Abstract
In Alzheimer’s disease (AD), a single-nucleotide polymorphism in the gene encoding brain-derived neurotrophic factor (BDNFVal66Met) is associated with worse impact of primary AD pathology (beta-amyloid, Aβ) on neurodegeneration and cognitive decline, rendering BDNFVal66Met an important modulating factor of cognitive impairment in AD. However, the effect of BDNFVal66Met on functional networks that may underlie cognitive impairment in AD is poorly understood. Using a cross-validation approach, we first explored in subjects with autosomal dominant AD (ADAD) from the Dominantly Inherited Alzheimer Network (DIAN) the effect of BDNFVal66Met on resting-state fMRI assessed functional networks. In seed-based connectivity analysis of six major large-scale networks, we found a stronger decrease of hippocampus (seed) to medial-frontal connectivity in the BDNFVal66Met carriers compared to BDNFVal homozogytes. BDNFVal66Met was not associated with connectivity in any other networks. Next, we tested whether the finding of more pronounced decrease in hippocampal-medial-frontal connectivity in BDNFVal66Met could be also found in elderly subjects with sporadically occurring Aβ, including a group with subjective cognitive decline (N = 149, FACEHBI study) and a group ranging from preclinical to AD dementia (N = 114, DELCODE study). In both of these independently recruited groups, BDNFVal66Met was associated with a stronger effect of more abnormal Aβ-levels (assessed by biofluid-assay or amyloid-PET) on hippocampal-medial-frontal connectivity decreases, controlled for hippocampus volume and other confounds. Lower hippocampal-medial-frontal connectivity was associated with lower global cognitive performance in the DIAN and DELCODE studies. Together these results suggest that BDNFVal66Met is selectively associated with a higher vulnerability of hippocampus-frontal connectivity to primary AD pathology, resulting in greater AD-related cognitive impairment.
Similar content being viewed by others
Introduction
Brain-derived neurotrophic factor (BDNF) is part of the neurotrophin family, playing a critical role in synaptic plasticity and repair throughout the lifespan [1,2,3]. BDNF is expressed beyond the neocortex particularly in the hippocampus [4], where BDNF is essential for long-term potentiation (LTP) underlying hippocampus-related memory [1, 3, 5]. Hippocampus-dependent memory impairment is a key feature of Alzheimer’s disease (AD), where beta-amyloid (Aβ) and tau are hallmark pathologies affecting the hippocampus early in AD [6,7,8]. Post-mortem studies showed decreased hippocampal BDNF levels among other brain regions in patients with AD dementia or mild cognitive impairment (MCI) [9, 10]. These observations raise the important question whether altered BDNF levels modulate the impact of AD pathology on brain function. Preclinical research in transgenic AD mouse models showed that genetic delivery or overexpression of BDNF improved hippocampus LTP [11, 12] and alleviated the impact of amyloid or tau pathology on cell loss and memory [12, 13], suggesting protective BDNF effects in AD. However, in humans the role of BDNF in modulating the impact of AD pathology is not well understood.
A single-nucleotide polymorphism (SNP) (rs6265) resulting in the substitution of Valine by Methionine in the BDNF pro-domain (i.e. BDNFVal66Met) leads to reduced synaptic BDNF release [14, 15]. Because BDNFVal66Met shows a relatively high prevalence ranging from 29% in European to 72% in Asian countries [16], BDNFVal66Met may constitute a genetic factor that modifies functional brain alterations in a substantial portion of AD subjects [17,18,19,20,21]. Although neuroimaging studies in AD showed only small BDNFVal66Met effects on hippocampal volume [22, 23], several studies have found BDNFVal66Met-related reductions in hippocampal FDG-PET metabolism and stronger memory impairment in both autosomal dominant and sporadic AD [17, 19,20,21]. In line with the results in transgenic AD mouse models [12, 13], these results suggest that BDNFVal66Met influences in particular the impact of AD pathology on hippocampal function and memory. Yet, the impact of BDNFVal66Met at the level of functional networks that support cognitive function is unknown. Thus, our major aim was to assess whether BDNFVal66Met moderates the impact of AD pathology on functional connectivity within major networks and associated cognitive impairment in AD.
The brain is composed of major large-scale functional networks that become engaged during cognitive tasks. Resting-state fMRI studies assessing the integrity of these networks in AD have shown reduced connectivity between the hippocampus and medial-frontal and posterior-parietal regions of the default-mode network (DMN) [24,25,26,27,28]. Higher Aβ-levels were associated with lower hippocampus connectivity underlying memory impairment in preclinical and prodromal AD patients [29, 30]. Here, we hypothesized that in BDNFVal66Met carriers, the impact of higher Aβ pathology on hippocampus connectivity and memory impairment is worse compared to homozygous BDNFVal carriers. We tested this hypothesis in three independently recruited samples of subjects with either genetically caused AD or biomarker evidence of Aβ. In a discovery sample of 115 mutation carriers with autosomal dominant AD (ADAD) and 91 familial non-carrier controls from the Dominantly Inherited Alzheimer Network (DIAN) [31], we explored in which functional network BDNFVal66Met carriers showed altered connectivity compared to homozygous BDNFVal carriers.
For our hypothesis-driven analysis we assessed connectivity alterations in the hippocampal network, and for exploratory purposes, in a set of other major functional networks (i.e. DMN, dorsal attention [DAN], salience [SAL], fronto-parietal control [CON]) that are part of the canonical set of resting-state networks associated with higher cognitive function and previously found altered in AD [32, 33]. We specifically conducted the discovery analyses in ADAD, since the study design of DIAN allows to examine the effect of BDNFVal66Met on connectivity changes in genetically caused AD where aging-related comorbidities such as cerebrovascular changes are unlikely. Based on the results on the association between BDNFVal66Met and functional network changes in ADAD, confirmatory analyses were subsequently conducted in elderly subjects with biomarker evidence of sporadic AD, including 149 subjects with subjective cognitive decline (SCD) from the Spanish FACEHBI study [34] and in 114 subjects ranging from cognitively normal to AD dementia assessed within the German DELCODE study [35]. The inclusion of these additional samples with sporadic AD pathophysiology provided the opportunity to test whether any effects of BDNFVal66Met on connectivity can be generalized towards the more common age-related form of accumulating AD pathology in elderly subjects. Since BDNFVal66Met has been previously associated with greater cognitive impairment in both autosomal dominant and sporadic AD [17, 19, 20], we tested in a last step whether any observed BDNFVal66Met-related functional network alterations are associated with worse global cognition and episodic memory.
Methods
Participants
DIAN
One hundred and fifteen carriers of ADAD-causing mutations in genes PSEN1 (n = 82) and PSEN2 (n = 12) or APP (n = 21), and 91 non-carrier (NC) siblings were included from DIAN data freeze 10 [31]. Beyond DIAN inclusion criteria, the current study required availability of BDNFVal66Met genotype, resting-state fMRI, T1-structural MRI, and cognitive assessments. No selection bias (i.e. demographic differences between the included subjects and excluded subjects) was found (p > 0.05) for age, gender, or education. As a marker of AD severity, we applied the estimated years from symptom onset (EYO), defined as the difference between a participants age at examination and the parental age of symptom onset for ADAD mutation carriers, as described previously [36,37,38]. As an additional marker of amyloid pathology, we employed global PiB-PET SUVR scores (normalized to the whole cerebellum, available in a subsample of 100 mutation carriers and 87 non-carriers) provided by the DIAN core, where subjects were binarized as Aβ+ at a SUVR threshold >1.31 following recommendations by the DIAN core. For details on PiB-PET assessment in DIAN, please refer to a previous publication [39]. BDNFVal66Met genotyping was performed on whole-blood samples using the Infinium HumanExomeCore V1.0 Beadchip (Illumina, Inc.) at Washington University. Each participant provided written informed consent. Local ethical approval was obtained at each DIAN site.
