Abstract
Cancer stem cells (CSC) appear to have increased metastatic potential, but mechanisms underlying this are poorly defined. Here we show that VEGFA induction of Sox2 promotes EMT and tumor metastasis. In breast lines and primary cancer culture, VEGFA rapidly upregulates SOX2 expression, leading to SNAI2 induction, EMT, increased invasion and metastasis. We show Sox2 downregulates miR-452, which acts as a novel metastasis suppressor to directly target the SNAI2 3′-untranslated region (3′-UTR). VEGFA stimulates Sox2- and Slug-dependent cell invasion. VEGFA increases lung metastasis in vivo, and this is abrogated by miR-452 overexpression. Furthermore, SNAI2 transduction rescues metastasis suppression by miR-452. Thus, in addition to its angiogenic action, VEGFA upregulates Sox2 to drive stem cell expansion, together with miR-452 loss and Slug upregulation, providing a novel mechanism whereby cancer stem cells acquire metastatic potential. Prior work showed EMT transcription factor overexpression upregulates CSC. Present work indicates that stemness and metastasis are a two-way street: Sox2, a major mediator of CSC self-renewal, also governs the metastatic process.
Similar content being viewed by others
Introduction
VEGFA is a cytokine that regulates vascular development during embryogenesis and the formation of new blood vessels from pre-existing vascular networks.1, 2, 3 VEGFA, secreted by cancer and stromal cells, stimulates endothelial cell invasion and vessel formation.4 Without new blood vessel formation, tumor size is restrained due to limited nutrient and oxygen supply. VEGFA is expressed in a variety of tumors and its overexpression is associated with poor prognosis and death from metastasis.5, 6, 7 VEGFA functions are not restricted to vasculogenesis and angiogenesis.8 Autocrine VEGFA cooperates with EGFR to drive tumor development9 and VEGFA has also been shown to drive tumor metastasis.4, 10, 11 Indeed, patients with metastatic breast cancer have higher circulating VEGFA levels than those without metastasis.12
Bevacizumab, a humanized monoclonal antibody that targets VEGFA, has been applied for the treatment of breast and other malignancies. However, trials in metastatic breast cancer have yielded variable results and the role of this drug is controversial.13, 14, 15 Recent work sheds light on the limited results of bevacizumab in most cancers. Hypoxia caused by inhibition of angiogenesis, upregulates VEGFA expression, contributing to aggressive disease recurrence.16, 17 VEGFA was recently shown to increase tumor-initiating stem cell abundance in skin18 and breast cancers,19, 20 and in glioblastoma.21, 22 The high local VEGFA induced by hypoxia following bevacizumab treatment would thus also promote expansion of the tumor cell subset with the greatest ability to initiate and disseminate tumors.
Cancer stem cells (CSCs) show greater motility and metastatic potential than the bulk tumor cell population and have been postulated to be drivers of tumor metastasis,23, 24, 25 but the mechanisms underlying this are not fully characterized. Metastasis requires cell invasion and escape from the primary tumor into the vasculature followed by colonization of secondary sites. Tumor invasion and intravasation are enabled by the epithelial to mesenchymal transition (EMT), a process in which epithelial cells lose polarity and intracellular adhesion, and acquire motility and invasiveness.26, 27, 28, 29 The EMT is regulated by diverse molecular networks including TGF-β, Notch, Wnt, Hedgehog and NF-κB signaling pathways, all of which have central roles in cancer invasion and metastasis.30 Downregulated expression of the cell adhesion molecule, E-cadherin, is critical for acquisition of the EMT phenotype and tumor invasion.31 Many EMT transcription factors repress CDH1, the gene encoding E-cadherin, directly or indirectly. Snail,32, 33 Slug,34 Zeb135 and Zeb236 can bind the CDH1 promoter and repress its transcription, whereas other factors such as Twist,37 Goosecoid38 and fork-head box protein C2 (FOXC2)39 repress CDH1 indirectly. Slug, whose expression correlates strongly with loss of E-cadherin, is an important EMT mediator in breast cancer cell models.40
The EMT program has been linked to the initiation and/or maintenance of CSCs. Enforced expression of EMT transcription factors has been shown to increase cancer stem cell abundance, and stem-like cells exhibit EMT properties such as increased expression of mesenchymal markers and EMT transcription factors, suggesting a link between cancer stem cells and the EMT process.41, 42 However, pathways governing the relationship between cancer stem cells and EMT are not fully defined. VEGFA not only increases the tumor-initiating stem cell population in several different murine and human cancer models,18, 19, 20, 21, 22 but is also known to induce EMT and metastasis.43, 44, 45 Our prior work showed that VEGFA rapidly activates STAT3 to induce SOX2 and increase the CSC population in breast and lung models.19 Here, we investigated whether upregulation of Sox2 by VEGFA might have a role not only in CSC expansion but also contribute to the activation of EMT and metastasis.
MicroRNAs (miRNAs) are small, noncoding RNAs that regulate transcriptional and post-transcriptional gene expression. Approximately 70% of all genes are regulated by miRNA in eukaryotes.46, 47 miRNAs carry out important functions in development, differentiation, cell cycle progression and apoptosis. Mature miRNAs bind complementary sequences in the 3′-untranslated region (3′-UTR) of target genes and repress gene expression by inducing mRNA degradation and/or translational inhibition.48, 49 In cancers, miRNA expression is deregulated by amplification, deletion, mutation and epigenetic silencing.50, 51, 52 Many miRNAs act as either oncogenes or tumor suppressors to regulate malignant transformation and metastatic progression.52 MiRNAs modulate the metastatic process by targeting metastasis suppressor genes or by repressing metastasis promoting genes.53 Several miRNAs regulate EMT transcription factors including Zeb1, Zeb2 and Snail.54, 55, 56 Indeed, several miRNAs that target EMT transcription factors, such as miR-200 that targets Zeb154, 57, 58, 59 and miR-34 that targets Snail,60 also repress cancer stem cell self-renewal.47
Here, we identify a novel pathway in which Sox2, a stem cell driver upregulated by VEGFA,19 contributes to the activation of EMT. VEGFA leads to induction of the stem cell transcription factor gene SOX2. Sox2, in turn, mediates repression of miR-452, which is shown to directly target the 3′-UTR of SNAI2, leading to EMT and breast cancer metastasis.
Results
VEGFA induces EMT and an increase in motility and invasion in breast models
In addition to its angiogenic effects, VEGFA promotes cancer stem cell expansion.19 VEGFA also drives cancer invasion and metastasis in experimental models.43, 45 Cancer stem cell expansion is linked to, and potentially driven by, upregulation of EMT transcription factors,41, 42 but whether stem cell drivers can also promote EMT has not been fully investigated. To investigate whether VEGFA-mediated CSC expansion might also be linked to EMT activation and metastasis, we tested the effect of VEGFA on motility and invasion in aggressive ER-negative breast cancer models. Since our earlier work showed a prolonged 7-day exposure to VEGFA caused an irreversible increase in stem-like cells,19 all experiments used 7 days of VEGFA (10 ng/ml), unless otherwise indicated. VEGFA-treated MDA-MB-231 showed faster migration on wound-healing assays and increased matrigel invasion compared with controls (Figures 1a and b). Results were validated in an ER, PR and Her2 (triple) negative primary breast cancer-derived line, SUM149PT (Supplementary Figure S1a and b).
VEGFA induces EMT and an increase in motility and invasion. (a) MDA-MB-231 was pre-treated for 24 h or 7 days ±10 ng/ml VEGFA followed by scratch wounding of a confluent monolayer. Photomicrographs were taken at 0 and 12 h and mean migration±s.e.m. graphed versus controls (C). (b) MDA-MB-231 pre-treated for 24 h or 7 days±VEGFA were recovered for real-time matrigel invasion assays. Data graphed represent mean +/− s.e.m. for 3 replicates. (c–e) EMT marker expression was compared by QPCR after 7 days ±10 ng/ml VEGFA (V) in MDA-MB-231 (c), SUM149PT (d) and MCF12A (e). All graphs show mean±s.e.m. Mean values were compared by Student's t test. * denotes P<0.05 for test versus control.
