Abstract
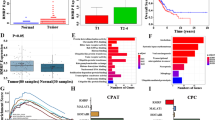
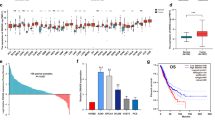
Long non-coding RNAs (lncRNAs) are reported to play key roles in tumorigenesis. However, the mechanisms underlying lncRNA-mediated regulation of RNA-binding protein phase separation in tumorigenesis have not been completely elucidated. In this study, an oncogenic lncRNA MELTF-AS1 was identified using systematic data analysis, screening, and verification. MELTF-AS1 was markedly upregulated in non-small cell lung cancer (NSCLC). High MELTF-AS1 levels were associated with advanced tumor-node-metastasis stage (TNM), high tumor size, and decreased survival time. Functionally, MELTF-AS1 regulated cell proliferation and metastasis in vitro and in vivo. RNA sequencing analysis revealed that MELTF-AS1 knockdown specifically modulated genes associated with cell proliferation, apoptosis, and migration. Mechanistically, at the genome level, copy number amplification promoted MELTF-AS1 expression. At the transcriptional level, the transcription factor SP1 directly activated MELTF-AS1 transcription by binding to its promoter. Furthermore, MELTF-AS1 could directly bind and drive the phase separation of YBX1, which was an RNA-binding protein and involved in tumorigenesis, thus activating ANXA8 transcription and promoting tumorigenesis of NSCLC. Aberrant activation of ANXA8 and promotion of tumorigenesis have been found in a variety of tumors. These novel findings demonstrated the critical role of MELTF-AS1-driven phase separation-mediated transcriptional regulation and provided a potential novel diagnostic and therapeutic target for NSCLC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2021. CA Cancer J Clin. 2021;71:7–33.
Petrelli NJ, Winer EP, Brahmer J, Dubey S, Smith S, Thomas C, et al. Clinical Cancer Advances 2009: major research advances in cancer treatment, prevention, and screening−a report from the American Society of Clinical Oncology. J Clin Oncol. 2009;27:6052–69.
Camps C, del Pozo N, Blasco A, Blasco P, Sirera R. Importance of quality of life in patients with non-small-cell lung cancer. Clin Lung Cancer. 2009;10:83–90.
Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12:861–74.
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, et al. Landscape of transcription in human cells. Nature. 2012;489:101–8.
Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Disco. 2011;1:391–407.
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10:155–9.
Gutschner T, Hammerle M, Eissmann M, Hsu J, Kim Y, Hung G, et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013;73:1180–9.
Chen Z, Li JL, Lin S, Cao C, Gimbrone NT, Yang R, et al. cAMP/CREB-regulated LINC00473 marks LKB1-inactivated lung cancer and mediates tumor growth. J Clin Invest. 2016;126:2267–79.
Zhang E, Han L, Yin D, He X, Hong L, Si X, et al. H3K27 acetylation activated-long non-coding RNA CCAT1 affects cell proliferation and migration by regulating SPRY4 and HOXB13 expression in esophageal squamous cell carcinoma. Nucleic Acids Res. 2017;45:3086–101.
Yin D, Lu X, Su J, He X, De W, Yang J, et al. Long noncoding RNA AFAP1-AS1 predicts a poor prognosis and regulates non-small cell lung cancer cell proliferation by epigenetically repressing p21 expression. Mol Cancer. 2018;17:92.
Zhang E, He X, Zhang C, Su J, Lu X, Si X, et al. A novel long noncoding RNA HOXC-AS3 mediates tumorigenesis of gastric cancer by binding to YBX1. Genome Biol. 2018;19:154.
Liu X, Lu X, Zhen F, Jin S, Yu T, Zhu Q, et al. LINC00665 induces acquired resistance to gefitinib through recruiting EZH2 and activating PI3K/AKT pathway in NSCLC. Mol Ther Nucleic Acids. 2019;16:155–61.
