Abstract
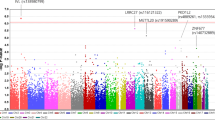
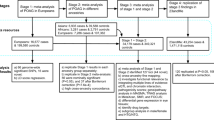
Exfoliation syndrome (XFS) is the most common known risk factor for secondary glaucoma and a major cause of blindness worldwide. Variants in two genes, LOXL1 and CACNA1A, have previously been associated with XFS. To further elucidate the genetic basis of XFS, we collected a global sample of XFS cases to refine the association at LOXL1, which previously showed inconsistent results across populations, and to identify new variants associated with XFS. We identified a rare protective allele at LOXL1 (p.Phe407, odds ratio (OR) = 25, P = 2.9 × 10−14) through deep resequencing of XFS cases and controls from nine countries. A genome-wide association study (GWAS) of XFS cases and controls from 24 countries followed by replication in 18 countries identified seven genome-wide significant loci (P < 5 × 10−8). We identified association signals at 13q12 (POMP), 11q23.3 (TMEM136), 6p21 (AGPAT1), 3p24 (RBMS3) and 5q23 (near SEMA6A). These findings provide biological insights into the pathology of XFS and highlight a potential role for naturally occurring rare LOXL1 variants in disease biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Vesti, E. & Kivelä, T. Exfoliation syndrome and exfoliation glaucoma. Prog. Retin. Eye Res. 19, 345–368 (2000).
Ritch, R. & Schlötzer-Schrehardt, U. Exfoliation syndrome. Surv. Ophthalmol. 45, 265–315 (2001).
Leske, M.C. et al. Factors for glaucoma progression and the effect of treatment: the early manifest glaucoma trial. Arch. Ophthalmol. 121, 48–56 (2003).
Ritch, R., Schlötzer-Schrehardt, U. & Konstas, A.G. Why is glaucoma associated with exfoliation syndrome? Prog. Retin. Eye Res. 22, 253–275 (2003).
Olawoye, O.O. et al. Exfoliation syndrome in Nigeria. Middle East Afr. J. Ophthalmol. 19, 402–405 (2012).
Kivelä, T. Ocular pseudoexfoliation syndrome and life span: act 2. EBioMedicine 2, 640–641 (2015).
Ritch, R. Ocular and systemic manifestations of exfoliation syndrome. J. Glaucoma 23 (Suppl. 1), S1–S8 (2014).
Ritch, R. Exfoliation syndrome—the most common identifiable cause of open-angle glaucoma. J. Glaucoma 3, 176–177 (1994).
Allingham, R.R. et al. Pseudoexfoliation syndrome in Icelandic families. Br. J. Ophthalmol. 85, 702–707 (2001).
Orr, A.C. et al. Exfoliation syndrome: clinical and genetic features. Ophthalmic Genet. 22, 171–185 (2001).
Thorleifsson, G. et al. Common sequence variants in the LOXL1 gene confer susceptibility to exfoliation glaucoma. Science 317, 1397–1400 (2007).
Williams, S.E. et al. Major LOXL1 risk allele is reversed in exfoliation glaucoma in a black South African population. Mol. Vis. 16, 705–712 (2010).
Wiggs, J.L. & Pasquale, L.R. Expression and regulation of LOXL1 and elastin-related genes in eyes with exfoliation syndrome. J. Glaucoma 23 (Suppl. 1), S62–S63 (2014).
Aung, T. et al. A common variant mapping to CACNA1A is associated with susceptibility to exfoliation syndrome. Nat. Genet. 47, 387–392 (2015).
Al Olama, A.A. et al. A meta-analysis of 87,040 individuals identifies 23 new susceptibility loci for prostate cancer. Nat. Genet. 46, 1103–1109 (2014).
Liu, J.Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Paternoster, L. et al. Multi-ancestry genome-wide association study of 21,000 cases and 95,000 controls identifies new risk loci for atopic dermatitis. Nat. Genet. 47, 1449–1456 (2015).