Acquisition of an EMT phenotype is critical for metastasis. Mesenchymal markers (vimentin, fibronectin and N-cadherin) were upregulated by VEGFA in MDA-MB-231 and SUM149PT lines and in the immortal but not malignantly transformed human mammary epithelial line, MCF12A. Epithelial markers, including one or both of E-cadherin and Zo-1, were decreased in all three cell lines (Figures 1c–e), compatible with a VEGFA-induced EMT.
VEGFA increases motility and invasion by upregulating Slug in breast cancer cells
Expression of major EMT-driving transcription factors (EMT-TFs), Slug, Snail, Zeb1 and Zeb2, was induced over a 7-day VEGFA exposure. The temporal patterns of EMT-TF upregulation during prolonged VEGFA exposure for MDA-MB-231 and MCF12A are shown in Supplementary Figures S2A and B. Of these, SNAI2, which encodes Slug, was the most strongly induced after 7 days, and was thus investigated further (Figures 2a and b). SNAI2 knockdown (Figure 2c) prevented VEGFA-mediated increases in cell motility and invasion (Figures 2d and e), indicating VEGFA increases migration and invasion via Slug. VEGFA also upregulated SNAI2 expression in SUM149PT cells, and SNAI2 knockdown inhibited VEGF-driven invasion in this second model (Figures 2f and g).
VEGFA increases motility and invasion via slug induction in breast cancer cells. (a and b) SNAI1, SNAI2, ZEB1 and ZEB2 expression levels were assayed by QPCR at indicated times after 10 ng/ml VEGFA in MDA-MB-231 (a) and in MCF12A (b). (c–e) The MDA-MB-231 cells were transduced with siSNAI2 or control siRNA (C) for 48 h. The cells were then treated ±10 ng/ml VEGFA for 7 days and then recovered for assays of SNAI2 and Slug expression by QPCR and western, respectively (c) and assays of migration (d) and matrigel invasion (e). (f and g) The SUM149PT cells were transduced with siSNAI2 or control siRNA for 48 h then treated ±10 ng/ml VEGFA for 7 days followed by assays of SNAI2 and Slug expression by QPCR and western, respectively (f) and by real-time matrigel invasion assay (g). All graphs show mean±s.e.m. Mean values were compared by Student's t test. * denotes P<0.05 for test versus control.
Sox2 is required for VEGFA-driven Slug upregulation and for increased motility and invasion
Sox2 drives self-renewal in both embryonic stem cells and in several cancer stem cell models61, 62 and is a key mediator of VEGFA-driven CSC expansion.19 CSC are thought to be drivers of tumor metastasis and exhibit greater motility and metastatic potential than bulk tumor cells.23, 24, 63 Notably, EMT-TF overexpression leads to expansion of cells with stem cell characteristics.41, 42 Here we tested whether the reverse is also true and whether the embryonic stem cell factor, Sox2, might mediate VEGFA-driven EMT. Upregulation of SOX2 expression by VEGFA occurs rapidly, within 1 h in MDA-MB-231 and SUM149PT (Figure 3a) and remains elevated for at least 7 days.19 SOX2 induction precedes that of SNAI2 by several days. SNAI2/Slug upregulation by VEGFA was prevented by SOX2 knockdown (Figures 3b and c, and Supplementary Figure S3) and Sox2 was also required for VEGFA-mediated increase in cell motility and invasion (Figure 3d) in MDA-MB-231. Findings were validated in the SUM149PT line (Figures 3e and f). Moreover, SOX2 overexpression was sufficient to increase Slug expression (Figure 3g), invasion and migration in the absence of VEGFA stimulation (Figure 3h). Thus, the rapid VEGFA-STAT3-mediated induction of SOX219 not only precedes, but is required for that of SNAI2 and for the increased migration and invasion following VEGFA exposure in both MDA-MB-231 and SUM149PT.
Sox2 is required for VEGFA-driven increases in slug, motility and invasion. (a) MDA-MB-231 (top) and SUM149PT (bottom) were treated ±10 ng/ml VEGFA and SOX2 expression quantitated by QPCR. (b) siSOX2 or control siRNA (C) transfected MDA-MB-231 cells were treated ±10 ng/ml VEGFA for 7 days followed by QPCR for SNAI2 expression. (c) MDA-MB-231 transduced with either siSOX2 or control siRNA were treated±VEGFA for 7 days before western blot for Sox2 and Slug. (d) MDA-MB-231 transduced with either siSOX2 or control siRNA for 48 h were treated±VEGFA for 7 days followed by assays of migration (top) and matrigel invasion (bottom). (e and f) SUM149PT cells transduced with siSOX2 or control siRNA for 48 h were treated±VEGFA for 7 days followed by QPCR for SNAI2 (e) and real-time matrigel invasion assay (f). (g) MDA-MB-231 cells were transfected with Sox2 or control vector (C), and Sox2 and Slug expression were assayed by western. (h) MDA-MB-231 transduced with Sox2 overexpression or control vector and migration (top) and matrigel invasion (bottom) were assayed using Xcelligence. All graphs show mean±s.e.m. Mean values were compared by Student's t test. * denotes P<0.05 for test versus control or versus indicated condition.
Although SOX2 siRNA significantly decreased VEGFA-induced SNAI2, SOX2 siRNA-transduced cells still showed a modest but significant increase of SNAI2 by VEGFA (Figure 3b right). This may reflect incomplete SOX2 knockdown by transient siRNASOX2 transfection (Figure 3b, left). It is also possible that additional mechanisms govern VEGFA action on Slug.
miR-452 downregulation is required for VEGFA-mediated increases in Slug and invasion
The SNAI2 promoter contains a single, putative Sox2 consensus motif, but Sox2 binding to this motif was not detected after VEGFA treatment. Notably, several studies of global Sox2 DNA binding by ChIP-sequencing also failed to show stable binding of Sox2 to the SNAI2 promoter.64, 65, 66, 67 These findings, and our observation that Sox2 is upregulated by VEGFA within hours (Figure 3a), but SNAI2 expression only increases several days later (Figures 2a and b) suggested that Sox2-mediated SNAI2 induction is indirect.
Since Sox2 is known to induce several miRNAs to drive stem cell self-renewal, we investigated whether a miRNA-driven mechanism might govern Slug upregulation. A miRNA screen of MDA-MB-231 cells before and after VEGFA treatment was performed. Among over 700 miRNAs, 47 miRNAs were significantly downregulated by VEGFA (see Supplementary Figure S4). The miRNA target prediction software TargetScan (Version 6.2) was used to identify miRNAs decreased by VEGFA that could potentially target SNAI2. Of four potential candidates, miR-452 had the highest probability score for targeting SNAI2 and was investigated herein. A second miRNA target prediction database (microT-CDS version 5.0) verified miR-452 as a putative regulator of SNAI2 expression. VEGFA downregulated miR-452 in both MDA-MB-231 and SUM149PT. miR-452 was decreased within 6–12 h and reduced levels persisted after 7 days of VEGFA treatment (Figure 4a, bottom panel). To further validate this finding, DT22, an early passage culture derived from a triple-negative primary human breast cancer was tested. This culture has been extensively validated and its gene expression and tumor marker profiles resemble those of the cancer from which it was derived.68 Prolonged exposure of DT22 to VEGFA over 7 days also led to miR-452 loss (Figure 4a, top right). SOX2 knockdown abrogated the VEGFA-driven loss of miR-452, indicating that Sox2 is required for miR-452 downregulation by VEGFA (Figure 4b). SOX2 overexpression (Figure 3g) also reduced miR-452 in MDA-MB-231 cells (Figure 4c). To test if downregulation of miR-452 is required for VEGFA to increase invasion, miR-452 was transduced into MDA-MB-231 and stable clones derived (Supplementary Figure S5, top panel). The miR-452 overexpression abrogated VEGFA-driven SNAI2 upregulation (Figure 4d) and prevented the VEGFA-driven increase in matrigel invasion (Figure 4f). Furthermore, the inhibition of miR-452 by transfection of a miR-452 antagomir increased SNAI2 expression (Figure 4g) and was sufficient to increase matrigel invasion (Figure 4h). Thus, miR-452 is required for Sox2-driven Slug upregulation and is critical for VEGFA-driven cell motility and invasion.