DELCODE
One hundred and fourteen older adults (>60 years) were included from the German multicenter DELCODE cohort on sporadic AD (data freeze 1, N = 366) [35]. Beyond DELCODE inclusion criteria, subjects had to have available BDNFVal66Met data, resting-state fMRI and T1-structural MRI, cerebrospinal fluid (CSF)-assessed Aβ-levels (Aβ42/40 ratio), and cognitive assessments. Testing for selection bias yielded no significant (p > 0.05) differences between the selected and excluded subjects for age, gender, and education. Subjects met classification criteria for cognitively normal (CN), SCD (i.e. normal cognitive performance and SCD between the last 6 months and 5 years), MCI, or AD dementia, following previously described procedures [35, 37]. Continuous CSF Aβ42/40 ratio levels, i.e. a close correlate of brain Aβ-levels [40], were used as a marker of AD pathology. For descriptive purposes Aβ-positivity was defined as CSF Aβ42/40<0.1, following pre-established cut-points [41] used within DELCODE [37]. BDNFVal66Met genotyping was performed on whole-blood samples using commercially available TaqMan probes (ThermoFischer). All subjects provided written informed consent; local ethical approval was obtained at each DELCODE site.
FACEHBI
One hundred and forty-nine older adults (>50 years) were included from the Spanish monocentric FACEHBI cohort (n = 214) collected at the Fundació ACE Alzheimer Treatment and Research Center in Barcelona. All subjects met research criteria for SCD as defined by the FACEHBI core (i.e. coexistence of subjective cognitive complaints defined as a score >7 on the memory failures in everyday life questionnaire [42], and normal performance in a comprehensive neuropsychological battery) [43]. For a detailed sample description and inclusion criteria please see a previous publication [34]. For the current study, subjects were included based on availability of BDNFVal66Met genotype, resting-state fMRI, T1-structural MRI 18[F]-Florbetaben amyloid-PET and cognitive data. No significant differences (p > 0.05) were found in age, gender, or education between the selected and excluded subjects. AD pathology was assessed using continuous measures of global 18[F]-Florbetaben PET SUVR scores, which were additionally stratified into Aβ-positive/negative for descriptive purposes, using a standard uptake value ratio (SUVR) threshold >1.45, following previous recommendations [44]. The assessment of global Aβ-PET SUVR has been conducted by the FACEHBI core and was described in detail previously [45]. The BDNFVal66Met genotype was extracted from GWAS data performed on whole-blood samples with the Illumina Infinium Omni Express Exome-8v1.3 chip. All subjects provided written informed consent; ethical approval was obtained by the FACEHBI principal investigators.
Neuropsychology
As a primary measure of cognition, we used the Mini-Mental State Exam (MMSE) [46] which was consistently available in all samples, thus facilitating across cohort comparability. As a secondary measure of cognition, we used episodic memory as assessed by the delayed recall score of the Wechsler logical memory scale [47], which was available in DIAN and DELCODE sample but not in FACEHBI.
MRI acquisition
All MRI scans were recorded on 3T scanners for DIAN and DELCODE, and a 1.5T scanner for FACEHBI. Within the multicenter studies DIAN or DELCODE, scanning protocols were harmonized across participating sites for each study. Here, we included resting-state fMRI and high-resolution T1-structural MRI scans. Detailed sequence parameters can be found in Supplementary Table 1.
Preprocessing of structural and resting-state fMRI
The same pre-established SPM12-based pipeline previously described by us [37, 48, 49], including motion correction, nuisance regression (motion, white-matter, and CSF), censoring of high-motion frames, spatial normalization, and smoothing, was applied to each cohort (DIAN, DELCODE, FACEHBI) separately so no data were pooled across cohorts during the entire analyses. For details on MRI preprocessing please see Supplementary Methods.
Assessment of seed-based connectivity
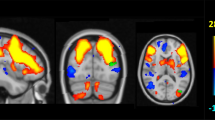
We used a seed-based connectivity approach previously described by us [37, 48, 49] in order to generate subject-specific Fisher z-transformed connectivity maps of hippocampal seeds and canonical resting-state networks (DMN, DAN, SAL, CON) for each subject from preprocessed resting-state fMRI data. For details on anatomical location of seed regions of interest (ROIs) and network topology please see Fig. 1 and Supplementary Methods.
Surface renderings of significant seed-based connectivity (p < 0.001, FWE cluster corrected at p < 0.05) in the DIAN discovery sample for the primary hippocampal seed ROIs and the secondary seed ROIs to derive connectivity for the control network (CON), dorsal attention network (DAN), default mode network (DMN), and salience network (SAL)
Statistical analysis
Within each sample, we compared baseline group differences using two-sample t-tests or ANOVAS (for >2 groups) for continuous measures and Chi-squared tests for categorical measures. We further tested BDNFVal66Met deviations from the Hardy–Weinberg equilibrium within each sample. Significant seed-based connectivity in DIAN was mapped by conducting voxel-wise t-tests against zero on the subject-specific connectivity maps (α = 0.001 and FWE cluster correction).
Next, we computed the voxel-wise interaction BDNFVal66Met × EYO on seed-based connectivity maps in the 115 ADAD mutation carriers (DIAN), controlling for gender, center, education, family affiliation, and subject-specific gray matter volume of the seed ROI that was extracted from spatially normalized and Jacobian-scaled gray matter images. The rationale to use EYO as a marker of disease severity in DIAN is that (1) EYO is a valid and reliable marker of global AD pathology and (2) that usage of EYO facilitates the comparability with previous studies in this cohort. In line with previous studies in DIAN we did not control for age in the model [37, 38], as age is highly correlated with EYO and may mask variance of interest and/or induce multicollinearity. Clusters showing a significant BDNFVal66Met × EYO interaction (α = 0.001 and FWE cluster correction at α = 0.05) were binarized to extract subject-specific mean connectivity values from individual connectivity maps (henceforth referred to as BDNF-related connectivity) for later analyses. Using mean connectivity values, we further conducted confirmatory analyses in a subset of 100 mutation-carrier subjects with available global amyloid-PET SUVR levels instead of EYO, to ensure that our results were not specific for using EYO as a measure of ADAD severity. Specifically, we tested the interaction global amyloid-PET SUVR × BDNFVal66Met on connectivity, controlling for gender, center, education, family affiliation, and gray matter volume of the respective seed ROI.