Ehret, G.B. et al. The genetics of blood pressure regulation and its target organs from association studies in 342,415 individuals. Nat. Genet. 48, 1171–1184 (2016).
Liu, C. et al. Meta-analysis identifies common and rare variants influencing blood pressure and overlapping with metabolic trait loci. Nat. Genet. 48, 1162–1170 (2016).
Surendran, P. et al. Trans-ancestry meta-analyses identify rare and common variants associated with blood pressure and hypertension. Nat. Genet. 48, 1151–1161 (2016).
Hauser, M.A. et al. Genetic variants and cellular stressors associated with exfoliation syndrome modulate promoter activity of a lncRNA within the LOXL1 locus. Hum. Mol. Genet. 24, 6552–6563 (2015).
Chen, H. et al. Ethnicity-based subgroup meta-analysis of the association of LOXL1 polymorphisms with glaucoma. Mol. Vis. 16, 167–177 (2010).
Fan, B.J. et al. LOXL1 promoter haplotypes are associated with exfoliation syndrome in a U.S. Caucasian population. Invest. Ophthalmol. Vis. Sci. 52, 2372–2378 (2011).
Ozaki, M. et al. Association of LOXL1 gene polymorphisms with pseudoexfoliation in the Japanese. Invest. Ophthalmol. Vis. Sci. 49, 3976–3980 (2008).
Tanito, M. et al. LOXL1 variants in elderly Japanese patients with exfoliation syndrome/glaucoma, primary open-angle glaucoma, normal tension glaucoma, and cataract. Mol. Vis. 14, 1898–1905 (2008).
Pasutto, F. et al. Association of LOXL1 common sequence variants in German and Italian patients with pseudoexfoliation syndrome and pseudoexfoliation glaucoma. Invest. Ophthalmol. Vis. Sci. 49, 1459–1463 (2008).
Mori, K. et al. LOXL1 genetic polymorphisms are associated with exfoliation glaucoma in the Japanese population. Mol. Vis. 14, 1037–1040 (2008).
Nakano, M. et al. Novel common variants and susceptible haplotype for exfoliation glaucoma specific to Asian population. Sci. Rep. 4, 5340 (2014).
Flannick, J. et al. Loss-of-function mutations in SLC30A8 protect against type 2 diabetes. Nat. Genet. 46, 357–363 (2014).
Bonnefond, A. et al. Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Nat. Genet. 44, 297–301 (2012).
Majithia, A.R. et al. Rare variants in PPARG with decreased activity in adipocyte differentiation are associated with increased risk of type 2 diabetes. Proc. Natl. Acad. Sci. USA 111, 13127–13132 (2014).
Majithia, A.R. et al. Prospective functional classification of all possible missense variants in PPARG. Nat. Genet. 48, 1570–1575 (2016).
Do, R. et al. Exome sequencing identifies rare LDLR and APOA5 alleles conferring risk for myocardial infarction. Nature 518, 102–106 (2015).
Luo, Y. et al. Exploring the genetic architecture of inflammatory bowel disease by whole-genome sequencing identifies association at ADCY7. Nat. Genet. 49, 186–192 (2017).
Raychaudhuri, S. et al. A rare penetrant mutation in CFH confers high risk of age-related macular degeneration. Nat. Genet. 43, 1232–1236 (2011).
Seddon, J.M. et al. Rare variants in CFI, C3 and C9 are associated with high risk of advanced age-related macular degeneration. Nat. Genet. 45, 1366–1370 (2013).
Fritsche, L.G. et al. A large genome-wide association study of age-related macular degeneration highlights contributions of rare and common variants. Nat. Genet. 48, 134–143 (2016).
Schlötzer-Schrehardt, U., von der Mark, K., Sakai, L.Y. & Naumann, G.O. Increased extracellular deposition of fibrillin-containing fibrils in pseudoexfoliation syndrome. Invest. Ophthalmol. Vis. Sci. 38, 970–984 (1997).
