Abstract
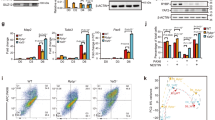
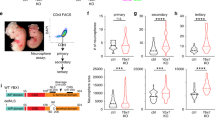
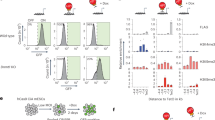
Temporal regulation of embryonic neurogenesis is controlled by hypostable transcription factors. The mechanism of the process is unclear. Here we show that the RNase III Drosha and DGCR8 (also known as Pasha), key components of the microRNA (miRNA) microprocessor, have important functions in mouse neurogenesis. Loss of microprocessor in forebrain neural progenitors resulted in a loss of stem cell character and precocious differentiation whereas Dicer deficiency did not. Drosha negatively regulated expression of the transcription factors Neurogenin 2 (Ngn2) and NeuroD1 whereas forced Ngn2 expression phenocopied the loss of Drosha. Neurog2 mRNA contains evolutionarily conserved hairpins with similarities to pri-miRNAs, and associates with the microprocessor in neural progenitors. We uncovered a Drosha-dependent destabilization of Neurog2 mRNAs consistent with microprocessor cleavage at hairpins. Our findings implicate direct and miRNA-independent destabilization of proneural mRNAs by the microprocessor, which facilitates neural stem cell (NSC) maintenance by blocking accumulation of differentiation and determination factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Eulalio, A., Huntzinger, E. & Izaurralde, E. Getting to the root of miRNA-mediated gene silencing. Cell 132, 9–14 (2008).
Filipowicz, W., Bhattacharyya, S.N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat. Rev. Genet. 9, 102–114 (2008).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Lai, E.C. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat. Genet. 30, 363–364 (2002).
Gregory, R.I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Han, J. et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Denli, A.M., Tops, B.B., Plasterk, R.H., Ketting, R.F. & Hannon, G.J. Processing of primary microRNAs by the microprocessor complex. Nature 432, 231–235 (2004).
Chendrimada, T.P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005).
Okamura, K., Hagen, J.W., Duan, H., Tyler, D.M. & Lai, E.C. The mirtron pathway generates microRNA-class regulatory RNAs in Drosophila. Cell 130, 89–100 (2007).
Ruby, J.G., Jan, C.H. & Bartel, D.P. Intronic microRNA precursors that bypass Drosha processing. Nature 448, 83–86 (2007).
Babiarz, J.E., Ruby, J.G., Wang, Y., Bartel, D.P. & Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 22, 2773–2785 (2008).
Yang, J.S. & Lai, E.C. Alternative miRNA biogenesis pathways and the interpretation of core miRNA pathway mutants. Mol. Cell 43, 892–903 (2011).
Ender, C. et al. A human snoRNA with microRNA-like functions. Mol. Cell 32, 519–528 (2008).
Cheloufi, S., Dos Santos, C.O., Chong, M.M. & Hannon, G.J. A dicer-independent miRNA biogenesis pathway that requires Ago catalysis. Nature 465, 584–589 (2010).
Cifuentes, D. et al. A novel miRNA processing pathway independent of Dicer requires Argonaute2 catalytic activity. Science 328, 1694–1698 (2010).
Yang, J.S. et al. Conserved vertebrate mir-451 provides a platform for Dicer-independent, Ago2-mediated microRNA biogenesis. Proc. Natl. Acad. Sci. USA 107, 15163–15168 (2010).
De Pietri Tonelli, D. et al. miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development 135, 3911–3921 (2008).
Babiarz, J.E. et al. A role for noncanonical microRNAs in the mammalian brain revealed by phenotypic differences in Dgcr8 versus Dicer1 knockouts and small RNA sequencing. RNA 17, 1489–1501 (2011).
Georgi, S.A. & Reh, T.A. Dicer is required for the transition from early to late progenitor state in the developing mouse retina. J. Neurosci. 30, 4048–4061 (2010).
Kaneko, H. et al. DICER1 deficit induces Alu RNA toxicity in age-related macular degeneration. Nature 471, 325–330 (2011).
Anthony, T.E., Klein, C., Fishell, G. & Heintz, N. Radial glia serve as neuronal progenitors in all regions of the central nervous system. Neuron 41, 881–890 (2004).