VEGFA- and Sox2-driven miR-452 downregulation mediates Slug upregulation and breast cancer cell invasion, and miR-452 directly targets the SNAI2 3′-UTR. (a) miR-452 expression (QPCR) was compared±VEGFA for 7 days in MDA-MB-231, SUM149PT and DT22 primary breast cancer culture (top) miR-452 expression levels were assayed by QPCR at indicated times in MDA-MB-231 (bottom). (b) The siSOX2 or control siRNA (C) transduced MDA-MB-231 cells were treated±VEGFA for 7 days and miR-452 expression assayed. (c) MDA-MB-231 was transduced with Sox2 or control vector, and miR-452 expression was assayed by QPCR. (d–f) miRNA control or miR-452 vector transduced cells were treated±VEGFA for 7 days before assays of Slug expression by western (d) and SNAI2 expression by QPCR (e) and real-time matrigel invasion (f). (g and h) MDA-MB-231 were transfected with miR-452 antagomir (inhibitor) or antagomir control followed by assays of SNAI2 by QPCR (g) and of invasion as above (h). (i) Sequence alignment of human miR-452 seed regions with SNAI2 3′-UTR. (j) 293 T and MDA-MB-231 were transfected with SNAI2 3′-UTR luciferase reporter together with miR-452 precursor or control miRNA plasmid and luciferase activity assayed after 48 h. (k) SNAI2 3′-UTR luciferase reporter plasmid bearing mutations in all three putative miR-452 binding sites show loss of luciferase regulation by transfected miR-452 precursor plasmid after 48 h. (l) miR-452 overexpressing MDA-MB-231 was transduced with either human SNAI2 or control vector, then treated±VEGFA for 7 days followed by real-time matrigel invasion assay. All graphs show mean±s.e.m. Mean values were compared by Student's t test. * denotes P<0.05 for test versus control or versus indicated condition.
miR-452 directly targets the SNAI2 3′-UTR to decrease Slug
Stable overexpression of miR-452 decreased SNAI2 expression (Figure 4e and Supplementary Figure S5, top) and miR-452 antagomir transfection increased SNAI2 levels in MDA-MB-231 (Figure 4g and Supplementary Figure S5, bottom). miRNAs commonly regulate mRNA expression by binding to the 3′-UTR. There are three putative miR-452 binding sites within the 3′-UTR of SNAI2 (Figure 4i). To investigate whether miR-452 directly targets the 3′-UTR of SNAI2 to repress Slug expression, a reporter assay was performed using the 3′-UTR of SNAI2 to drive luciferase expression. 293T and MDA-MB-231 cells were transfected with a human SNAI2 3′-UTR luciferase reporter plasmid together with plasmids encoding either the miR-452 precursor or control miRNA, and luciferase activity was measured after 48 h. miR-452 transfection significantly reduced luciferase activity, indicating miR-452 targets the SNAI2 3′-UTR to repress Slug expression (Figure 4j).
A mutant SNAI2 3′-UTR luciferase vector was constructed in which all three putative miR-452 binding sites were mutationally disrupted. When this mutated vector was co-transfected into 293T cells with the miR-452 precursor plasmid, luciferase activity was not impaired. Thus one or more of these sites is required for miR-452 to inhibit SNAI2 expression (Figure 4k).
To test whether SNAI2 overexpression could rescue the inhibitory effect of miR-452 on VEGFA-induced invasion, miR-452 overexpressing MDA-MB-231 cells were transduced with either control vector or human SNAI2 cDNA lacking the 3′-UTR region. As noted above, miR-452 overexpression abrogated the increased invasion by VEGFA (Figure 4f). Overexpression of this 3′-UTR-deficient SNAI2 vector rescued the inhibitory effect of miR-452 on cell invasion (Figure 4l), consistent with the notion that miR-452 targets SNAI2. Thus, VEGFA-mediated miR-452 downregulation is critical for the induction of SNAI2 and for Slug action on cell motility and invasion.
Repression of miR-452 is required for VEGFA-dependent cancer metastasis in vivo
Although VEGFA has been shown to drive cancer metastasis,45, 69 mechanisms thereof are largely unknown. To test whether VEGFA drives metastasis in vivo through regulation of miR-452 and Slug, MDA-MB-231 cells were pre-treated with VEGFA for 1 week before injection by tail vein into nude mice, without further VEGFA treatment after tumor cell injection. The animals were monitored by in vivo imaging system. VEGFA pre-treated cells gave rise to a significant increase in lung tumor establishment as measured by tumor bioluminescence on in vivo imaging system over the next 5 weeks compared with mock-treated cells (Figures 5a–d). VEFG also increased green fluorescence of lung tumors (Figures 5e). miR-452 overexpressing cells failed to respond to VEGFA, and showed no effect of VEGFA pre-treatment on tumor metastasis. Notably, transduction of a SNAI2 cDNA vector lacking the 3′-UTR into miR-452 overexpressing cells overcame the effect of miR-452 to inhibit VEGFA-stimulated metastasis in vivo (Figures 5a–e). VEGFA treatment does not affect MDA-MB-231 cell cycle progression or population growth.19 Overexpression of miR-452 and SNAI2 did not change cell proliferation (Supplementary Figure S6), thus differences in the metastatic tumor burdens of each group are not due to differences in growth rates.
miR-452 repression is required for VEGFA-induced cancer metastasis in vivo. (a) MDA-MB-231-luc expressing the indicated vectors were pre-treated ±VEGFA and injected via tail vein into nude mice as described in the ‘Materials and methods’ section. Representative bioluminescence images of tumor bearing mice at 0 and 5 weeks are shown. The color scale depicts photon flux (photons/s) from xenografted mice. (b) Mean bioluminescence/time of lung metastasis in xenografted mice, graphed as normalized photon flux/time. (c) Mean bioluminescence at 5 weeks (d) Representative images and photomicrographs of lung tumors from indicated groups. (e) Representative immunofluorescence images (× 4) of GFP-positive metastasis observed immediately ex vivo. (f) Mean lung weights. All graphs show mean±s.e.m. Mean values were compared by Student's t test. * denotes P<0.05 for test versus control or versus indicated condition.
The animals injected with VEGFA pre-treated cells showed extensive areas of confluent tumor growth in the lungs on microscopic analysis, precluding accurate enumeration of tumor nodules (Figures 5d). As a second measure of lung tumor burden, lung weights were measured. The lung weights were significantly increased in animals injected with VEGFA-pretreated cells, whereas those of mice injected with VEGFA-treated miR-452 overexpressing cells were not increased compared with controls. Finally, SNAI2 transduction into the miR-452 overexpressing cells yielded similar lung weights to those in the VEGFA-treated group (Figure 5f). Thus miR-452 repression is required not only for VEGFA-dependent Slug upregulation in vitro, but also for increased cancer metastasis in vivo.