Schlötzer-Schrehardt, U., Zenkel, M., Küchle, M., Sakai, L.Y. & Naumann, G.O. Role of transforming growth factor-β1 and its latent form binding protein in pseudoexfoliation syndrome. Exp. Eye Res. 73, 765–780 (2001).
Schlötzer-Schrehardt, U. Molecular pathology of pseudoexfoliation syndrome/glaucoma—new insights from LOXL1 gene associations. Exp. Eye Res. 88, 776–785 (2009).
Lucero, H.A. & Kagan, H.M. Lysyl oxidase: an oxidative enzyme and effector of cell function. Cell. Mol. Life Sci. 63, 2304–2316 (2006).
Liu, X. et al. Elastic fiber homeostasis requires lysyl oxidase–like 1 protein. Nat. Genet. 36, 178–182 (2004).
Atienza, J.M. et al. Dynamic and label-free cell-based assays using the real-time cell electronic sensing system. Assay Drug Dev. Technol. 4, 597–607 (2006).
Solly, K., Wang, X., Xu, X., Strulovici, B. & Zheng, W. Application of real-time cell electronic sensing (RT–CES) technology to cell-based assays. Assay Drug Dev. Technol. 2, 363–372 (2004).
Urcan, E. et al. Real-time xCELLigence impedance analysis of the cytotoxicity of dental composite components on human gingival fibroblasts. Dent. Mater. 26, 51–58 (2010).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Gaulton, K.J. et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat. Genet. 47, 1415–1425 (2015).
Maller, J.B. et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Liu, Y. et al. Serial analysis of gene expression (SAGE) in normal human trabecular meshwork. Mol. Vis. 17, 885–893 (2011).
Wagner, A.H. et al. Exon-level expression profiling of ocular tissues. Exp. Eye Res. 111, 105–111 (2013).
Wistow, G. et al. NEIBank: genomics and bioinformatics resources for vision research. Mol. Vis. 14, 1327–1337 (2008).
Bowes Rickman, C. et al. Defining the human macula transcriptome and candidate retinal disease genes using EyeSAGE. Invest. Ophthalmol. Vis. Sci. 47, 2305–2316 (2006).
Springelkamp, H. et al. ARHGEF12 influences the risk of glaucoma by increasing intraocular pressure. Hum. Mol. Genet. 24, 2689–2699 (2015).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Westra, H.J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Orchard, S. et al. The MIntAct project—IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 42, D358–D363 (2014).
Lee, P.H., O'Dushlaine, C., Thomas, B. & Purcell, S.M. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics 28, 1797–1799 (2012).
Barker, H.E., Cox, T.R. & Erler, J.T. The rationale for targeting the LOX family in cancer. Nat. Rev. Cancer 12, 540–552 (2012).
Braunsmann, C. et al. Evaluation of lamina cribrosa and peripapillary sclera stiffness in pseudoexfoliation and normal eyes by atomic force microscopy. Invest. Ophthalmol. Vis. Sci. 53, 2960–2967 (2012).
Schlötzer-Schrehardt, U.M., Dörfler, S. & Naumann, G.O. Corneal endothelial involvement in pseudoexfoliation syndrome. Arch. Ophthalmol. 111, 666–674 (1993).
Sauna, Z.E. & Kimchi-Sarfaty, C. Understanding the contribution of synonymous mutations to human disease. Nat. Rev. Genet. 12, 683–691 (2011).
Shabalina, S.A., Spiridonov, N.A. & Kashina, A. Sounds of silence: synonymous nucleotides as a key to biological regulation and complexity. Nucleic Acids Res. 41, 2073–2094 (2013).
Hunt, R.C., Simhadri, V.L., Iandoli, M., Sauna, Z.E. & Kimchi-Sarfaty, C. Exposing synonymous mutations. Trends Genet. 30, 308–321 (2014).
Zenkel, M., Kruse, F.E., Naumann, G.O. & Schlötzer-Schrehardt, U. Impaired cytoprotective mechanisms in eyes with pseudoexfoliation syndrome/glaucoma. Invest. Ophthalmol. Vis. Sci. 48, 5558–5566 (2007).