After the discovery analysis in DIAN, cluster maps of significant EYO × BDNFVal66Met interactions were forward applied to the DELCODE and FACEHBI groups to extract mean seed ROI to cluster connectivity values for validation analysis including the interaction BDNFVal66Met × amyloid load on connectivity. As a measure of amyloid load, we applied continuous CSF measures of the Aβ42/40 ratio for DELCODE and continuous global amyloid-PET SUVR scores for FACEHBI. Due to the skewed distribution of global amyloid-PET SUVRs, the scores were Box-Cox transformed prior to analysis, after which there was no deviation from a normal distribution. Using separate linear models for DELCODE and FACEHBI, we then tested the Aβ × BDNFVal66Met two-way interaction on connectivity, controlling for age, gender, education, ROI gray matter volume, ApoE4 status (as well as center and diagnosis for DELCODE).
We further extracted connectivity to clusters in the ADAD non-carrier group (DIAN), to assess potential BDNFVal66Met-related connectivity changes in relatively young subjects unaffected by ADAD (i.e. BDNFVal66Met × Age on connectivity, controlling for gender, family and gray matter volume of the respective seed ROI). The rationale for this analysis is, that the BDNFVal66Met genotype may also have an effect on connectivity changes in subjects unaffected by ADAD.
Next, we tested separately within the ADAD subjects as well as in DELCODE and FACEHBI whether BDNF-related hippocampal connectivity (i.e. seed connectivity to clusters that showed a significant BDNFVal66Met × EYO interaction in DIAN) moderated AD-related cognitive decreases. For DELCODE and FACEHBI, we used robust linear mixed-effects models, where we tested within each sample the two-way interaction between BDNF-related hippocampal-medio-frontal connectivity and amyloid on (1) episodic memory or (2) global cognition controlling for gender and hippocampal volume. Due to the study design of DELCODE we additionally included diagnosis as a fixed effect and center as a random effect in the respective statistical models. MMSE values were box-cox transformed prior to analyses to minimize skew. For DIAN, analogous regression analyses were run using MMSE or episodic memory as dependent variables, testing the two-way interaction between hippocampal-medial-frontal connectivity and EYO, controlling for family affiliation (to control for shared genetic background among DIAN family members), gender, and hippocampal volume.
All voxel-wise analyses were computed in SPM12 and restricted to group-specific gray matter masks. Statistical analyses were conducted using R statistical software. Effects were considered significant when meeting a two-tailed alpha threshold of α = 0.05.
Results
See Table 1 for sample characteristics. Patterns of significant seed-based connectivity in the DIAN discovery sample are shown in Fig. 1. No BDNFVal66Met deviations from Hardy–Weinberg equilibrium were found.
Discovery analysis: Interaction BDNFVal66Met × EYO on hippocampal connectivity in ADAD (DIAN)
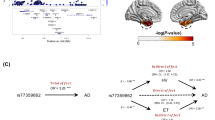
We found a significant BDNFVal66Met × EYO interaction on connectivity between the right hippocampus and the bilateral medial-frontal cortex (henceforth referred to right hippocampal-medial-frontal connectivity), a key region of the DMN (MNI: x = −2, y = 33, z = 14, t(108) = 4.77, blue and purple areas in Fig. 2a). When illustrating the interaction effect using mean connectivity values of that cluster, BDNFVal was associated with relatively stable right hippocampal-medial-frontal connectivity levels across EYO, whereas BDNFVal66Met was associated with a decline in right hippocampal-medial-frontal connectivity across EYO (Fig. 2b). For the left hippocampus seed ROI, the BDNFVal66Met × EYO interaction was also detected for the bilateral medial-frontal cortex (Fig. 2e, MNI: x = 4, y = 49, z = −10, t(109) = 3.93), where BDNFVal was associated with relatively preserved connectivity across EYO compared to BDNFVal66Met (Fig. 2f). For the sake of consistency with the subsequent validation analyses in DELCODE and FACEHBI using continuous biomarkers of Aβ as the marker of AD pathology (see below), we repeated the analyses using global amyloid-PET levels instead of EYO as the predictor variable in a subset of 100 ADAD subjects with available PiB-PET. Results of the regression analysis showed a congruent interaction between global amyloid-PET SUVR and BDNFVal66Met on both left (t(94) = 3.074, β/SE = 0.066/0.021, p = 0.002) and right (t(94) = 3.873, β/SE = 0.083/0.021, p < 0.001) hippocampus-medial-frontal connectivity.
Results of the voxel-wise interaction analysis of EYO × BDNFVal66Met on hippocampal connectivity in the DIAN-MC subjects for the right (a, b) and left (e, f) hippocampus seed ROI (i.e. discovery). In panels a and e, red areas correspond to regions showing significant seed-based hippocampal connectivity, while purple and blue regions indicate the boundaries of the significant EYO × BDNFVal66Met interaction. Validation analyses of the BDNFVal66Met × Aβ interaction in DELCODE and FACEHBI using the DIAN-MC derived medial-frontal ROI are shown in panels c and g for DELCODE and d and h for FACEHBI. DIAN-MC = mutation carriers with autosomal dominant AD from DIAN
No Age × BDNFVal66Met interaction effects for right or left hippocampal connectivity were detected in the ADAD non-carriers, suggesting specificity for ADAD. Also, no main effect of age on hippocampal connectivity was found in this group, suggesting that hippocampus connectivity remains relatively stable across age in the absence of ADAD. This view is further confirmed by an exploratory assessed ADAD mutation (i.e. carriers vs. non-carriers) × age interaction on both left (t(196) = −2.087, β/SE = −0.005/0.002, p = 0.038) and right (t(196) = −2.201, β/SE = −0.006/0.003, p = 0.029) hippocampus-medial-frontal connectivity, when controlling for gender, seed ROI volume, family, and BDNFVal66Met. Here, ADAD mutation carriers show decreasing connectivity across age (which is highly correlated with EYO) while non-carriers remain relatively stable. These results suggest that the hippocampus-medial-frontal connectivity decreases as observed in the ADAD group are driven by AD pathology and do not reflect normal age-related connectivity changes.
In addition, none of the other resting-state networks (DMN, DAN, SAL, CON) showed a significant BDNFVal66Met × EYO interaction, both in ADAD mutation carriers and non-carriers (where we used age instead of EYO).
Validation analyses: BDNFVal66Met and hippocampal connectivity in subjects with sporadic Aβ pathology (DELCODE and FACEHBI)
To validate our findings of the association between BDNFVal66Met and hippocampal-medial-frontal connectivity in ADAD, we assessed the interaction BDNFVal66Met × Aβ on hippocampus-medial-frontal connectivity (Fig. 2a, e) in the independent DELCODE and FACEHBI samples. Here, we employed Aβ-levels to assess AD severity, since Aβ is the defining feature of sporadic AD pathology in the mostly non-demented DELCODE and FACEHBI subjects.
By considering the clinical spectrum of CN, SCD, MCI, and AD dementia in DELCODE, we found a significant BDNFVal66Met × CSF Aβ42/40 interaction on both right hippocampal-medial-frontal (β/SE = −0.404/0.179, p = 0.026; Fig. 2c) and left hippocampal-medial-frontal connectivity (β/SE = −0.355/0.169, p = 0.037; Fig. 2g), where BDNFVal was associated with more stable connectivity across a decreasing CSF Aβ42/40 ratio compared to BDNFVal66Met. In FACEHBI (SCD subjects), significant BDNFVal66Met × global amyloid-PET SUVR interactions were observed on right hippocampal-medial-frontal (β/SE = 0.301/0.170, p = 0.039; Fig. 2d) and left hippocampal-medial-frontal connectivity (β/SE = 0.333/0.195, p = 0.045; Fig. 2h). In exploratory analyses, no BDNFVal66Met × global amyloid-PET SUVR interactions were found for any of the other functional networks in both DELCODE or FACEHBI. These results suggest congruent effects of BDNFVal66Met on hippocampal connectivity in all three samples. Regression results are summarized in Table 2.
