Abstract
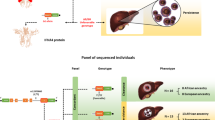
Host–viral genetic interaction has a key role in hepatitis C infection (HCV) and maybe in the viral selection. In a preliminary GWAS analysis, we identified BTN3A2 rs9104 to be associated with HCV genotype 1. Therefore, our aim was to determine the influence of BTN family on the selection of HCV genotype. We performed a fine-mapping analysis of BTN gene region in a cohort of chronic HCV infection (N=841), validating significant results in another independent chronic HCV infection cohort (N=637), according to selection of viral genotype. BTN3A2 rs9104, BTN3A2 rs733528, BTN2A1 rs6929846, BTN2A1 rs7763910 and BTN3A3 rs13220495 were associated with viral genotype selection. Interestingly, BTN3A2 rs9104 GG genotype was closely related to genotype 1 infection (80.7% (394/488) compared with genotype 3 infection (53.5% (23/43); P=0.0001) in patients harboring IL28B-CT/TT genotype, although this effect was not observed in IL28B-CC genotype. Similarly, BTN3A3 rs13220495 CC genotype was linked to genotype 3 infection (100% (32/32)) compared to genotype 1 (87.3% (137/157); P=0.028) in patients harboring IL28B-CC genotype, but did not in IL28B-CT/TT genotype. Genetic variants in the butyrophilin family genes may alter susceptibility to infection, selecting HCV genotype and influencing disease progression. BTN3A2 rs9104 was strongly associated with genotype 1 infection and the haplotype BTN3A3 rs13220495 CC+IL28B genotype CC was universal in patients with hepatitis C genotype 3a.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Grebely J, Page K, Sacks-Davis R, van der Loeff MS, Rice TM, Bruneau J . InC3 Study Group. The effects of female sex, viral genotype, and IL28B genotype on spontaneous clearance of acute hepatitis C virus infection. Hepatology 2014; 59: 109–120.
Saito T, Ueno Y . Transmission of hepatitis C virus: self-limiting hepatitis or chronic hepatitis? World J Gastroenterol 2013; 19: 6957–6961.
Romero V, Azocar J, Zúñiga J, Clavijo OP, Terreros D, Gu X et al. Interaction of NK inhibitory receptor genes with HLA-C and MHC class II alleles in Hepatitis C virus infection outcome. Mol Immunol 2008; 45: 2429–2436.
Montes-Cano MA, Caro-Oleas JL, Romero-Gómez M, Diago M, Andrade R, Carmona I et al. HLA-C and KIR genes in hepatitis C virus infection. Hum Immunol 2005; 66: 1106–1109.
Cubero M, Gregori J, Esteban JI, García-Cehic D, Bes M, Perales C et al. Identification of host and viral factors involved in a dissimilar resolution of a hepatitis C virus infection. Liver Int 2014; 34: 896–906.
Thomas DL, Thio CL, Martin MP, Qi Y, Ge D, O'Huigin C et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature 2009; 461: 798–801.
Zhu YZ, Qian XJ, Zhao P, Qi ZT . How hepatitis C virus invades hepatocytes: the mystery of viral entry. World J Gastroenterol 2014; 20: 3457–3467.
Rojas L, Ampuero J, Del Campo JA, Garcia-Lozano RJ, Solá R, Forns X et al. Fine mapping of the butyrophilin genomics region: Role in hepatitis C virus infection (HCV). J Hepatol 2014; 60: S139.
Bekker V, Chanock SJ, Yeager M, Hutchinson AA, von Hahn T, Chen S et al. Genetic variation in CLDN1 and susceptibility to hepatitis C virus infection. J Viral Hepat 2010; 17: 192–200.
Viken MK, Blomhoff A, Olsson M, Akselsen HE, Pociot F, Nerup J et al. Reproducible association with type 1 diabetes in the extended class I region of the major histocompatibility complex. Genes Immun 2009; 10: 323–333.
Le Page C, Marineau A, Bonza PK, Rahimi K, Cyr L, Labouba I et al. BTN3A2 expression in epithelial ovarian cancer is associated with higher tumor infiltrating T cells and a better prognosis. PLoS One 2012; 7: e38541.
Bamne M, Wood J, Chowdari K, Watson AM, Celik C, Mansour H et al. Evaluation of HLA polymorphisms in relation to schizophrenia risk and infectious exposure. Schizophr Bull 2012; 38: 1149–1154.
Montes-Cano MA, García-Lozano JR, Abad-Molina C, Romero-Gómez M, Barroso N, Aguilar-Reina J et al. Interleukin-28B genetic variants and hepatitis virus infection by different viral genotypes. Hepatology 2010; 52: 33–37.
Ampuero J, Romero-Gómez M, Reddy KR . Review article: HCV genotype 3–the new treatment challenge. Aliment Pharmacol Ther. 2014; 39: 686–698.
Horner SM, Gale M Jr . Regulation of hepatic innate immunity by hepatitis C virus. Nat Med 2013; 19: 879–888.
Khakoo SI, Thio CL, Martin MP, Brooks CR, Gao X, Astemborski J et al. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science 2004; 305: 872–874.
Terilli RR, Cox AL . Immunity and hepatitis C: a review. Curr HIV/AIDS Rep 2013; 10: 51–58.
Amadei B, Urbani S, Cazaly A, Fisicaro P, Zerbini A, Ahmed P et al. Activation of natural killer cells during acute infection with hepatitis C virus. Gastroenterology 2010; 138: 1536–1545.
Messal N, Mamessier E, Sylvain A, Celis-Gutierrez J, Thibult ML, Chetaille B et al. Differential role for CD277 as a co-regulator of the immune signal in T and NK cells. Eur J Immunol 2011; 41: 3443–3454.
Acknowledgements
This work has been supported by Grant #P013/01192 of the Instituto de Salud Carlos III of the Spanish Government.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Genes and Immunity website
Supplementary information
Rights and permissions
About this article
Cite this article
Ampuero, J., del Campo, J., Rojas, L. et al. Fine-mapping butyrophilin family genes revealed several polymorphisms influencing viral genotype selection in hepatitis C infection. Genes Immun 16, 297–300 (2015). https://doi.org/10.1038/gene.2015.14
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2015.14