Korolchuk, V.I., Menzies, F.M. & Rubinsztein, D.C. Mechanisms of cross-talk between the ubiquitin–proteasome and autophagy–lysosome systems. FEBS Lett. 584, 1393–1398 (2010).
Kraft, C., Peter, M. & Hofmann, K. Selective autophagy: ubiquitin-mediated recognition and beyond. Nat. Cell Biol. 12, 836–841 (2010).
Want, A. et al. Autophagy and mitochondrial dysfunction in tenon fibroblasts from exfoliation glaucoma patients. PLoS One 11, e0157404 (2016).
Schlötzer-Schrehardt, U. & Naumann, G.O. Ocular and systemic pseudoexfoliation syndrome. Am. J. Ophthalmol. 141, 921–937 (2006).
Kivelä, T., Hietanen, J. & Uusitalo, M. Autopsy analysis of clinically unilateral exfoliation syndrome. Invest. Ophthalmol. Vis. Sci. 38, 2008–2015 (1997).
Guan, W. et al. Genome-wide association study of plasma N6 polyunsaturated fatty acids within the Cohorts for Heart and Aging Research in Genomic Epidemiology consortium. Circ Cardiovasc Genet 7, 321–331 (2014).
Lambert, J.C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat. Genet. 45, 1452–1458 (2013).
Nalls, M.A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson's disease. Nat. Genet. 46, 989–993 (2014).
Cohen, J.C., Boerwinkle, E., Mosley, T.H. Jr. & Hobbs, H.H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Jonsson, T. et al. A mutation in APP protects against Alzheimer's disease and age-related cognitive decline. Nature 488, 96–99 (2012).
Nioi, P. et al. Variant ASGR1 associated with a reduced risk of coronary artery disease. N. Engl. J. Med. 374, 2131–2141 (2016).
Barrett, J.C. et al. Genome-wide association study of ulcerative colitis identifies three new susceptibility loci, including the HNF4A region. Nat. Genet. 41, 1330–1334 (2009).
Anderson, C.A. et al. Data quality control in genetic case–control association studies. Nat. Protoc. 5, 1564–1573 (2010).
Mells, G.F. et al. Genome-wide association study identifies 12 new susceptibility loci for primary biliary cirrhosis. Nat. Genet. 43, 329–332 (2011).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Verhoeven, V.J. et al. Genome-wide meta-analyses of multiancestry cohorts identify multiple new susceptibility loci for refractive error and myopia. Nat. Genet. 45, 314–318 (2013).
Kiryluk, K. et al. Discovery of new risk loci for IgA nephropathy implicates genes involved in immunity against intestinal pathogens. Nat. Genet. 46, 1187–1196 (2014).
Bronson, P.G. et al. Common variants at PVT1, ATG13–AMBRA1, AHI1 and CLEC16A are associated with selective IgA deficiency. Nat. Genet. 48, 1425–1429 (2016).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Kooner, J.S. et al. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat. Genet. 43, 984–989 (2011).
Blake, J.A., Bult, C.J., Eppig, J.T., Kadin, J.A. & Richardson, J.E. The Mouse Genome Database: integration of and access to knowledge about the laboratory mouse. Nucleic Acids Res. 42, D810–D817 (2014).
Foo, J.N. et al. Analysis of non-synonymous-coding variants of Parkinson's disease–related pathogenic and susceptibility genes in East Asian populations. Hum. Mol. Genet. 23, 3891–3897 (2014).
Cheng, T.H. et al. Five endometrial cancer risk loci identified through genome-wide association analysis. Nat. Genet. 48, 667–674 (2016).