VEGFA, SOX2, SNAI2, miR-452 and GABRE expression and prognosis in primary breast cancers
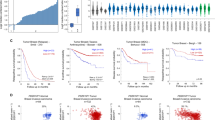
Our in vitro and in vivo models suggest a mechanism in which VEGFA induces EMT and metastasis by activating Sox2, resulting in de-repression of SNAI2 through loss of miR-452 (Figure 6a). To validate our findings in vivo, we tested whether high VEGFA alone or together with high SOX2, SNAI2 and decreased miR-452 expression might identify prognostic subsets of primary human breast cancers. miR-452 is expressed as an intronic transcript from the GABRE gene locus.70 Pearson’s correlation analysis of two independent breast cancer data sets, the METABRIC and Enerly data sets,71, 72 respectively, showed that miR-452 expression correlated strongly with that of its parent transcript, GABRE (R2 values of 0.484 and 0.786 in the METABRIC and Enerly data sets, respectively; Figure 6b), indicating GABRE can be used as a surrogate for miR-452 expression in data sets, such as KM Plotter, that lack microRNA data.
Prognostic value of VEGFA, SOX2, SNAI2, miR-452 and GABRE expression in breast cancer patients. (a) Model of mechanism by which VEGFA increases breast cancer invasion and metastasis through Sox2, miR-452 and Slug. (b) Correlation of miR-452 and GABRE expression in the METABRIC (left) and Enerly (right) primary breast cancer data sets; R2 values for Pearson's correlation are indicated. (c and d) Kaplan–Meier plots of distant metastasis-free survival (DMFS) of all breast cancer patients (c) and those with ER-negative breast cancer (d), stratified by VEGFA expression alone or with sequential inclusion of downstream pathway genes (SOX2, GABRE, SNAI2); data were analyzed and plots generated using the KM Plotter web tool (http://kmplot.com/analysis/). Patients with available clinical data: DMFS, n=1609. (e) Kaplan–Meier survival plot of disease-specific survival (DSS) of breast cancer patients from the METABRIC data set classified by top quartile VEGFA expression (left) or the top 25% patients with the highest mean expression of both VEGFA and SOX2 were classified as ‘high’ (right). Hazard ratios, HR (95% CI), were determined using Univariate Cox Proportional Hazards analysis and the P-values from the log-rank test are shown in each graph.
As Sox2 regulates SNAI2 expression via miR-452, we next tested whether high VEGFA expression (top quartile) alone or in combination with high SOX2, high SNAI2 and the lowest quartile GABRE/miR-452 expression was associated with distant metastasis-free breast cancer survival (DMFS) in the KM Plotter data set. Differences between groups are shown by graphed Kaplan–Meier curves and hazard ratios (HRs) from Univariate Cox Proportional Hazards analysis. Log-rank comparison of outcome curves was done and P-values are presented in each graph. Of 1609 primary breast cancers in the KM Plotter data set, those with high VEGFA expression alone showed significantly poorer DMFS (n=1609, HR (95% confidence interval [CI])=1.45 [1.17–1.81]; P=0.00082; Figure 6c). We next tested elevated VEGFA and SOX2 expression, and then evaluated tumors with high levels of VEGFA, SOX2 and SNAI2, and low GABRE (a surrogate for miR-452) expression. Tumors in the top quartile of both VEGFA and SOX2 expression showed significantly worse DMFS (HR for recurrence [95% CI]=1.9 [1.36–2.65]; P=0.00014; Figure 6c), while those with high expression of VEGFA, SOX2 and SNAI2 and the lowest quartile GABRE expression showed an even greater risk of relapse (HR [95% CI]=2.03 [1.46–2.84]; P=2.0e−05; Figure 6c).
A similar analysis showed VEGFA expression was of greater prognostic importance in breast cancers defined as ER-negative by clinical ER protein immunohistochemistry (n=170 with both VEGFA and ER data). Although the top quartile VEGFA expression associated with shorter DMFS (HR [95% CI]=2.87 [1.69–4.86]; P= 4.2e−05; Supplementary Figure S7), median cutoffs were used to classify ‘high’ or ‘low’ expressers due to the reduced sample size. Using median cutoffs, high VEGFA associated with shorter DMFS (HR [95% CI]=1.71 [1.01–2.87]; P=0.042; Figure 6d). ER-negative cancers with high VEGFA and SOX2 expression showed significantly shorter DMFS (HR=2.72 [95% CI, 1.11–6.7]; P=0.023; Figure 6d). Remarkably, among ER-negative breast cancers, those with high VEGFA, SOX2 and SNAI2 together with low GABRE expression showed a 4.54-fold higher risk of metastasis, with DMFS (HR [95% CI]=4.54 [1.67–12.35]; P=0.0011, Figure 6d). These KM Plotter data identify a very aggressive population within all cancers and in ER-negative breast cancers, in which the mechanistic pathway identified herein appears to be activated.
To validate these findings in an independent patient group, a similar analysis was carried out for disease-specific survival (DSS) in the METABRIC breast cancer data set (n=1286). Analysis of this second independent patient cohort confirmed the prognostic significance of VEGFA and showed that elevation of both VEGFA and SOX2 expression associated with a worse survival than did VEGFA elevation alone. High VEGFA alone conferred a 1.69-fold higher risk of death (DSS HR (95% CI) =1.69 (1.25–2.3), P=0.000595: Figure 6e, left), and METABRIC cancers in the top quartile of both VEGFA and SOX2 had even worse outcome (DSS HR (95% CI) =1.76 (1.3–2.38), P=0.000199; Figure 6e, right).
Discussion
VEGFA is best known as an angiogenic agent,73 but it also promotes cancer invasion and metastasis through mechanisms that are not fully understood. VEGFA not only creates a vascular niche for expanding stem cells,21 it was recently shown to increase the stem-like cell population in certain human malignancies, including breast cancer.18, 19, 20, 22 Hypoxia, caused by angiogenesis inhibitors, stimulates VEGFA gene expression, and would thus contribute to CSC expansion74 and disease recurrence and progression.16, 17
CSC have been implicated as drivers of tumor metastasis, however, the molecular pathways linking stemness and induction of metastasis are not fully elucidated. Populations bearing surface CSC markers75, 76, 77 or that are enriched for ALDH1 activity78 have been shown to have greater motility, invasiveness and metastatic potential than the bulk of the cancer population. Recent work in a pancreatic model showed EMT and dissemination may precede overt tumor invasion.79 Circulating tumor cells could be detected during in situ tumor growth before overt invasion. Circulating tumor cells bearing the CSC marker, CD44+, showed much more aggressive self-renewal and tumor-generating potential than CD44+-positive cells from the primary tumor site, indicating that escape of stem-like cells from the primary tumor environment is linked to increased self-renewal potential.79
Sox2 is an important mediator of self-renewal in embryonic stem cells and is an oncogenic driver of CSC in several cancer models, including breast cancer.19, 25, 61, 62 Our prior work showed that VEGFA mediates CSC expansion via STAT3-driven SOX2 induction in breast and lung cancer models.19 Proinflammatory cytokines that are upregulated on breast cancer cell invasion into fat also induce SOX2 to drive CSC self-renewal.25 SOX2 knockdown can decrease both CSC and experimental lung metastasis62 and SOX2 expression is associated with colon cancer metastasis.80 Present work reveals Sox2 is necessary for VEGFA-driven SNAI2 induction, EMT and invasion of breast cancer cells and provides a mechanistic link between VEGFA-stimulated CSC expansion via SOX2 induction,19 and the upregulation of metastatic potential.