BDNFVal66Met-related hippocampal-medial-frontal connectivity moderates the effect of AD severity on cognition
We next tested whether BDNF-related hippocampus-medial-frontal connectivity was associated with cognition. In DIAN, ADAD mutation carriers showed for both right and left hippocampal-medial-frontal connectivity (i.e. Figure 2a, e) a significant two-way interaction with EYO on MMSE, when controlling for gender, education, family affiliation, respective hippocampal volume and center (right: β/SE = 0.202/0.047, p < 0.001, left: β/SE = 0.177/0.068, p = 0.009). As hypothesized, individuals with lower right and left hippocampal-medial-frontal connectivity showed steeper MMSE decreases across EYO than individuals with higher FC. A hippocampal-medial-frontal connectivity × EYO interaction was also detected for logical memory recall (right: β/SE = 0.144/0.071, p = 0.045, Fig. 3b; left: β/SE = 0.160/0.077, p = 0.041), with worse memory impairment at lower hippocampal-medial-frontal connectivity.
Scatterplots illustrating the interaction between right (a–d) and left (e–h) hippocampus to medial-frontal connectivity and AD severity (i.e. EYO or CSF Aβ42/40 ratio) on cognition. Color groupings were based on median split and are for illustrational purposes only, since the underlying linear mixed models were computed using continuous measures. DIAN-MC = mutation carriers with autosomal dominant AD from DIAN
In DELCODE, we found a significant two-way interaction such that individuals with lower right hippocampal-medial-frontal (β/SE = −0.222/0.080, p = 0.007; Fig. 3c) or left hippocampal-medial-frontal connectivity (β/SE = −0.205/0.079, p = 0.011) showed stronger MMSE reductions as a function of more abnormal CSF Aβ42/40-levels, controlled for diagnosis among other potentially confounding variables. All results remained consistent when additionally controlling for ApoE4 status. No interaction was found, however, for logical memory (Fig. 3d, Supplementary Table 2). In the FACEHBI sample, no hippocampal-medial-frontal connectivity × global amyloid-PET SUVR two-way interaction effects on MMSE were found, probably due to lack of sufficient variance of cognitive performance in this cognitively normal SCD group.
Discussion
Our major finding, observed across three independent cohorts, was that BDNFVal66Met was associated with stronger AD pathology-related decreases of hippocampal-medial-frontal connectivity. Furthermore, we found that lower hippocampal-medial-frontal connectivity was associated with stronger reductions in global cognition in both ADAD and symptomatic sporadic AD. Thus, our results suggest that the variation in BDNFVal66Met moderates the effect of AD pathology on hippocampus functional connectivity. Although the moderating effects of BDNFVal66Met-related hippocampus connectivity on cognition were not strong in the current study, our results provide preliminary evidence that hippocampal-mediofrontal-connectivity alterations linked to BDNFVal66Met may contribute to global cognitive impairment in AD.
In the current study, the association between BDNFVal66Met and decreased hippocampal-medial-frontal connectivity has been observed across different samples including non-symptomatic and symptomatic (MCI and dementia) AD stages, suggesting robust effects of BDNFVal66Met on regional hippocampal connectivity alterations across the AD spectrum. These results remained after accounting for hippocampus atrophy, suggesting that the association between BDNFVal66Met and hippocampal connectivity alterations cannot be fully explained by hippocampal neuronal loss. Strikingly, the association between BDNFVal66Met and stronger connectivity decreases was confined to the hippocampal networks since exploratory analyses of other major functional networks did not yield any significant effects. These findings are consistent with previous reports suggesting that BDNF-related alterations occur specifically in the hippocampus rather than other brain regions. In healthy subjects, BDNFVal66Met was associated with reduced hippocampal activation during an encoding task, but showed no effect on a wider task-activated cortical network [50]. Furthermore, in ADAD, more pronounced FDG-PET hypometabolism within the hippocampus, but not precuneus, was observed in BDNFVal66Met-carriers compared to BDNFVal homozygotes [20]. Together, these results suggest that BDNFVal66Met is specifically associated with hippocampal alterations. Given that the hippocampus is among the earliest regions to be affected by AD [51, 52], showing early connectivity decreases [32, 53], the design of the current study may have favored statistical power to detect BDNFVal66Met effects specifically within the hippocampus as most subjects had only mild AD. Strikingly, the moderating effects of BDNFVal66Met on the association between AD severity and hippocampal connectivity decreases were strongest in the ADAD group, as compared to the sporadic AD groups. This pronunciation of effects in ADAD might be due sample specific effects in the DIAN cohort. Thus, the effect size in the validation samples reflects probably a more likely approximation of the true effect size. Here, a large-scale study would be needed in order to conduct sensitivity analyses of any potential factors that may influence the observed effect size.
Connectivity between the hippocampus and medial-frontal cortex is supported anatomically by direct neuronal connections [54]. Major mono-synaptic bidirectional connections between the orbitofrontal cortex and the peri- and entorhinal cortices, i.e. a major connection to the hippocampus, are constituted by the uncinate fasciculus as found in both post-mortem tracer studies and in vivo diffusion-tensor imaging studies [55]. Functional imaging studies have shown that hippocampal-orbitofrontal connections are part of a network that is activated during cognitive tasks tapping episodic [56,57,58] and working memory [59, 60]. Alterations in hippocampal-prefrontal connectivity have been previously associated with reduced episodic- or working memory in AD [55, 61, 62] or psychiatric diseases including schizophrenia [63, 64], suggesting that this anatomically supported functional network is critical for a wider range of cognitive abilities. We found that BDNFVal66Met-related hippocampal-medial-frontal connectivity was associated with greater impairment in episodic memory and global cognition in ADAD, and stronger associations between Aβ (as measured by CSF Aβ42/40) and impaired global cognition in a group covering the AD spectrum (DELCODE). However, no effect of BDNFVal66Met-related hippocampal-medial-frontal connectivity on the association between Aβ (as assessed by amyloid-PET) and global cognition was found in subjects with SCD (FACEHBI). The variability of findings between studies may be due to small variability in cognitive performance and AD-related cognitive changes in SCD, requiring larger sample sizes to detect any small effects of BDNFVal66Met in cognitively normal subjects. Together, the current results suggest that BDNFVal66Met enhances the vulnerability of this hippocampal-frontal memory network to the effects of Aβ and may thus worsen cognitive decline in symptomatic AD.