Browning, B.L. & Browning, S.R. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 84, 210–223 (2009).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Acknowledgements
This research is supported by the Biomedical Research Council, Agency for Science, Technology and Research, Singapore (to C.C.K.), by the Glaucoma Foundation of New York (to C.C.K.), by grants from the National Medical Research Council, Singapore NMRC/TCR/008-SERI/2013 and NMRC/STAR/0023/2014 to T.A. and NMRC/CBRG/0032/2013 to E.N.V.), by the Interdisziplinäres Zentrum für Klinische Forschung (IZKF-E23) from Germany to F.P. and U.S.-S., and by grants from the US National Institutes of Health: UM1 CA186107, R01 CA49449, R01 AR056291, R01 CA131332, P01 CA055075, R01 CA134958 and R01 EY015473 to J.L.W. For XFS cases in the US GWAS data set, genotyping services were provided through a grant to J.L.W. (HG008597) by the Center for Inherited Disease Research (CIDR). CIDR is fully funded through a federal contract from the US National Institutes of Health to The Johns Hopkins University, contract HHSN268201200008I. The Beijing, China, collection was supported by funding from the National Natural Science Foundation of China (81030016 and 81570837), the Program of Beijing Scholars (2013), the National Major Scientific and Technological Special Project for Significant New Drugs Development (2011ZX09302-007-05) and Leading Talents–High-Level Talents of the Health System of Beijing (2009-1-05) to N. Wang. We dedicate this article to the ophthalmologist Eva Forsman from Finland who passed away after diagnosing all of the exfoliation syndrome cases from Finland.
Author information
Authors and Affiliations
Contributions
C.C.K., F.P., J.L.W., T.A. and M.O. jointly conceived the project. M.C.L., U.S.-S., M.Z., D.B., Y.F.C., X.Y.N., A.W.O.C., E.N.V., S.R.G., A.S.Y.C. and Y.C. conducted functional biological experiments. G.T., R.P.I., K.P.B., Z.L., G.P., S.S., J.N.C.B., S.U., Z.Y., L. Huang, J.N.F., R.Q.S., K.S.S., P. Kraft, I.J., A.G., M.A.P.-V., A.M.H., E.N.V., C.-Y.C. and J.L.H. conducted statistical analysis. S. Raychaudhuri provided critical input on statistical analysis. Z.X., S.Q.M., H.M.S., X.Y.C., S.Q.P. and K.K.H. conducted genotyping and sequencing experiments. T.A., M.O., T.M., A.H., S.E.W., Y.S.A., A.C.O., S. Nakano, K. Mori, A.P.C., K.H., S. Manabe, S. Kazama, T. Zarnowski, K.I., M. Irkec, M.C.-P., K. Sugiyama, P. Schlottmann, S.F.L., H.L., Y.N., M.B., K.H.P., S.C.C., K.Y., J.C.Z., J.B.J., R.S.K., S.A.P., N. Kalpana, R.G., L.V., T.D., D.P.E., L.d.J.M., M.P., S. Moghimi, R.I., D.B.-H., P. Kappelgaard, B. Wirostko, S.T., D.G., K.B., W.L.G., X.C., J.S., H.J., L.J., C.Q., H.Z., X.L., B.Z., Y.-X.W., L.X., S.L., P.R., G.C., S.T., G.M., N. Weisschuh, U.H., U.-C.W.-L., C.M., P. Founti, A. Chatzikyriakidou, T.P., E.A., A.L., R.S., N.P., V.S., R.V., C. Shivkumar, N. Kobakhidze, M.R.K., A.N.B., S.Y., A.I., H.N., N. Khatibi, A.F., C.L., L.D., T.R., P. Frezzotti, D.P., E.S., P. Manunta, Y.M., K. Miyata, T.H., E.C., S.I., A.Y., M.Y., Y.K., T.O., T. Sakurai, T. Sugimoto, H.C., M.A., M. Inatani, M.M., N.G., F.M., N.Y., Y.I., M.U., C. Sotozono, J.W.J., M.S., K.H.P., J.A., M.C.-A., S.M.E., A. Rafei, V.H.K.Y., M.I.K., O.O.O., A.O.A., I.U., A.O., N.K.-A., C.T., Y.S., W.S., S.O., N.J.U., I.A., H.A., F.A., E.K.