EMT arising from overexpression of various EMT-TFs has been shown to increase tumor-initiating cell abundance;41, 42 moreover, stem-like cells exhibit EMT properties such as increased mesenchymal markers and EMT transcription factor expression81 suggesting an intimate relationship between CSCs and EMT. Mammary cell lines overexpressing various EMT-TFs showed PLCγ-mediated PKC activation leading to a c-Jun/Fra1-induced CSC transcriptional program.82 TGF-β and TNFα pathways interact to drive both EMT and upregulate breast CSC properties.83 Elegant in vitro and in vivo studies in a Trp53-null mouse breast cancer model showed cross-talk between transformed mesenchymal cells and tumor-initiating subpopulations. The mesenchymal cells produced stimulatory ligands driving CSC surface receptors to increase tumorigenicity and metastasis via both Wnt/Fzd7 and CXCL12/CXCR4 pathways,84 suggesting that heterogeneous cell populations with differing stem cell self-renewal may interact with each other to drive pathways governing both self-renewal and metastasis.
miRNAs regulate many processes central to oncogenesis.47 Several miRNAs oppose EMT by targeting EMT transcription factors.85 miR-200 targets Zeb1 and Zeb254,57–59 as does miR-138,86 and Snail is targeted by miR-30a56 and miR-34.60 A number of miRNAs not only regulate EMT but also serve as key CSC regulators. For example, miR-200 not only inhibits EMT by suppressing Zeb1/2, but also downregulates stem-like cells by targeting Bmi187, 88 and the Notch pathway.89 In addition to its action on Snail, miR-34a also decreases CSC by targeting Myc90 and downregulates CD44 expression to decrease prostate CSC.91 miR-128-2 targets both the EMT mediator Snail, and CSC drivers Nanog, KLF492 and Bmi1.93
Here we identify miR-452 at the interface between VEGFA-activated CSC self-renewal and EMT, providing a novel connection between VEGFA, induction of the embryonic stem cell transcription factor, Sox2 and EMT. Sox2 not only governs CSC expansion, but also mediates acquisition of EMT and metastatic potential. VEGFA-induced SOX2 expression is required not only for CSC expansion,19 but also for VEGFA-mediated SNAI2 induction. VEGFA increased SOX2 expression within an hour, but EMT-TF levels, and in particular that of Slug, rose over several days, suggesting that Sox2 affects Slug indirectly, via an intermediary mechanism. Our miRNA screen identified miR-452 as a putative metastasis suppressor, significantly downregulated by VEGFA in MDA-MB-231 cells. miR-452 is downregulated in breast cancers compared with normal breast tissue.94 We show miR-452 targets SNAI2 directly. MiR-452 overexpression abrogated the VEGFA-mediated upregulation of SNAI2 expression and cell invasion, and miR-452 antagomir was sufficient to upregulate SNAI2 expression and cell invasion. Sox2 upregulation by VEGFA mediates the loss of miR-452, and miR-452 loss is required for Slug upregulation and for VEGFA-driven cell motility, invasion and metastasis in breast cancer models. Notably, miR-452 expression correlates inversely with glioblastoma survival and inhibits glioma stem cells and tumorigenesis by targeting CSC mediators, Bmi1, LEF1 and TCF4.95 Thus, miR-452 may not only regulate Sox2-driven EMT via Slug, but may also serve a dual role at the interface between EMT and CSC regulation.
The pathway linking VEGFA to Sox2 upregulation, miR-452 loss and SNAI2 induction is supported by our analysis of two major data sets including over 2500 primary human breast cancers. Although high intratumor VEGFA levels detected by immunohistochemistry have been linked to poor breast cancer outcome, most studies have been small and results controversial.96, 97 Our analysis showed breast cancers in the highest quartile of VEGFA expression fare worse than all others, and the prognostic value of high VEGFA levels is increased by sequential addition of high SOX2, SNAI2 and decreased GABRE (a surrogate for miR-452) expression. This finding is less important for its prognostic significance than it is as a confirmation of the molecular pathway identified herein. Among aggressive breast cancers expressing high VEGFA, those with SOX2 overexpression define an even more aggressive subgroup in the two independent data sets evaluated.
VEGFA is a critical mediator of tumor progression. It acts to generate a vascular niche for CSC through autocrine and paracrine action on both tumor and microenvironmental components, and links CSC self-renewal to the acquisition of metastatic potential. To date, targeting VEGFA has had limited success in cancer, and this may be due to anti-angiogenics causing tumor hypoxia, leading to upregulation of both VEGFA and CSC. Since treatment of metastasis is the final therapeutic frontier, it is hoped that mechanistic insights linking VEGFA to tumor initiation and the acquisition of metastatic potential will ultimately generate new strategies for VEGF pathway-targeted intervention.
Materials and methods
Cell lines and reagents
Luciferase-tagged MDA-MB-231 (from J Massague, MSKCC, New York, NY, USA), Lenti-X 293 T cells (from Clontech, Mountain View, CA, USA) and MCF12A line were cultured as described.19 Both lines were verified by STR profiling. SUM149PT cells were provided by S Ethier (Medical University of South Carolina, Charleston, SC, USA) and cultured in Ham's F12 medium with 5% FBS, 5 μg/ml insulin, 1 μg/ml hydrocortisone and 1 mm HEPES.98 All lines were mycoplasma free.
Scratch assay
MDA-MB-231 cells were seeded into six-well plates, grown to confluence and wound-healing scratch assays were performed as in Larrea et al.,99 and the cells were photomicrographed after 12 h using an Olympus CKX41 microscope.
Transwell migration and invasion assays
Real-time cell analysis from Xcelligence (ACEA Biosciences, San Diego, CA, USA) was used for automated transwell migration and invasion from serum-free toward serum-containing medium was as described100 and plotted as mean cell index±s.e.m. for at least three independent wells per group.
Quantitative real-time PCR and miRNA RT–PCR
QPCR was performed at least thrice and mean Ct values normalized to GAPDH or 18S values. mRNA isolation used miRNeasy mini kit (Qiagen, Hilden, Germany) and cDNA synthesys used NcodemiR First-Strand cDNA synthesis kit (Invitrogen, Carlsbad, CA, USA). miR-452 levels were assayed by QPCR. PCR primers for EMT markers and transcription factors assayed, and for miR-452 are in Supplementary Figure S8.
miRNA screen and antagomiR
MDA-MB-231 cells were treated ±10 ng/ml VEGFA for 7 days followed by Ready-to-Use PCR microRNA array, Human panel I+II in 384-well plates from Exiqon (Woburn, MA, USA). miRCURY LNA miR-452 antagomir and miRCURY LNA miRNA antagomir control were purchased from Exiqon (Woburn, MA, USA) and transduced per manufacturer.
siRNA analysis and western blots
SiRNA pools of three to five target-specific 19–25 nucleotide siRNAs designed to knockdown Slug/Sox2 and control siRNAs were purchased from Santa Cruz Biotechnology (Dallas, TX, USA) and used per manufacturer. Western blots were as described19 using antibodies: anti-Slug (#9585) and Sox2 (#3579) from Cell Signaling (Denver, CO, USA); β-actin (#A1978) from Sigma-Aldrich (St Louis, MO, USA).
Lentivirus production and transduction
Human SNAI2 (EX-T1290-Lv155), control vector (EX-NEG-Lv155), hsa-mir-452 (HmiR0407-MR03) and miRNA scrambled control clone (CmiR0001-MR03) lentivirus vectors were purchased from GeneCopoeia (Rockville, MD, USA). Lentivirus vectors were co-transfected with Delta VPR and CMV VSVG plasmids (Addgene, Cambridge, MA, USA) into Lenti-X 293T cells with Lipofectamine Plus. Viral supernatants at 48 h were concentrated by ultracentrifugation for 2 h at 22 000 r.p.m. at 4 °C. MDA-MB-231 was infected with virus in polybrene (10 μg/ml) as described.19 Stable expression was confirmed by GFP fluorescence visualization and western.
Luciferase assays
293 T and MDA-MB-231 were transfected with human SNAI2 3′-UTR luciferase reporter (GeneCopoeia, Rockville MD, USA) plasmid together with miR-452 or control miRNA plasmid. After 48 h, Firefly and Renilla luciferase reporter activity luciferase activity was measured using Luc-Pair Duo-Luciferase Assay Kit 2.0 (GeneCopoeia, Rockville MD, USA) per manufacturer's instructions.
Experimental lung metastasis assay
MDA-MB-231-luc and controls transduced with miR-452 or miR-452+SNAI2 were pre-treated ±VEGFA for 7 days, before injection of 5 × 105cells via tail vein into 4–6-week female Balb/C nude mice as described.19 Each experimental group contained 10 animals. The mice were imaged by in vivo imaging system (Xenogen, Caliper, Hopkinton, MA, USA) and bioluminescence (photon flux) was quantified with time as described.100 All animal work was carried out in compliance with the Institutional Animal Care and Use Committee in the University of Miami.