Gotz, M. & Huttner, W.B. The cell biology of neurogenesis. Nat. Rev. Mol. Cell Biol. 6, 777–788 (2005).
Kageyama, R., Niwa, Y., Shimojo, H., Kobayashi, T. & Ohtsuka, T. Ultradian oscillations in Notch signaling regulate dynamic biological events. Curr. Top. Dev. Biol. 92, 311–331 (2010).
Gohlke, J.M. et al. Characterization of the proneural gene regulatory network during mouse telencephalon development. BMC Biol. 6, 15 (2008).
Kageyama, R., Ohtsuka, T., Shimojo, H. & Imayoshi, I. Dynamic regulation of Notch signaling in neural progenitor cells. Curr. Opin. Cell Biol. 21, 733–740 (2009).
Masamizu, Y. et al. Real-time imaging of the somite segmentation clock: revelation of unstable oscillators in the individual presomitic mesoderm cells. Proc. Natl. Acad. Sci. USA 103, 1313–1318 (2006).
Guillemot, F. Spatial and temporal specification of neural fates by transcription factor codes. Development 134, 3771–3780 (2007).
Miyata, T., Kawaguchi, D., Kawaguchi, A. & Gotoh, Y. Mechanisms that regulate the number of neurons during mouse neocortical development. Curr. Opin. Neurobiol. 20, 22–28 (2010).
Makeyev, E.V., Zhang, J., Carrasco, M.A. & Maniatis, T. The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 27, 435–448 (2007).
Lutolf, S., Radtke, F., Aguet, M., Suter, U. & Taylor, V. Notch1 is required for neuronal and glial differentiation in the cerebellum. Development 129, 373–385 (2002).
Imayoshi, I., Sakamoto, M., Yamaguchi, M., Mori, K. & Kageyama, R. Essential roles of Notch signaling in maintenance of neural stem cells in developing and adult brains. J. Neurosci. 30, 3489–3498 (2010).
Han, J. et al. Posttranscriptional crossregulation between Drosha and DGCR8. Cell 136, 75–84 (2009).
Salomoni, P. & Calegari, F. Cell cycle control of mammalian neural stem cells: putting a speed limit on G1. Trends Cell Biol. 20, 233–243 (2010).
Nguyen, L. et al. p27kip1 independently promotes neuronal differentiation and migration in the cerebral cortex. Genes Dev. 20, 1511–1524 (2006).
Cheung, T.H. et al. Maintenance of muscle stem-cell quiescence by microRNA-489. Nature 482, 524–528 (2012).
Wang, Y. & Blelloch, R. Cell cycle regulation by MicroRNAs in embryonic stem cells. Cancer Res. 69, 4093–4096 (2009).
Gantier, M.P. et al. Analysis of microRNA turnover in mammalian cells following Dicer1 ablation. Nucleic Acids Res. 39, 5692–5703 (2011).
Kawase-Koga,, Y. et al. RNaase-III enzyme Dicer maintains signaling pathways for differentiation and survival in mouse cortical neural stem cells. J. Cell Sci. 123, 586–594 (2010).
Guillemot, F. Cellular and molecular control of neurogenesis in the mammalian telencephalon. Curr. Opin. Cell Biol. 17, 639–647 (2005).
Nieto, M., Schuurmans, C., Britz, O. & Guillemot, F. Neural bHLH genes control the neuronal versus glial fate decision in cortical progenitors. Neuron 29, 401–413 (2001).
Ganesan, G. & Rao, S.M. A novel noncoding RNA processed by Drosha is restricted to nucleus in mouse. RNA 14, 1399–1410 (2008).
Chong, M.M. et al. Canonical and alternate functions of the microRNA biogenesis machinery. Genes Dev. 24, 1951–1960 (2010).
Karginov, F.V. et al. Diverse endonucleolytic cleavage sites in the mammalian transcriptome depend upon microRNAs, Drosha, and additional nucleases. Mol. Cell 38, 781–788 (2010).
Shenoy, A. & Blelloch, R. Genomic analysis suggests that mRNA destabilization by the microprocessor is specialized for the auto-regulation of Dgcr8. PLoS ONE 4, e6971 (2009).