Sabari BR, Dall'Agnese A, Boija A, Klein IA, Coffey EL, Shrinivas K. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science. 2018;361: eaar3958.
Banani SF, Lee HO, Hyman AA, Rosen MK. Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol. 2017;18:285–98.
Jiang S, Fagman JB, Chen C, Alberti S, Liu B. Protein phase separation and its role in tumorigenesis. Elife. 2020;9: e60264.
Alberti S, Dormann D. Liquid-liquid phase separation in disease. Annu Rev Genet. 2019;53:171–94.
Lin C, Yang L. Long noncoding RNA in cancer: wiring signaling circuitry. Trends Cell Biol. 2018;28:287–301.
Schimke RT. The search for early genetic events in tumorigenesis: an amplification paradigm. Cancer Cells. 1990;2:149–51.
Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J, et al. The landscape of somatic copy-number alteration across human cancers. Nature. 2010;463:899–905.
Lin M, Smith LT, Smiraglia DJ, Kazhiyur-Mannar R, Lang JC, Schuller DE, et al. DNA copy number gains in head and neck squamous cell carcinoma. Oncogene. 2006;25:1424–33.
La-Touche S, Lemetre C, Lambros M, Stankiewicz E, Ng CK, Weigelt B, et al. DNA copy number aberrations, and human papillomavirus status in penile carcinoma. Clinico-pathological correlations and potential driver genes. PLoS ONE. 2016;11:e0146740.
Ricciotti RW, Baraff AJ, Jour G, Kyriss M, Wu Y, Liu Y, et al. High amplification levels of MDM2 and CDK4 correlate with poor outcome in patients with dedifferentiated liposarcoma: a cytogenomic microarray analysis of 47 cases. Cancer Genet. 2017;218-219:69–80.
Li L, Davie JR. The role of Sp1 and Sp3 in normal and cancer cell biology. Ann Anat. 2010;192:275–83.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98–W102.
Yu XX, Murray SF, Pandey SK, Booten SL, Bao D, Song XZ, et al. Antisense oligonucleotide reduction of DGAT2 expression improves hepatic steatosis and hyperlipidemia in obese mice. Hepatology. 2005;42:362–71.
Yang F, Wei J, Zhang S, Zhang X. Shrimp miR-S8 suppresses the stemness of human melanoma stem-like cells by targeting the transcription factor YB-1. Cancer Res. 2017;77:5543–53.
Kosnopfel C, Sinnberg T, Schittek B. Y-box binding protein 1−a prognostic marker and target in tumour therapy. Eur J Cell Biol. 2014;93:61–70.
Goodarzi H, Liu X, Nguyen HC, Zhang S, Fish L, Tavazoie SF. Endogenous tRNA-derived fragments suppress breast cancer progression via YBX1 displacement. Cell. 2015;161:790–802.
Shibahara K, Sugio K, Osaki T, Uchiumi T, Maehara Y, Kohno K, et al. Nuclear expression of the Y-box binding protein, YB-1, as a novel marker of disease progression in non-small cell lung cancer. Clin Cancer Res. 2001;7:3151–5.
Evdokimova V, Tognon C, Ng T, Ruzanov P, Melnyk N, Fink D, et al. Translational activation of snail1 and other developmentally regulated transcription factors by YB-1 promotes an epithelial-mesenchymal transition. Cancer Cell. 2009;15:402–15.
Li D, Liu X, Zhou J, Hu J, Zhang D, Liu J, et al. Long noncoding RNA HULC modulates the phosphorylation of YB-1 through serving as a scaffold of extracellular signal-regulated kinase and YB-1 to enhance hepatocarcinogenesis. Hepatology. 2017;65:1612–27.
Deng SJ, Chen HY, Ye Z, Deng SC, Zhu S, Zeng Z, et al. Hypoxia-induced LncRNA-BX111 promotes metastasis and progression of pancreatic cancer through regulating ZEB1 transcription. Oncogene. 2018;37:5811–28.