Statistical analysis and expression analysis of VEGFA, SOX2, SNAI2, GABRE genes and miR-452
The METABRIC data set contains gene expression data for 2136 and microRNA expression for 1448 primary breast cancer samples, with both available in 1302 samples, together with clinical information and DSS outcome data.71, 101 METABRIC and the independent Enerly primary breast cancer data set, containing 101 cases,72 were used to identify a correlation between miR-452 and GABRE expression by Pearson’s correlation. The KM plotter data set contains gene expression from primary human breast cancers (n=2553) and was used for the analysis of distant metastasis-free survival (DMFS).
For clinical outcome analysis, expression quartiles were used to test whether VEGFA expression alone, or with the other genes, associated with poor DSS (METABRIC) or with poor DMFS (KM plotter) using Kaplan–Meier analysis and Univariate Cox proportional hazards analysis identified hazard ratios with 95% CI. DSS or DMFS curves were also compared using the log-rank test and the P-value from this analysis was displayed in each graph. Data analysis was performed using R statistical software or by using the KM plotter web tool, as in Gyorffy et al.102 Mihaly et al.103
For in vitro work, data are graphed from ⩾3 biologic experiments as means±s.e.m. Means were compared with two-tailed Student’s t-tests. P-values <0.05 were considered statistically significant. Statistical differences of a real-time cell analysis data between invasion rates used the ‘Compare Growth Curves’ function (http://bioinf.wehi.edu.au/software/compareCurves/).
References
Olsson AK, Dimberg A, Kreuger J, Claesson-Welsh L . VEGF receptor signalling—in control of vascular function. Nat Rev Mol Cell Biol 2006; 7: 359–371.
Leung DW, Cachianes G, Kuang WJ, Goeddel DV, Ferrara N . Vascular endothelial growth factor is a secreted angiogenic mitogen. Science 1989; 246: 1306–1309.
Tischer E, Gospodarowicz D, Mitchell R, Silva M, Schilling J, Lau K et al. Vascular endothelial growth factor: a new member of the platelet-derived growth factor gene family. Biochem Biophys Res Commun 1989; 165: 1198–1206.
Hicklin DJ, Ellis LM . Role of the vascular endothelial growth factor pathway in tumor growth and angiogenesis. J Clin Oncol 2005; 23: 1011–1027.
Goel HL, Mercurio AM . VEGF targets the tumour cell. Nat Rev Cancer 2013; 13: 871–882.
Berns EM, Klijn JG, Look MP, Grebenchtchikov N, Vossen R, Peters H et al. Combined vascular endothelial growth factor and TP53 status predicts poor response to tamoxifen therapy in estrogen receptor-positive advanced breast cancer. Clin Cancer Res 2003; 9: 1253–1258.
Manders P, Beex LV, Tjan-Heijnen VC, Geurts-Moespot J, Van Tienoven TH, Foekens JA et al. The prognostic value of vascular endothelial growth factor in 574 node-negative breast cancer patients who did not receive adjuvant systemic therapy. Br J Cancer 2002; 87: 772–778.
Senger DR . Vascular endothelial growth factor: much more than an angiogenesis factor. Mol Biol Cell 2010; 21: 377–379.
Lichtenberger BM, Tan PK, Niederleithner H, Ferrara N, Petzelbauer P, Sibilia M . Autocrine VEGF signaling synergizes with EGFR in tumor cells to promote epithelial cancer development. Cell 2010; 140: 268–279.
Hanahan D, Folkman J . Patterns and emerging mechanisms of the angiogenic switch during tumorigenesis. Cell 1996; 86: 353–364.
Mercurio AM, Lipscomb EA, Bachelder RE . Non-angiogenic functions of VEGF in breast cancer. J Mammary Gland Biol Neoplasia 2005; 10: 283–290.
Adams J, Carder PJ, Downey S, Forbes MA, MacLennan K, Allgar V et al. Vascular endothelial growth factor (VEGF) in breast cancer: comparison of plasma, serum, and tissue VEGF and microvessel density and effects of tamoxifen. Cancer Res 2000; 60: 2898–2905.
Sullivan LA, Brekken RA . The VEGF family in cancer and antibody-based strategies for their inhibition. MAbs 2010; 2: 165–175.
Shih T, Lindley C . Bevacizumab: an angiogenesis inhibitor for the treatment of solid malignancies. Clin Ther 2006; 28: 1779–1802.
Montero AJ, Escobar M, Lopes G, Gluck S, Vogel C . Bevacizumab in the treatment of metastatic breast cancer: friend or foe? Curr Oncol Rep 2012; 14: 1–11.
Paez-Ribes M, Allen E, Hudock J, Takeda T, Okuyama H, Vinals F et al. Antiangiogenic therapy elicits malignant progression of tumors to increased local invasion and distant metastasis. Cancer Cell 2009; 15: 220–231.
Ebos JM, Lee CR, Cruz-Munoz W, Bjarnason GA, Christensen JG, Kerbel RS . Accelerated metastasis after short-term treatment with a potent inhibitor of tumor angiogenesis. Cancer Cell 2009; 15: 232–239.
Beck B, Driessens G, Goossens S, Youssef KK, Kuchnio A, Caauwe A et al. A vascular niche and a VEGF-Nrp1 loop regulate the initiation and stemness of skin tumours. Nature 2011; 478: 399–403.
Zhao D, Pan C, Sun J, Gilbert C, Drews-Elger K, Azzam DJ et al. VEGF drives cancer-initiating stem cells through VEGFR-2/Stat3 signaling to upregulate Myc and Sox2. Oncogene 2014; 34: 3107–3119.
Goel HL, Pursell B, Chang C, Shaw LM, Mao J, Simin K et al. GLI1 regulates a novel neuropilin-2/alpha6beta1 integrin based autocrine pathway that contributes to breast cancer initiation. EMBO Mol Med 2013; 5: 488–508.
Bao S, Wu Q, Sathornsumetee S, Hao Y, Li Z, Hjelmeland AB et al. Stem cell-like glioma cells promote tumor angiogenesis through vascular endothelial growth factor. Cancer Res 2006; 66: 7843–7848.
Hamerlik P, Lathia JD, Rasmussen R, Wu Q, Bartkova J, Lee M et al. Autocrine VEGF-VEGFR2-Neuropilin-1 signaling promotes glioma stem-like cell viability and tumor growth. J Exp Med 2012; 209: 507–520.
Malanchi I, Santamaria-Martinez A, Susanto E, Peng H, Lehr HA, Delaloye JF et al. Interactions between cancer stem cells and their niche govern metastatic colonization. Nature 2012; 481: 85–89.
Takebe N, Harris PJ, Warren RQ, Ivy SP . Targeting cancer stem cells by inhibiting Wnt, Notch, and Hedgehog pathways. Nat Rev Clin Oncol 2011; 8: 97–106.
Picon-Ruiz M, Pan C, Drews-Elger K, Jang K, Besser A, Zhao D et al. Interactions between adipocytes and breast cancer cells stimulate cytokine production and drive Src/Sox2/miR-302b mediated malignant progression. Cancer Res 2016; 76: 491–504.
Thiery JP . Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer 2002; 2: 442–454.
Kalluri R, Weinberg RA . The basics of epithelial-mesenchymal transition. J Clin Invest 2009; 119: 1420–1428.
Micalizzi DS, Farabaugh SM, Ford HL . Epithelial-mesenchymal transition in cancer: parallels between normal development and tumor progression. J Mammary Gland Biol Neoplasia 2010; 15: 117–134.
Thiery JP, Acloque H, Huang RY, Nieto MA . Epithelial-mesenchymal transitions in development and disease. Cell 2009; 139: 871–890.