While we caution that the current results do not provide evidence for causal effects of BDNF genetic variants on functional connectivity, we encourage future studies to examine the mechanisms that underlie the potential effects of BDNFVal66Met in AD. One possibility is that BDNFVal66Met is associated with disturbed BDNF secretion, since preclinical in vitro and in vivo studies showed that BDNFVal66Met is associated with impaired intracellular BDNF trafficking, entailing decreased synaptic BDNF levels [14]. Since LTP may enhance connectivity between neural populations detectable by fMRI [65] and BDNFVal66Met-related reduced BDNF bioavailability impairs LTP [15], BDNFVal66Met may contribute to higher susceptibility of hippocampal connectivity to detrimental Aβ-effects. Note that we detected no difference between BDNFVal66Met carriers and BDNFVal-carriers in Aβ-biomarker levels. These findings are consistent with previous reports of the absence of altered BDNFVal66Met effects on CSF Aβ1-42 in ADAD [20, 21] or sporadic AD [17, 66]. These results are also in line with previous findings in transgenic AD mouse models, where lentiviral BDNF-gene delivery reduced cell loss and improved learning performance, but did not alter Aβ-burden [13]. Similarly, reduced BDNF levels were found in the P301L transgenic mouse model of tau pathology, where adeno-virus induced restoration of BDNF levels alleviated synaptic degeneration and spatial memory deficits, but did not attenuate tau pathology [12]. Together, these findings suggest that BDNF is not associated with alterations in the primary pathology itself, rather lower BDNF (as proxied by BDNFVal66Met) may be associated with enhanced susceptibility of hippocampus connectivity to the effects of AD pathology.
For drawing conclusions based on the current results, some caveats should be considered. We caution that BDNFVal66Met effects on BDNF levels in the brain of AD patients are not clear, and we did not assess BDNF protein levels directly. Previous studies reported decreased brain BDNF mRNA levels in AD mouse models [67] and AD patients [10] as well as decreases of CSF-derived BDNF levels in AD patients, but CSF or serum BDNF levels were not altered by BDNFVal66Met [68, 69]. The latter studies, however, included mostly healthy controls and only few MCI and AD cases, rendering the results difficult to interpret. BDNFVal66Met effects on CSF or serum-derived BDNF levels in MCI and AD remain to be investigated in future studies.
It should be acknowledged that we included a total of three different international studies using slightly different inclusion/exclusion criteria, MRI hardware, and scanning protocols, which were not a priori harmonized across studies and can thus be considered a “naturalistic” sample of AD subjects. However, the heterogeneity in the sampling may also entail increased variability in the variable measurements and limited comparability. For instance, the FACEHBI 1.5 T scanning protocol is potentially less sensitive to detect BOLD signal changes compared to the DIAN and DELCODE 3 T scanning protocols [70]. In order to enhance comparability between studies, we applied harmonized MRI preprocessing and analysis pipelines and selected uniformly available cognitive tests. Importantly, however, rather than pooling data across all samples, where variability of the data may hamper the estimation of the regression parameters, we used a cross-validation approach in order to test the robustness and external validity of our initial analysis in the ADAD sample. Thus, our highly consistent findings across cohorts reduce the likelihood of our results being driven by technical assessment procedures or selection criteria.
A further potential limitation is that ADAD and sporadic AD may show slight differences in disease development. While ADAD and sporadic AD share core neuropathological and clinical features [71] and functional network changes [72] (for a review, see [73]), several differences have been reported as well: Compared to sporadic AD, ADAD is associated with an earlier symptom onset [74], stronger striatal amyloid accumulation [75], higher likelihood of non-memory cognitive deficits [74], higher prevalence of psychosis and hallucinations [71], spastic paraparesis and motor symptoms [76, 77]. Given such differences between ADAD and sporadic AD, it is possible that our approach of using the data from ADAD as the discovery sample may entail missing some BDNFVal66Met-associated brain network effects that are specific to sporadic AD. To address this caveat, we tested in exploratory secondary analyses the interaction effect of BDNFVal66Met × Aβ on each resting-state network in both the DELODE and FACEHBI studies. The absence of any interaction effect of the BDNFVal66Met genotype by Aβ biomarker levels on any network other than the hippocampal network supports the conclusion that the effects of BDNFVal66Met on exclusively hippocampal connectivity are consistent across samples, despite potential clinico-pathological differences between ADAD and sporadic AD.
Lastly, we did not assess tau pathology in the current study. Tau deposition emerges first in the entorhinal cortex and hippocampus [78] and has been shown to be correlated with cognitive performance and hippocampus connectivity in humans [79]. Given previous preclinical evidence that BDNF may modulate the neurotoxic effects of tau [12], BDNFVal66Met may also alter the effect of tau on hippocampus function. Additionally, previous studies of ADAD have reported that BDNFVal66Met-carriers show faster CSF tau and p-tau181 increases [21]. The recent development of tau-PET will allow future studies to assess regional tau levels in the brain [80], and provide an opportunity to test the potential modulating effects of BDNFVal66Met on hippocampus connectivity in future studies.
Conclusions
In summary, our results suggest that BDNFVal66Met is associated with increased impairment of hippocampal-medial-frontal connectivity cortex in the presence of AD pathology.
Our current results are consistent with previous studies suggesting that BDNFVal66Met may moderate the detrimental effects of Aβ on hippocampal connectivity and memory impairment in AD. A critical future question will be, whether the effects of BDNFVal66Met show a disease stage-dependent peak, i.e. at the preclinical, prodromal, or dementia stage of AD. We thus encourage future studies to assess this question as soon as large enough data become available for each subgroup. Our results have clinical implications: BDNF levels are modifiable in humans and could thus become a promising treatment target to enhance resilience against the impact of neurotoxic primary AD pathology [9]. Alterations of hippocampal connectivity can be a potential outcome measure to assess the effect of BDNF-targeting drugs in AD. Increased BDNF levels after aerobic exercise have been observed in subjects with neurological disorders [81], and this may mediate the effect of exercise on enhanced synaptic plasticity and hippocampus-dependent learning [82]. Enhancing BDNF levels in the brain may thus constitute a promising secondary-preventive approach to attenuate AD-effects on brain integrity and cognitive function [9, 83].
References
Bekinschtein P, Cammarota M, Katche C, Slipczuk L, Rossato JI, Goldin A, et al. BDNF is essential to promote persistence of long-term memory storage. Proc Natl Acad Sci USA. 2008;105:2711–6.
Song M, Martinowich K, Lee FS. BDNF at the synapse: why location matters. Mol Psychiatry. 2017;22:1370–5.
Hall J, Thomas KL, Everitt BJ. Rapid and selective induction of BDNF expression in the hippocampus during contextual learning. Nat Neurosci. 2000;3:533–5.
Murer MG, Yan Q, Raisman-Vozari R. Brain-derived neurotrophic factor in the control human brain, and in Alzheimer's disease and Parkinson's disease. Prog Neurobiol. 2001;63:71–124.
Lu B, Nagappan G, Lu Y. BDNF and synaptic plasticity, cognitive function, and dysfunction. Handb Exp Pharmacol. 2014;220:223–50.
Scheff SW, Price DA, Schmitt FA, Mufson EJ. Hippocampal synaptic loss in early Alzheimer's disease and mild cognitive impairment. Neurobiol Aging. 2006;27:1372–84.
de Wilde MC, Overk CR, Sijben JW, Masliah E. Meta-analysis of synaptic pathology in Alzheimer's disease reveals selective molecular vesicular machinery vulnerability. Alzheimers Dement. 2016;12:633–44.
Schuff N, Woerner N, Boreta L, Kornfield T, Shaw LM, Trojanowski JQ, et al. MRI of hippocampal volume loss in early Alzheimer's disease in relation to ApoE genotype and biomarkers. Brain. 2009;132(Pt 4):1067–77.