Shimojo, H., Ohtsuka, T. & Kageyama, R. Oscillations in notch signaling regulate maintenance of neural progenitors. Neuron 58, 52–64 (2008).
Heng, J.I. et al. Neurogenin 2 controls cortical neuron migration through regulation of Rnd2. Nature 455, 114–118 (2008).
Kim, V.N., Han, J. & Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139 (2009).
Kadener, S. et al. Genome-wide identification of targets of the drosha-pasha/DGCR8 complex. RNA 15, 537–545 (2009).
Noctor, S.C., Martinez-Cerdeno, V., Ivic, L. & Kriegstein, A.R. Cortical neurons arise in symmetric and asymmetric division zones and migrate through specific phases. Nat. Neurosci. 7, 136–144 (2004).
Heo, I. et al. TUT4 in concert with Lin28 suppresses microRNA biogenesis through pre-microRNA uridylation. Cell 138, 696–708 (2009).
Basak, O. & Taylor, V. Identification of self-replicating multipotent progenitors in the embryonic nervous system by high Notch activity and Hes5 expression. Eur. J. Neurosci. 25, 1006–1022 (2007).
Harfe, B.D., McManus, M.T., Mansfield, J.H., Hornstein, E. & Tabin, C.J. The RNaseIII enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb. Proc. Natl. Acad. Sci. USA 102, 10898–10903 (2005).
Chong, M.M., Rasmussen, J.P., Rudensky, A.Y. & Littman, D.R. The RNAseIII enzyme Drosha is critical in T cells for preventing lethal inflammatory disease. J. Exp. Med. 205, 2005–2017 (2008).
Stump, G. et al. Notch1 and its ligands Delta-like and Jagged are expressed and active in distinct cell populations in the postnatal mouse brain. Mech. Dev. 114, 153–159 (2002).
Giachino, C., Basak, O. & Taylor, V. Isolation and manipulation of mammalian neural stem cells in vitro. Methods Mol. Biol. 482, 143–158 (2009).
Nyfeler, Y. et al. Jagged1 signals in the postnatal subventricular zone are required for neural stem cell self-renewal. EMBO J. 24, 3504–3515 (2005).
Acknowledgements
We thank N. Kim (Seoul National University) for Drosha constructs, F. Guillemot (UK National Institute for Medical Research) for Neurog2 and shRNA constructs, and P. Herrare for beta2/NeuroD promoter, members of the Taylor laboratory for helpful discussions, and F. Sager for excellent technical assistance. P.K. and S.L. were PhD students of the International Max Planck Research School Molecular and Cellular Biology and of the Faculty of Biology, University of Freiburg, M.A.V. is a PhD student of the Department of Biomedical Science, University of Sheffield. This work was supported by the Deutsche Forschungsgemeinschaft (TA–310–1, TA–310–2 and SFB592 to V.T.).
Author information
Authors and Affiliations
Contributions
P.K. and M.A.V. performed the experiments, analyzed the data and wrote the paper, S.L. cloned the Hes5∷Cre construct and performed neurosphere assays, M.M. analyzed the miRNA array data, M.M.W.C. and D.R.L. provided the floxed Drosha mice, G.M.H. and S.A.W. helped with the CLIP analysis. V.T. designed the project, performed experiments, analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1–3 (PDF 8509 kb)
Rights and permissions
About this article
Cite this article
Knuckles, P., Vogt, M., Lugert, S. et al. Drosha regulates neurogenesis by controlling Neurogenin 2 expression independent of microRNAs. Nat Neurosci 15, 962–969 (2012). https://doi.org/10.1038/nn.3139
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3139
This article is cited by
-
Cathepsin B S-nitrosylation promotes ADAR1-mediated editing of its own mRNA transcript via an ADD1/MATR3 regulatory axis
Cell Research (2023)
-
Non-canonical functions of the microprocessor
Nature Reviews Molecular Cell Biology (2021)
-
HOTAIRM1 regulates neuronal differentiation by modulating NEUROGENIN 2 and the downstream neurogenic cascade
Cell Death & Disease (2020)
-
Dgcr8 knockout approaches to understand microRNA functions in vitro and in vivo
Cellular and Molecular Life Sciences (2019)
-
Neuronal activity regulates DROSHA via autophagy in spinal muscular atrophy
Scientific Reports (2018)