-J., U.L., I.L., V.C., R.P.G., G.S.M., S. Roy, E.D., E. Silke, A. Rao, P. Sahay, P. Fornero, O.C., D.S., T. Zompa, R.A.M., E. Souzeau, P. Mitchell, J.J.W., A.W.H., M.C., J.G.C., S.Y.A., E.L.A., A.E., V.V., G.K., R.F., S.A.A.-O., O.O., L.A.A., B.C., R.H., S.-L.H., F.A.E.-D., R.G.-S., F.M.-T., A. Salas, K.P., L. Hansapinyo, B. Wanichwecharugruang, N.K., A. Sakuntabhai, H.X.N., G.T.T.N., T.V.N., W.Z., A.B., D.S.K., M.L.H., S.D., S. Herms, S. Heegaard, M.M.N., S. Moebus, R.M.R., A.Z., T.R.C., M.R., L.A., M.G., H.G.-I., P.P.R.-C., L.F.-V.C., C.O., N.T., E.A., B.B., D.A., B.K., M.R.W., A.L.C., Y.L., P.C., L. Herndon, R.W.K., J.K., K.C., C.J.C., A. Crandall, L.M.Z., T.Y.W., M.N., S. Kinoshita, A.I.d.H., E.V., J.H.F., R.K.L., A.J.S., B.J.S., N. Wang, D.C., R.Q., T. Kivela, A. Reis, F.E.K., R.N.W., L.R.P., F.J., R.R.A., R.R., T. Kubota, S. Micheal, F.T., J.E.C., K.A.-A., M.H., J.H.K., S. Nelson, D.M., P. Sundaresan, M.D. and K.T. were involved in sample collection, phenotyping and processing. U.T., G.T. and K. Stefansson supervised, conducted and provided analysis of deCODE data. The manuscript was written by C.C.K., with critical input from T.A., T.K., U.T., J.L.W., L.R.P. and F.P. All co-authors approved the manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–23, Supplementary Tables 1–5, 8–13 and 15–17, and Supplementary Note. (PDF 6393 kb)
Supplementary Table 6
Genome-wide significant (P < 5 × 10−8) SNPs emerging from the LOXL1 deep sequencing effort after fixed-effects meta-analysis is performed. (XLSX 14 kb)
Supplementary Table 7
Details of all 63 amino acid substitutions (excluding the well-known rs3825942[G>A] for p.Gly153Asp and rs1048661[T>G] for p.Leu141Arg) that were detected from the deep resequencing of LOXL1 in 5,566 exfoliation syndrome cases and 6,279 controls from nine countries. (XLSX 14 kb)
Supplementary Table 14
LD regions around each of the seven genome-wide significant loci for exfoliation syndrome. (XLSX 13 kb)
Supplementary Data 1
Phased LOXL1 haplotypes from deep resequencing data. (XLSX 23 kb)
Supplementary Data 2
Summary statistics for the genome-wide association study. (ZIP 34845 kb)
Supplementary Data 3
INRICH pathway analysis. (XLSX 23 kb)
Rights and permissions
About this article
Cite this article
Aung, T., Ozaki, M., Lee, M. et al. Genetic association study of exfoliation syndrome identifies a protective rare variant at LOXL1 and five new susceptibility loci. Nat Genet 49, 993–1004 (2017). https://doi.org/10.1038/ng.3875
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3875
This article is cited by
-
Analysis of genetically determined gene expression suggests role of inflammatory processes in exfoliation syndrome
BMC Genomics (2023)
-
A Comparison of Genomic Advances in Exfoliation Syndrome and Primary Open-Angle Glaucoma
Current Ophthalmology Reports (2021)
-
Exfoliation syndrome: association with systemic diseases—the Maccabi glaucoma study
Graefe's Archive for Clinical and Experimental Ophthalmology (2021)
-
Association of IL-10 gene promoter polymorphisms with susceptibility to pseudoexfoliation syndrome, pseudoexfoliative and primary open-angle glaucoma
BMC Medical Genetics (2020)
-
Glucocorticoid-induced cell-derived matrix modulates transforming growth factor β2 signaling in human trabecular meshwork cells
Scientific Reports (2020)