Huber MA, Kraut N, Beug H . Molecular requirements for epithelial-mesenchymal transition during tumor progression. Curr Opin Cell Biol 2005; 17: 548–558.
Onder TT, Gupta PB, Mani SA, Yang J, Lander ES, Weinberg RA . Loss of E-cadherin promotes metastasis via multiple downstream transcriptional pathways. Cancer Res 2008; 68: 3645–3654.
Batlle E, Sancho E, Franci C, Dominguez D, Monfar M, Baulida J et al. The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat Cell Biol 2000; 2: 84–89.
Cano A, Perez-Moreno MA, Rodrigo I, Locascio A, Blanco MJ, del Barrio MG et al. The transcription factor snail controls epithelial-mesenchymal transitions by repressing E-cadherin expression. Nat Cell Biol 2000; 2: 76–83.
Bolos V, Peinado H, Perez-Moreno MA, Fraga MF, Esteller M, Cano A . The transcription factor Slug represses E-cadherin expression and induces epithelial to mesenchymal transitions: a comparison with Snail and E47 repressors. J Cell Sci 2003; 116: 499–511.
Eger A, Aigner K, Sonderegger S, Dampier B, Oehler S, Schreiber M et al. DeltaEF1 is a transcriptional repressor of E-cadherin and regulates epithelial plasticity in breast cancer cells. Oncogene 2005; 24: 2375–2385.
Comijn J, Berx G, Vermassen P, Verschueren K, van GL, Bruyneel E et al. The two-handed E box binding zinc finger protein SIP1 downregulates E-cadherin and induces invasion. Mol Cell 2001; 7: 1267–1278.
Yang J, Mani SA, Donaher JL, Ramaswamy S, Itzykson RA, Come C et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 2004; 117: 927–939.
Hartwell KA, Muir B, Reinhardt F, Carpenter AE, Sgroi DC, Weinberg RA . The Spemann organizer gene, Goosecoid, promotes tumor metastasis. Proc Natl Acad Sci USA 2006; 103: 18969–18974.
Mani SA, Yang J, Brooks M, Schwaninger G, Zhou A, Miura N et al. Mesenchyme Forkhead 1 (FOXC2) plays a key role in metastasis and is associated with aggressive basal-like breast cancers. Proc Natl Acad Sci USA 2007; 104: 10069–10074.
Hajra KM, Chen DY, Fearon ER . The SLUG zinc-finger protein represses E-cadherin in breast cancer. Cancer Res 2002; 62: 1613–1618.
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan A, Zhou AY et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008; 133: 704–715.
Morel AP, Lievre M, Thomas C, Hinkal G, Ansieau S, Puisieux A . Generation of breast cancer stem cells through epithelial-mesenchymal transition. PLoS ONE 2008; 3: e2888.
Bachelder RE, Wendt MA, Mercurio AM . Vascular endothelial growth factor promotes breast carcinoma invasion in an autocrine manner by regulating the chemokine receptor CXCR4. Cancer Res 2002; 62: 7203–7206.
Gonzalez-Moreno O, Lecanda J, Green JE, Segura V, Catena R, Serrano D et al. VEGF elicits epithelial-mesenchymal transition (EMT) in prostate intraepithelial neoplasia (PIN)-like cells via an autocrine loop. Exp Cell Res 2010; 316: 554–567.
Kim LS, Huang S, Lu W, Lev DC, Price JE . Vascular endothelial growth factor expression promotes the growth of breast cancer brain metastases in nude mice. Clin Exp Metastasis 2004; 21: 107–118.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Ceppi P, Peter ME . MicroRNAs regulate both epithelial-to-mesenchymal transition and cancer stem cells. Oncogene 2014; 33: 269–278.
Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N, Nishikura K et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 2005; 436: 740–744.
Diederichs S, Haber DA . Dual role for argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 2007; 131: 1097–1108.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435: 834–838.
Garzon R, Calin GA, Croce CM . MicroRNAs in Cancer. Annu Rev Med 2009; 60: 167–179.
Iorio MV, Croce CM . MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol Med 2012; 4: 143–159.
Pencheva N, Tavazoie SF . Control of metastatic progression by microRNA regulatory networks. Nat Cell Biol 2013; 15: 546–554.
Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep 2008; 9: 582–589.
Gregory PA, Bracken CP, Smith E, Bert AG, Wright JA, Roslan S et al. An autocrine TGF-beta/ZEB/miR-200 signaling network regulates establishment and maintenance of epithelial-mesenchymal transition. Mol Biol Cell 2011; 22: 1686–1698.
Kumarswamy R, Mudduluru G, Ceppi P, Muppala S, Kozlowski M, Niklinski J et al. MicroRNA-30a inhibits epithelial-to-mesenchymal transition by targeting Snai1 and is downregulated in non-small cell lung cancer. Int J Cancer 2012; 130: 2044–2053.
Park SM, Gaur AB, Lengyel E, Peter ME . The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 2008; 22: 894–907.
Korpal M, Lee ES, Hu G, Kang Y . The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 2008; 283: 14910–14914.
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 2008; 10: 593–601.
Kim NH, Kim HS, Li XY, Lee I, Choi HS, Kang SE et al. A p53/miRNA-34 axis regulates Snail1-dependent cancer cell epithelial-mesenchymal transition. J Cell Biol 2011; 195: 417–433.
Leis O, Eguiara A, Lopez-Arribillaga E, Alberdi MJ, Hernandez-Garcia S, Elorriaga K et al. Sox2 expression in breast tumours and activation in breast cancer stem cells. Oncogene 2011; 31: 1354–1365.
Xiang R, Liao D, Cheng T, Zhou H, Shi Q, Chuang TS et al. Downregulation of transcription factor SOX2 in cancer stem cells suppresses growth and metastasis of lung cancer. Br J Cancer 2011; 104: 1410–1417.
Azzam DJ, Zhao D, Sun J, Minn AJ, Ranganathan P, Drews-Elger K et al. Triple negative breast cancer initiating cell subsets differ in functional and molecular characteristics and in γ-secretase inhibitor drug responses. EMBO Mol Med 2013; 5: 1502–1522.
Fang X, Yu W, Li L, Shao J, Zhao N, Chen Q et al. ChIP-seq and functional analysis of the SOX2 gene in colorectal cancers. OMICS 2010; 14: 369–384.
Boyer LA, Lee TI, Cole MF, Johnstone SE, Levine SS, Zucker JP et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005; 122: 947–956.
Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet 2008; 40: 499–507.
Engelen E, Akinci U, Bryne JC, Hou J, Gontan C, Moen M et al. Sox2 cooperates with Chd7 to regulate genes that are mutated in human syndromes. Nat Genet 2011; 43: 607–611.
Drews-Elger K, Brinkman JA, Miller P, Shah SH, Harrell JC, da Silva TG et al. Primary breast tumor-derived cellular models: characterization of tumorigenic, metastatic, and cancer-associated fibroblasts in dissociated tumor (DT) cultures. Breast Cancer Res Treat 2014; 144: 503–517.
Hirakawa S, Kodama S, Kunstfeld R, Kajiya K, Brown LF, Detmar M . VEGF-A induces tumor and sentinel lymph node lymphangiogenesis and promotes lymphatic metastasis. J Exp Med 2005; 201: 1089–1099.
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM et al. The human genome browser at UCSC. Genome Res 2002; 12: 996–1006.
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 2012; 486: 346–352.
Enerly E, Steinfeld I, Kleivi K, Leivonen SK, Aure MR, Russnes HG et al. miRNA-mRNA integrated analysis reveals roles for miRNAs in primary breast tumors. PLoS ONE 2011; 6: e16915.
Ferrara N, Davis-Smyth T . The biology of vascular endothelial growth factor. Endocr Rev 1997; 18: 4–25.
Conley SJ, Gheordunescu E, Kakarala P, Newman B, Korkaya H, Heath AN et al. Antiangiogenic agents increase breast cancer stem cells via the generation of tumor hypoxia. Proc Natl Acad Sci USA 2012; 109: 2784–2789.
