Abstract
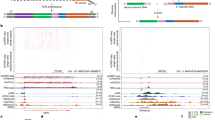
Gene expression in multiple individual cells from a tissue or culture sample varies according to cell-cycle, genetic, epigenetic and stochastic differences between the cells. However, single-cell differences have been largely neglected in the analysis of the functional consequences of genetic variation. Here we measure the expression of 92 genes affected by Wnt signaling in 1,440 single cells from 15 individuals to associate single-nucleotide polymorphisms (SNPs) with gene-expression phenotypes, while accounting for stochastic and cell-cycle differences between cells. We provide evidence that many heritable variations in gene function—such as burst size, burst frequency, cell cycle–specific expression and expression correlation/noise between cells—are masked when expression is averaged over many cells. Our results demonstrate how single-cell analyses provide insights into the mechanistic and network effects of genetic variability, with improved statistical power to model these effects on gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Nica, A.C. et al. The architecture of gene regulatory variation across multiple human tissues: the MuTHER study. PLoS Genet. 7, e1002003 (2011).
Li, H. & Deng, H. Systems genetics, bioinformatics and eQTL mapping. Genetica 138, 915–924 (2010).
Bak, P. et al. Self-organized criticality: an explanation of the 1/f noise. Phys. Rev. Lett. 59, 381–384 (1987).
Shalek, A.K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Nicolae, D.L. et al. Trait-associated SNPs are more likely to be eQTLs: annotation to enhance discovery from GWAS. PLoS Genet. 6, e1000888 (2010).
Islam, S. et al. Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq. Genome Res. 21, 1160–1167 (2011).
Ramskold, D. et al. Full-length mRNA-Seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Livak, K.J. et al. Methods for qPCR gene expression profiling applied to 1440 lymphoblastoid single cells. Methods 59, 71–79 (2013).
International HapMap 3 Consortium. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Coghlan, M.P. et al. Selective small molecule inhibitors of glycogen synthase kinase-3 modulate glycogen metabolism and gene transcription. Chem. Biol. 7, 793–803 (2000).
Dar, R.D. et al. Transcriptional burst frequency and burst size are equally modulated across the human genome. Proc. Natl. Acad. Sci. USA 109, 17454–17459 (2012).
Bengtsson, M., Stahlberg, A., Rorsman, P. & Kubista, M. Gene expression profiling in single cells from the pancreatic islets of Langerhans reveals lognormal distribution of mRNA levels. Genome Res. 15, 1388–1392 (2005).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule Sensitivity in single cells. Science 329, 533–538 (2010).
Bublik, D.R.R., Scolz, M., Triolo, G., Monte, M. & Schneider, C. Human GTSE-1 regulates p21(CIP1/WAF1) stability conferring resistance to paclitaxel treatment. J. Biol. Chem. 285, 5274–5281 (2010).
Choy, E. et al. Genetic analysis of human traits in vitro: drug response and gene expression in lymphoblastoid cell lines. PLoS Genet. 4, e1000287 (2008).
Im, H.K.K. et al. Mixed effects modeling of proliferation rates in cell-based models: consequence for pharmacogenomics and cancer. PLoS Genet. 8, e1002525 (2012).
Cuddapah, S. et al. Global analysis of the insulator binding protein CTCF in chromatin barrier regions reveals demarcation of active and repressive domains. Genome Res. 19, 24–32 (2009).
Hardy, R.R. & Hayakawa, K. B cell development pathways. Annu. Rev. Immunol. 19, 595–621 (2001).
Wu, B., Piatkevich, K.D., Lionnet, T., Singer, R.H. & Verkhusha, V.V. Modern fluorescent proteins and imaging technologies to study gene expression, nuclear localization, and dynamics. Curr. Opin. Cell Biol. 23, 310–317 (2011).
Siegel, A.F. Robust regression using repeated medians. Biometrika 69, 242–244 (1982).
Johnstone, I.M. & Velleman, P.F. The resistant line and related regression methods. J. Am. Stat. Assoc. 80, 1041–1054 (1985).
Sen, P.K. Estimates of the regression coefficient based on Kendall's Tau. J. Am. Stat. Assoc. 63, 1379–1389 (1968).
Acknowledgements
Many thanks to L. Toji at the Coriell Institute for her valuable input on the cell line growth and transformation characteristics. Also, thanks to the following people at Fluidigm: B. Jones for his overall support, G. Harris and D. Wang for their help with primer design, and the meticulous technical assistance of K. Datta and R. Mittal. C.H. and T.E. are funded by the Medical Research Council of the UK. T.E. is also funded by Leukaemia Lymphoma Research and EuroSyStem.
Author information
Authors and Affiliations
Contributions
Q.F.W. and C.H. conceived and designed the study. A.J.T. and T.E. ran the initial flow cytometry characterization and cell culture optimization. A.J.G. and D.W.S. ran the main study's cell culture and flow cytometry, further optimizing the sample characterization. K.J.L. designed and optimized the single-cell RNA assays, and generated the gene expression chip data. Q.F.W. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
K.L. is an employee of the Fluidigm Corporation.
Supplementary information
Supplementary Text and Figures
Supplementary Notes, Supplementary Figures 1–11 and Supplementary Tables 1–2 (PDF 2231 kb)
Supplementary Data
Supplementary Data (XLSX 944 kb)
Rights and permissions
About this article
Cite this article
Wills, Q., Livak, K., Tipping, A. et al. Single-cell gene expression analysis reveals genetic associations masked in whole-tissue experiments. Nat Biotechnol 31, 748–752 (2013). https://doi.org/10.1038/nbt.2642
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2642
This article is cited by
-
RNA recording in single bacterial cells using reprogrammed tracrRNAs
Nature Biotechnology (2023)
-
Transcriptomic and proteomic retinal pigment epithelium signatures of age-related macular degeneration
Nature Communications (2022)
-
Single-cell expression quantitative trait loci (eQTL) analysis of SLE-risk loci in lupus patient monocytes
Arthritis Research & Therapy (2021)
-
Single-cell transcriptomic profiling and characterization of endothelial progenitor cells: new approach for finding novel markers
Stem Cell Research & Therapy (2021)
-
Cell-to-cell expression dispersion of B-cell surface proteins is linked to genetic variants in humans
Communications Biology (2020)