Lu B, Nagappan G, Guan X, Nathan PJ, Wren P. BDNF-based synaptic repair as a disease-modifying strategy for neurodegenerative diseases. Nat Rev Neurosci. 2013;14:401–16.
Michalski B, Corrada MM, Kawas CH, Fahnestock M. Brain-derived neurotrophic factor and TrkB expression in the "oldest-old," the 90+ Study: correlation with cognitive status and levels of soluble amyloid-beta. Neurobiol Aging. 2015;36:3130–9.
Nagahara AH, Merrill DA, Coppola G, Tsukada S, Schroeder BE, Shaked GM, et al. Neuroprotective effects of brain-derived neurotrophic factor in rodent and primate models of Alzheimer's disease. Nat Med. 2009;15:331–7.
Jiao SS, Shen LL, Zhu C, Bu XL, Liu YH, Liu CH, et al. Brain-derived neurotrophic factor protects against tau-related neurodegeneration of Alzheimer's disease. Transl Psychiatry. 2016;6:e907.
Nagahara AH, Mateling M, Kovacs I, Wang L, Eggert S, Rockenstein E, et al. Early BDNF treatment ameliorates cell loss in the entorhinal cortex of APP transgenic mice. J Neurosci. 2013;33:15596–602.
Egan MF, Kojima M, Callicott JH, Goldberg TE, Kolachana BS, Bertolino A, et al. The BDNF val66met polymorphism affects activity-dependent secretion of BDNF and human memory and hippocampal function. Cell. 2003;112:257–69.
Hao R, Qi Y, Hou DN, Ji YY, Zheng CY, Li CY, et al. BDNF val66met polymorphism impairs hippocampal long-term depression by down-regulation of 5-HT3 receptors. Front Cell Neurosci. 2017;11:306.
Petryshen TL, Sabeti PC, Aldinger KA, Fry B, Fan JB, Schaffner SF, et al. Population genetic study of the brain-derived neurotrophic factor (BDNF) gene. Mol Psychiatry. 2010;15:810–5.
Lim YY, Villemagne VL, Laws SM, Ames D, Pietrzak RH, Ellis KA, et al. BDNF Val66Met, Abeta amyloid, and cognitive decline in preclinical Alzheimer's disease. Neurobiol Aging. 2013;34:2457–64.
Lim YY, Villemagne VL, Laws SM, Ames D, Pietrzak RH, Ellis KA, et al. Effect of BDNF Val66Met on memory decline and hippocampal atrophy in prodromal Alzheimer's disease: a preliminary study. PLoS One. 2014;9:e86498.
Lim YY, Villemagne VL, Laws SM, Pietrzak RH, Snyder PJ, Ames D, et al. APOE and BDNF polymorphisms moderate amyloid beta-related cognitive decline in preclinical Alzheimer's disease. Mol Psychiatry. 2015;20:1322–8.
Lim YY, Hassenstab J, Cruchaga C, Goate A, Fagan AM, Benzinger TL, et al. BDNF Val66Met moderates memory impairment, hippocampal function and tau in preclinical autosomal dominant Alzheimer's disease. Brain. 2016;139(Pt 10):2766–77.
Lim YY, Hassenstab J, Goate A, Fagan AM, Benzinger TLS, Cruchaga C, et al. Effect of BDNFVal66Met on disease markers in dominantly inherited Alzheimer's disease. Ann Neurol. 2018;84:424–35.
Kambeitz JP, Bhattacharyya S, Kambeitz-Ilankovic LM, Valli I, Collier DA, McGuire P. Effect of BDNF val(66)met polymorphism on declarative memory and its neural substrate: a meta-analysis. Neurosci Biobehav Rev. 2012;36:2165–77.
Harrisberger F, Spalek K, Smieskova R, Schmidt A, Coynel D, Milnik A, et al. The association of the BDNF Val66Met polymorphism and the hippocampal volumes in healthy humans: a joint meta-analysis of published and new data. Neurosci Biobehav Rev. 2014;42:267–78.
Park KH, Noh Y, Choi EJ, Kim H, Chun S, Son YD. Functional connectivity of the hippocampus in early- and vs. late-onset alzheimer's disease. J Clin Neurol. 2017;13:387–93.
Allen G, Barnard H, McColl R, Hester AL, Fields JA, Weiner MF, et al. Reduced hippocampal functional connectivity in Alzheimer disease. Arch Neurol. 2007;64:1482–7.
Greicius MD, Srivastava G, Reiss AL, Menon V. Default-mode network activity distinguishes Alzheimer's disease from healthy aging: evidence from functional MRI. Proc Natl Acad Sci USA. 2004;101:4637–42.
Tahmasian M, Pasquini L, Scherr M, Meng C, Forster S, Mulej Bratec S, et al. The lower hippocampus global connectivity, the higher its local metabolism in Alzheimer disease. Neurology. 2015;84:1956–63.
Sorg C, Riedl V, Muhlau M, Calhoun VD, Eichele T, Laer L, et al. Selective changes of resting-state networks in individuals at risk for Alzheimer's disease. Proc Natl Acad Sci USA. 2007;104:18760–5.
Pasquini L, Scherr M, Tahmasian M, Meng C, Myers NE, Ortner M, et al. Link between hippocampus' raised local and eased global intrinsic connectivity in AD. Alzheimers Dement. 2015;11:475–84.
La Joie R, Landeau B, Perrotin A, Bejanin A, Egret S, Pelerin A, et al. Intrinsic connectivity identifies the hippocampus as a main crossroad between Alzheimer's and semantic dementia-targeted networks. Neuron. 2014;81:1417–28.
Moulder KL, Snider BJ, Mills SL, Buckles VD, Santacruz AM, Bateman RJ, et al. Dominantly inherited Alzheimer network: facilitating research and clinical trials. Alzheimers Res Ther. 2013;5:48.
Jones DT, Knopman DS, Gunter JL, Graff-Radford J, Vemuri P, Boeve BF, et al. Cascading network failure across the Alzheimer's disease spectrum. Brain. 2016;139(Pt 2):547–62.
Thomas JB, Brier MR, Bateman RJ, Snyder AZ, Benzinger TL, Xiong C, et al. Functional connectivity in autosomal dominant and late-onset Alzheimer disease. JAMA Neurol. 2014;71:1111–22.
Rodriguez-Gomez O, Sanabria A, Perez-Cordon A, Sanchez-Ruiz D, Abdelnour C, Valero S, et al. FACEHBI: a prospective study of risk factors, biomarkers and cognition in a cohort of individuals with subjective cognitive decline. study rationale and research protocols. J Prev Alzheimers Dis. 2017;4:100–8.
Jessen F, Spottke A, Boecker H, Brosseron F, Buerger K, Catak C, et al. Design and first baseline data of the DZNE multicenter observational study on predementia Alzheimer's disease (DELCODE). Alzheimer's Res Ther. 2018;10:15.
Bateman RJ, Xiong C, Benzinger TL, Fagan AM, Goate A, Fox NC, et al. Clinical and biomarker changes in dominantly inherited Alzheimer's disease. N Engl J Med. 2012;367:795–804.
Franzmeier N, Duzel E, Jessen F, Buerger K, Levin J, Duering M, et al. Left frontal hub connectivity delays cognitive impairment in autosomal-dominant and sporadic Alzheimer's disease. Brain. 2018;141:1186–1200.
