-1 kills MYC-driven mouse and human lymphoma cells even when they bear mutations in p53. Genes Dev 2013; 28: 58–70.
Soucek L, Evan GI . The ups and downs of Myc biology. Curr Opin Genetics Dev 2010; 20: 91–95.
Evan GI, Wyllie AH, Gilbert CS, Littlewood TD, Land H, Brooks M et al. Induction of apoptosis in fibroblasts by c-myc protein. Cell 1992; 69: 119–128.
Strasser A, Elefanty AG, Harris AW, Cory S . Progenitor tumours from Em-bcl-2-myc transgenic mice have lymphomyeloid differentiation potential and reveal developmental differences in cell survival. EMBO J 1996; 15: 3823–3834.
Hobeika E, Thiemann S, Storch B, Jumaa H, Nielsen PJ, Pelanda R et al. Testing gene function early in the B cell lineage in mb1-cre mice. Proc Natl Acad Sci USA 2006; 103: 13789–13794.
McCormack MP, Forster A, Drynan L, Pannell R, Rabbitts TH . The LMO2 T-cell oncogene is activated via chromosomal translocations or retroviral insertion during gene therapy but has no mandatory role in normal T-cell development. Mol Cell Biol 2003; 23: 9003–9013.
Igarashi H, Gregory SC, Yokota T, Sakaguchi N, Kincade PW . Transcription from the RAG1 locus marks the earliest lymphocyte progenitors in bone marrow. Immunity 2002; 17: 117–130.
Adams JM, Cory S . The Bcl-2 protein family: arbiters of cell survival. Science 1998; 281: 1322–1326.
Schmidt-Supprian M, Rajewsky K . Vagaries of conditional gene targeting. Nat Immunol 2007; 8: 665–668.
Leverson JD, Zhang H, Chen J, Tahir SK, Phillips DC, Xue J et al. Potent and selective small-molecule MCL-1 inhibitors demonstrate on-target cancer cell killing activity as single agents and in combination with ABT-263 (navitoclax). Cell Death Dis 2015; 6: e1590.
Goodwin CM, Rossanese OW, Olejniczak ET, Fesik SW . Myeloid cell leukemia-1 is an important apoptotic survival factor in triple-negative breast cancer. Cell Death Differ 2015; 22: 2098–2106.
Adams JM, Harris AW, Pinkert CA, Corcoran LM, Alexander WS, Cory S et al. The c-myc oncogene driven by immunoglobulin enhancers induces lymphoid malignancy in transgenic mice. Nature 1985; 318: 533–538.
Okamoto T, Coultas L, Metcalf D, van Delft MF, Glaser SP, Takiguchi M et al. Enhanced stability of Mcl1, a prosurvival Bcl2 relative, blunts stress-induced apoptosis, causes male sterility, and promotes tumorigenesis. Proc Natl Acad Sci USA 2014; 111: 261–266.
Acknowledgements
We thank Prof S Cory for mice; Dr. LA O’Reilly for discussions; L Reid, J Mansheim, N Iannarella, S Allan, C D’Alessandro and G Siciliano for expert animal care; B Helbert for genotyping; J Corbin for automated blood analysis. This work was supported by grants and fellowships from the Cancer Council of Victoria (SG, ARDD), Leukaemia Foundation Australia (SG), the Lady Tata Memorial Trust (SG), Cure Brain Cancer Australia (AS), the National Health and Medical Research Council (Program Grant #1016701, NHMRC Australia Fellowship 1020363 to AS, Project Grant #1086291 to GLK), the Leukemia and Lymphoma Society (SOCR Grant #7001-03 to AS), Kay Kendall Leukemia Fund Intermediate Fellowship KKL331 (GLK), Cancer Council Victoria Grant-In-Aid #1086157 (GLK), Melbourne International Research and the Melbourne International Fee Remission Scholarship (University of Melbourne, SG) and Cancer Therapeutics CRC Top-up Scholarship (SG, ARDD). The estate of Anthony (Toni) Redstone OAM, University of Melbourne International Research and International Fee Remission Scholarships (SG), Australian Postgraduate Award (ARDD), and the operational infrastructure grants through the Australian Government IRIISS and the Victorian State Government OIS.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
SG, ARDD, GLK, PB, JMA and AS are employed by The Walter and Eliza Hall Institute, which receives milestone payments from Genentech and AbbVie for the development of ABT-199 for cancer therapy.
Additional information
Edited by G Melino
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Grabow, S., Kelly, G., Delbridge, A. et al. Critical B-lymphoid cell intrinsic role of endogenous MCL-1 in c-MYC-induced lymphomagenesis. Cell Death Dis 7, e2132 (2016). https://doi.org/10.1038/cddis.2016.43
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2016.43
This article is cited by
-
Synergistic apoptotic effect of Mcl-1 inhibition and doxorubicin on B-cell precursor acute lymphoblastic leukemia cells
Molecular Biology Reports (2022)
-
Ubiquitination and deubiquitination of MCL1 in cancer: deciphering chemoresistance mechanisms and providing potential therapeutic options
Cell Death & Disease (2020)
-
IBTK contributes to B-cell lymphomagenesis in Eμ-myc transgenic mice conferring resistance to apoptosis
Cell Death & Disease (2019)
-
Survival control of malignant lymphocytes by anti-apoptotic MCL-1
Leukemia (2016)