Todaro M, Gaggianesi M, Catalano V, Benfante A, Iovino F, Biffoni M et al. CD44v6 is a marker of constitutive and reprogrammed cancer stem cells driving colon cancer metastasis. Cell Stem Cell 2014; 14: 342–356.
Sheridan C, Kishimoto H, Fuchs RK, Mehrotra S, Bhat-Nakshatri P, Turner CH et al. CD44+/CD24- breast cancer cells exhibit enhanced invasive properties: an early step necessary for metastasis. Breast Cancer Res 2006; 8: R59.
Abraham BK, Fritz P, McClellan M, Hauptvogel P, Athelogou M, Brauch H . Prevalence of CD44+/CD24-/low cells in breast cancer may not be associated with clinical outcome but may favor distant metastasis. Clin Cancer Res 2005; 11: 1154–1159.
Charafe-Jauffret E, Ginestier C, Iovino F, Tarpin C, Diebel M, Esterni B et al. Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin Cancer Res 2010; 16: 45–55.
Rhim AD, Mirek ET, Aiello NM, Maitra A, Bailey JM, McAllister F et al. EMT and dissemination precede pancreatic tumor formation. Cell 2012; 148: 349–361.
Han X, Fang X, Lou X, Hua D, Ding W, Foltz G et al. Silencing SOX2 induced mesenchymal-epithelial transition and its expression predicts liver and lymph node metastasis of CRC patients. PLoS One 2012; 7: e41335.
Gupta PB, Chaffer CL, Weinberg RA . Cancer stem cells: mirage or reality? Nat Med 2009; 15: 1010–1012.
Tam WL, Lu H, Buikhuisen J, Soh BS, Lim E, Reinhardt F et al. Protein kinase C alpha is a central signaling node and therapeutic target for breast cancer stem cells. Cancer Cell 2013; 24: 347–364.
Polyak K, Weinberg RA . Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat Rev Cancer 2009; 9: 265–273.
Zhang M, Tsimelzon A, Chang CH, Fan C, Wolff A, Perou CM et al. Intratumoral heterogeneity in a Trp53-null mouse model of human breast cancer. Cancer Discov 2015; 5: 520–533.
Tavazoie SF, Alarcon C, Oskarsson T, Padua D, Wang Q, Bos PD et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 2008; 451: 147–152.
Liu X, Wang C, Chen Z, Jin Y, Wang Y, Kolokythas A et al. MicroRNA-138 suppresses epithelial-mesenchymal transition in squamous cell carcinoma cell lines. Biochem J 2011; 440: 23–31.
Shimono Y, Zabala M, Cho RW, Lobo N, Dalerba P, Qian D et al. Downregulation of miRNA-200c links breast cancer stem cells with normal stem cells. Cell 2009; 138: 592–603.
Wellner U, Schubert J, Burk UC, Schmalhofer O, Zhu F, Sonntag A et al. The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat Cell Biol 2009; 11: 1487–1495.
Brabletz S, Bajdak K, Meidhof S, Burk U, Niedermann G, Firat E et al. The ZEB1/miR-200 feedback loop controls Notch signalling in cancer cells. EMBO J 2011; 30: 770–782.
Yamamura S, Saini S, Majid S, Hirata H, Ueno K, Deng G et al. MicroRNA-34a modulates c-Myc transcriptional complexes to suppress malignancy in human prostate cancer cells. PLoS ONE 2012; 7: e29722.
Liu C, Kelnar K, Liu B, Chen X, Calhoun-Davis T, Li H et al. The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat Med 2011; 17: 211–215.
Qian P, Banerjee A, Wu ZS, Zhang X, Wang H, Pandey V et al. Loss of SNAIL regulated miR-128-2 on chromosome 3p22.3 targets multiple stem cell factors to promote transformation of mammary epithelial cells. Cancer Res 2012; 72: 6036–6050.
Godlewski J, Nowicki MO, Bronisz A, Williams S, Otsuki A, Nuovo G et al. Targeting of the Bmi-1 oncogene/stem cell renewal factor by microRNA-128 inhibits glioma proliferation and self-renewal. Cancer Res 2008; 68: 9125–9130.
van Schooneveld E, Wouters MC, Van dA I, Peeters DJ, Wildiers H, van Dam PA et al. Expression profiling of cancerous and normal breast tissues identifies microRNAs that are differentially expressed in serum from patients with (metastatic) breast cancer and healthy volunteers. Breast Cancer Res 2012; 14: R34.
Liu L, Chen K, Wu J, Shi L, Hu B, Cheng S et al. Downregulation of miR-452 promotes stem-like traits and tumorigenicity of gliomas. Clin Cancer Res 2013; 19: 3429–3438.
Arias-Pulido H, Chaher N, Gong Y, Qualls C, Vargas J, Royce M . Tumor stromal vascular endothelial growth factor A is predictive of poor outcome in inflammatory breast cancer. BMC Cancer 2012; 12: 298.
Sa-Nguanraksa D, Chuangsuwanich T, Pongpruttipan T, Charoenrat P . High vascular endothelial growth factor gene expression predicts poor outcome in patients with non-luminal A breast cancer. Mol Clin Oncol 2015; 3: 1103–1108.
Ethier S, Mahacek ML, Gullick WJ, Frank TS, Weber BL . Differential isolation of normal luminal mammary epithelial cells and breast cancer cells from primary and metastatic sites using selective media. Cancer Res 1993; 53: 627–635.
Larrea MD, Hong F, Wander SA, da Silva TG, Helfman D, Lannigan D et al. RSK1 drives p27Kip1 phosphorylation at T198 to promote RhoA inhibition and increase cell motility. Proc Natl Acad Sci USA 2009; 106: 9268–9273.
Wander S, Zhao D, Besser A, Hong F, Wei J, Ince T et al. PI3K/mTOR inhibition can impair tumor invasion and metastasis in vivo despite a lack of antiproliferative action in vitro: implications for targeted therapy. Breast Cancer Res Treat 2013; 138: 369–381.
Dvinge H, Git A, Graf S, Salmon-Divon M, Curtis C, Sottoriva A et al. The shaping and functional consequences of the microRNA landscape in breast cancer. Nature 2013; 497: 378–382.
Gyorffy B, Benke Z, Lanczky A, Balazs B, Szallasi Z, Timar J et al. RecurrenceOnline: an online analysis tool to determine breast cancer recurrence and hormone receptor status using microarray data. Breast Cancer Res Treat 2012; 132: 1025–1034.
Mihaly Z, Kormos M, Lanczky A, Dank M, Budczies J, Szasz MA et al. A meta-analysis of gene expression-based biomarkers predicting outcome after tamoxifen treatment in breast cancer. Breast Cancer Res Treat 2013; 140: 219–232.
Acknowledgements
This work was supported by funding from the Breast Cancer Research Foundation to JMS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kim, M., Jang, K., Miller, P. et al. VEGFA links self-renewal and metastasis by inducing Sox2 to repress miR-452, driving Slug. Oncogene 36, 5199–5211 (2017). https://doi.org/10.1038/onc.2017.4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.4
This article is cited by
-
NRP1 interacts with endoglin and VEGFR2 to modulate VEGF signaling and endothelial cell sprouting
Communications Biology (2024)
-
ID4-dependent secretion of VEGFA enhances the invasion capability of breast cancer cells and activates YAP/TAZ via integrin β3-VEGFR2 interaction
Cell Death & Disease (2024)
-
Establishing and Validating an Innovative Focal Adhesion-Linked Gene Signature for Enhanced Prognostic Assessment in Endometrial Cancer
Reproductive Sciences (2024)
-
MicroRNA-452: a double-edged sword in multiple human cancers
Clinical and Translational Oncology (2023)
-
A reliable mouse model of liver and lung metastasis by injecting esophageal cancer stem cells (CSCs) through tail-vein injection
Molecular Biology Reports (2023)