Suarez-Calvet M, Araque Caballero MA, Kleinberger G, Bateman RJ, Fagan AM, Morris JC, et al. Early changes in CSF sTREM2 in dominantly inherited Alzheimer's disease occur after amyloid deposition and neuronal injury. Sci Transl Med. 2016;8:369ra178.
Benzinger TL, Blazey T, Jack CR Jr., Koeppe RA, Su Y, Xiong C, et al. Regional variability of imaging biomarkers in autosomal dominant Alzheimer's disease. Proc Natl Acad Sci USA. 2013;110:E4502–4509.
Lewczuk P, Matzen A, Blennow K, Parnetti L, Molinuevo JL, Eusebi P, et al. Cerebrospinal fluid Abeta42/40 corresponds better than Abeta42 to amyloid PET in Alzheimer's disease. J Alzheimers Dis. 2017;55:813–22.
Janelidze S, Zetterberg H, Mattsson N, Palmqvist S, Vanderstichele H, Lindberg O, et al. CSF Abeta42/Abeta40 and Abeta42/Abeta38 ratios: better diagnostic markers of Alzheimer disease. Ann Clin Transl Neurol. 2016;3:154–65.
Lozoya-Delgado P, Ruiz-Sanchez de Leon JM, Pedrero-Perez EJ. [Validation of a cognitive complaints questionnaire for young adults: the relation between subjective memory complaints, prefrontal symptoms and perceived stress]. Rev Neurol. 2012;54:137–50.
Alegret M, Espinosa A, Valero S, Vinyes-Junque G, Ruiz A, Hernandez I, et al. Cut-off scores of a brief neuropsychological battery (NBACE) for Spanish individual adults older than 44 years old. PLoS ONE. 2013;8:e76436.
Villemagne VL, Ong K, Mulligan RS, Holl G, Pejoska S, Jones G, et al. Amyloid imaging with (18)F-florbetaben in Alzheimer disease and other dementias. J Nucl Med. 2011;52:1210–7.
Sanabria A, Alegret M, Rodriguez-Gomez O, Valero S, Sotolongo-Grau O, Monte-Rubio G, et al. The Spanish version of Face-Name Associative Memory Exam (S-FNAME) performance is related to amyloid burden in subjective cognitive decline. Sci Rep. 2018;8:3828.
Folstein MF, Folstein SE, McHugh PR. "Mini-mental state". A practical method for grading the cognitive state of patients for the clinician. J Psychiatr Res. 1975;12:189–98.
Kent P. The evolution of the Wechsler Memory Scale: a selective review. Appl Neuropsychol Adult. 2013;20:277–91.
Franzmeier N, Caballero MÁA, Taylor ANW, Simon-Vermot L, Buerger K, Ertl-Wagner B et al. Resting-state global functional connectivity as a biomarker of cognitive reserve in mild cognitive impairment. Brain Imaging Behav 2016;11:368–82.
Franzmeier N, Duering M, Weiner M, Dichgans M, Ewers M, Alzheimer's Disease Neuroimaging I. Left frontal cortex connectivity underlies cognitive reserve in prodromal Alzheimer disease. Neurology. 2017;88:1054–61.
Hariri AR, Goldberg TE, Mattay VS, Kolachana BS, Callicott JH, Egan MF, et al. Brain-derived neurotrophic factor val66met polymorphism affects human memory-related hippocampal activity and predicts memory performance. J Neurosci. 2003;23:6690–4.
Gordon BA, Blazey TM, Su Y, Hari-Raj A, Dincer A, Flores S, et al. Spatial patterns of neuroimaging biomarker change in individuals from families with autosomal dominant Alzheimer's disease: a longitudinal study. Lancet Neurol. 2018;17:241–50.
Araque Caballero MA, Brendel M, Delker A, Ren J, Rominger A, Bartenstein P, et al. Mapping 3-year changes in gray matter and metabolism in Abeta-positive nondemented subjects. Neurobiol Aging. 2015;36:2913–24.
Pasquini L, Scherr M, Tahmasian M, Meng C, Myers NE, Ortner M et al. Link between hippocampus' raised local and eased global intrinsic connectivity in AD. Alzheimers Dement 2014;11:475–84.
Ebeling U, von Cramon D. Topography of the uncinate fascicle and adjacent temporal fiber tracts. Acta Neurochir (Wien). 1992;115:143–8.
Von Der Heide RJ, Skipper LM, Klobusicky E, Olson IR. Dissecting the uncinate fasciculus: disorders, controversies and a hypothesis. Brain. 2013;136(Pt 6):1692–707.
Anderson KL, Rajagovindan R, Ghacibeh GA, Meador KJ, Ding M. Theta oscillations mediate interaction between prefrontal cortex and medial temporal lobe in human memory. Cereb Cortex. 2010;20:1604–12.
Spaniol J, Davidson PS, Kim AS, Han H, Moscovitch M, Grady CL. Event-related fMRI studies of episodic encoding and retrieval: meta-analyses using activation likelihood estimation. Neuropsychologia. 2009;47:1765–79.
Grady CL, McIntosh AR, Craik FI. Age-related differences in the functional connectivity of the hippocampus during memory encoding. Hippocampus. 2003;13:572–86.
Axmacher N, Schmitz DP, Wagner T, Elger CE, Fell J. Interactions between medial temporal lobe, prefrontal cortex, and inferior temporal regions during visual working memory: a combined intracranial EEG and functional magnetic resonance imaging study. J Neurosci. 2008;28:7304–12.
Harris AZ, Gordon JA. Long-range neural synchrony in behavior. Annu Rev Neurosci. 2015;38:171–94.
Wang L, Zang Y, He Y, Liang M, Zhang X, Tian L, et al. Changes in hippocampal connectivity in the early stages of Alzheimer's disease: evidence from resting state fMRI. Neuroimage. 2006;31:496–504.
Zhang Y, Simon-Vermot L, Araque Caballero MA, Gesierich B, Taylor AN, Duering M, et al. Enhanced resting-state functional connectivity between core memory-task activation peaks is associated with memory impairment in MCI. Neurobiol Aging. 2016;45:43–49.
Henseler I, Falkai P, Gruber O. Disturbed functional connectivity within brain networks subserving domain-specific subcomponents of working memory in schizophrenia: relation to performance and clinical symptoms. J Psychiatr Res. 2010;44:364–72.
Meyer-Lindenberg AS, Olsen RK, Kohn PD, Brown T, Egan MF, Weinberger DR, et al. Regionally specific disturbance of dorsolateral prefrontal-hippocampal functional connectivity in schizophrenia. Arch Gen Psychiatry. 2005;62:379–86.
Alvarez-Salvado E, Pallares V, Moreno A, Canals S. Functional MRI of long-term potentiation: imaging network plasticity. Philos Trans R Soc Lond B Biol Sci. 2014;369:20130152.
Boots EA, Schultz SA, Clark LR, Racine AM, Darst BF, Koscik RL, et al. BDNF Val66Met predicts cognitive decline in the Wisconsin Registry for Alzheimer's Prevention. Neurology. 2017;88:2098–106.
Peng S, Garzon DJ, Marchese M, Klein W, Ginsberg SD, Francis BM, et al. Decreased brain-derived neurotrophic factor depends on amyloid aggregation state in transgenic mouse models of Alzheimer's disease. J Neurosci. 2009;29:9321–9.
Li G, Peskind ER, Millard SP, Chi P, Sokal I, Yu CE, et al. Cerebrospinal fluid concentration of brain-derived neurotrophic factor and cognitive function in non-demented subjects. PLoS ONE. 2009;4:e5424.
Lim YY, Rainey-Smith S, Lim Y, Laws SM, Gupta V, Porter T, et al. BDNF Val66Met in preclinical Alzheimer's disease is associated with short-term changes in episodic memory and hippocampal volume but not serum mBDNF. Int Psychogeriatr. 2017;29:1825–34.
Krasnow B, Tamm L, Greicius MD, Yang TT, Glover GH, Reiss AL, et al. Comparison of fMRI activation at 3 and 1.5T during perceptual, cognitive, and affective processing. Neuroimage. 2003;18:813–26.
Day GS, Musiek ES, Roe CM, Norton J, Goate AM, Cruchaga C, et al. Phenotypic similarities between late-onset autosomal dominant and sporadic alzheimer disease: a single-family case-control study. JAMA Neurol. 2016;73:1125–32.
Chhatwal JP, Schultz AP, Johnson KA, Hedden T, Jaimes S, Benzinger TLS, et al. Preferential degradation of cognitive networks differentiates Alzheimer's disease from ageing. Brain. 2018;141:1486–500.
Bateman RJ, Aisen PS, De Strooper B, Fox NC, Lemere CA, Ringman JM, et al. Autosomal-dominant Alzheimer's disease: a review and proposal for the prevention of Alzheimer's disease. Alzheimers Res Ther. 2011;3:1.
Joshi A, Ringman JM, Lee AS, Juarez KO, Mendez MF. Comparison of clinical characteristics between familial and non-familial early onset Alzheimer's disease. J Neurol. 2012;259:2182–8.
Villemagne VL, Ataka S, Mizuno T, Brooks WS, Wada Y, Kondo M, et al. High striatal amyloid beta-peptide deposition across different autosomal Alzheimer disease mutation types. Arch Neurol. 2009;66:1537–44.
Crook R, Verkkoniemi A, Perez-Tur J, Mehta N, Baker M, Houlden H, et al. A variant of Alzheimer's disease with spastic paraparesis and unusual plaques due to deletion of exon 9 of presenilin 1. Nat Med. 1998;4:452–5.
Larner AJ, Doran M. Clinical phenotypic heterogeneity of Alzheimer's disease associated with mutations of the presenilin-1 gene. J Neurol. 2006;253:139–58.
Braak H, Alafuzoff I, Arzberger T, Kretzschmar H, Del Tredici K. Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta Neuropathol. 2006;112:389–404.
Schultz AP, Chhatwal JP, Hedden T, Mormino EC, Hanseeuw BJ, Sepulcre J, et al. Phases of hyperconnectivity and hypoconnectivity in the default mode and salience networks track with amyloid and tau in clinically normal individuals. J Neurosci. 2017;37:4323–31.
Marquie M, Normandin MD, Vanderburg CR, Costantino IM, Bien EA, Rycyna LG, et al. Validating novel tau positron emission tomography tracer [F-18]-AV-1451 (T807) on postmortem brain tissue. Ann Neurol. 2015;78:787–800.
Mackay CP, Kuys SS, Brauer SG. The effect of aerobic exercise on brain-derived neurotrophic factor in people with neurological disorders: a systematic review and meta-analysis. Neural Plast. 2017;2017:4716197.
Vaynman S, Ying Z, Gomez-Pinilla F. Hippocampal BDNF mediates the efficacy of exercise on synaptic plasticity and cognition. Eur J Neurosci. 2004;20:2580–90.
Zuccato C, Cattaneo E. Brain-derived neurotrophic factor in neurodegenerative diseases. Nat Rev Neurol. 2009;5:311–22.
Funding
This project was supported by The Dominantly Inherited Alzheimer’s Network (DIAN, UF1 AG032438) funded by the National Institute on Aging (NIA), the German Center for Neurodegenerative Diseases (DZNE), the NIHR Queen Square Dementia Biomedical Research Centre and the MRC Dementias Platform UK (MR/L023784/1 and MR/009076/1), and AMED under grant number JP17dk0207036 and JP17kk0205009. This manuscript has been reviewed by DIAN Study investigators for scientific content and consistency of data interpretation with previous DIAN Study publications. We acknowledge the altruism of the participants and their families and contributions of the DIAN research and support staff at each of the participating sites for their contributions to this study. The FACEHBI study is supported by Grifols®, Piramal®, Araclon Biotech®, Laboratorios Echevarne S.A. and Fundació ACE, Institut Català de Neurociències Aplicades. M.E.—Alzheimer Forschung Initiative & LMU excellent; J.C.—K23AG049087; B.A.G.—K01AG053474, Barnes-Jewish Hospital Foundation Willman Scholar Fund; Y.Y.L.—National Health and Medical Research Council (NHMRC) GNT1111603, GNT1147465.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
A.M.F. has received research funding from Biogen, Fujirebio, and Roche Diagnostics. She is a member of the scientific advisory boards for Roche, Genentech, and AbbVie and also consults for Araclon/Griffols and DiamiR.: Y.Y.L. has served as a scientific consultant to Biogen and Lundbeck; M.B. who has consulted or advisory board for Araclon, Grifols, Lilly, Nutricia, Roche and Servier. She received fees for lectures and funds for research from Araclon, Grifols, Nutricia, Roche and Servier. She has not received personal compensations from these organizations. A. Ruiz has consulted for Grifols and Landsteiner Genmed. He received funds for research and/or reimbursement of expenses for congresses attendance from Araclon, and Grifols. He has not received personal compensations from these organizations: T.B., Investigator, initiated research funding sponsored by Avid Radiopharmaceuticals (a wholly owned subsidiary of Eli Lilly) and Foundation for the NIH. Clinical trials sponsored by Avid Radiopharmaceuticals, Eli Lilly, Roche, Jaansen, Biogen, and NIH. Travel sponsored by the American Society for Neuroradiology, Alzheimer’s Association International Convention, NIH. The remaining authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Franzmeier, N., Ren, J., Damm, A. et al. The BDNFVal66Met SNP modulates the association between beta-amyloid and hippocampal disconnection in Alzheimer’s disease. Mol Psychiatry 26, 614–628 (2021). https://doi.org/10.1038/s41380-019-0404-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-019-0404-6
This article is cited by
-
Subject classification and cross-time prediction based on functional connectivity and white matter microstructure features in a rat model of Alzheimer’s using machine learning
Alzheimer's Research & Therapy (2023)
-
Brain-Derived Neurotrophic Factor rs6265 polymorphism is associated with severe cancer-related fatigue and neuropathic pain in female cancer survivors
Journal of Cancer Survivorship (2023)
-
Correlation between nocturnal intermittent hypoxemia and mild cognitive impairment in the older adult and the role of BDNF Val66Met polymorphism: a hospital-based cross-sectional study
Sleep and Breathing (2023)
-
Brain-derived neurotrophic factor in Alzheimer’s disease and its pharmaceutical potential
Translational Neurodegeneration (2022)
-
A review of brain imaging biomarker genomics in Alzheimer’s disease: implementation and perspectives
Translational Neurodegeneration (2022)