Imada K, Shiota M, Kohashi K, Kuroiwa K, Song Y, Sugimoto M, et al. Mutual regulation between Raf/MEK/ERK signaling and Y-box-binding protein-1 promotes prostate cancer progression. Clin Cancer Res. 2013;19:4638–50.
Roden C, Gladfelter AS. RNA contributions to the form and function of biomolecular condensates. Nat Rev Mol Cell Biol. 2021;22:183–95.
Wu M, Xu G, Han C, Luan PF, Xing YH, Nan F, et al. lncRNA SLERT controls phase separation of FC/DFCs to facilitate Pol I transcription. Science. 2021;373:547–55.
Ulitsky I, Bartel DP. lincRNAs: genomics, evolution, and mechanisms. Cell. 2013;154:26–46.
Stein T, Price KN, Morris JS, Heath VJ, Ferrier RK, Bell AK, et al. Annexin A8 is up-regulated during mouse mammary gland involution and predicts poor survival in breast cancer. Clin Cancer Res. 2005;11:6872–9.
Hata H, Tatemichi M, Nakadate T. Involvement of annexin A8 in the properties of pancreatic cancer. Mol Carcinog. 2014;53:181–91.
Sarkar A, Yang P, Fan YH, Mu ZM, Hauptmann R, Adolf GR, et al. Regulation of the expression of annexin VIII in acute promyelocytic leukemia. Blood. 1994;84:279–86.
Li C, Tan F, Pei Q, Zhou Z, Zhou Y, Zhang L, et al. Non-coding RNA MFI2-AS1 promotes colorectal cancer cell proliferation, migration and invasion through miR-574-5p/MYCBP axis. Cell Prolif. 2019;52:e12632.
Flippot R, Mouawad R, Spano JP, Roupret M, Comperat E, Bitker MO, et al. Expression of long non-coding RNA MFI2-AS1 is a strong predictor of recurrence in sporadic localized clear-cell renal cell carcinoma. Sci Rep. 2017;7:8540.
Wei Y, Wang Z, Zong Y, Deng D, Chen P, Lu J. LncRNA MFI2-AS1 promotes HCC progression and metastasis by acting as a competing endogenous RNA of miR-134 to upregulate FOXM1 expression. Biomed Pharmacother. 2020;125:109890.
Liu K, Liu J, Bo QF. MFI2-AS1 regulates the aggressive phenotypes in glioma by modulating MMP14 via a positive feedback loop. Eur Rev Med Pharm Sci. 2019;23:5884–95.
Tseng YY, Moriarity BS, Gong W, Akiyama R, Tiwari A, Kawakami H, et al. PVT1 dependence in cancer with MYC copy-number increase. Nature. 2014;512:82–86.
Wierstra I. Sp1: emerging roles−beyond constitutive activation of TATA-less housekeeping genes. Biochem Biophys Res Commun. 2008;372:1–13.
Jungert K, Buck A, von Wichert G, Adler G, Konig A, Buchholz M, et al. Sp1 is required for transforming growth factor-beta-induced mesenchymal transition and migration in pancreatic cancer cells. Cancer Res. 2007;67:1563–70.
Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152:1298–307.
Liu Y, Zhao J, Zhang W, Gan J, Hu C, Huang G, et al. lncRNA GAS5 enhances G1 cell cycle arrest via binding to YBX1 to regulate p21 expression in stomach cancer. Sci Rep. 2015;5:10159.
Boija A, Klein IA, Young RA. Biomolecular condensates and cancer. Cancer Cell. 2021;39:174–92.
Boija A, Klein IA, Sabari BR, Dall’Agnese A, Coffey EL, Zamudio AV, et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell. 2018;175:1842–55.
Munschauer M, Nguyen CT, Sirokman K, Hartigan CR, Hogstrom L, Engreitz JM, et al. The NORAD lncRNA assembles a topoisomerase complex critical for genome stability. Nature. 2018;561:132–6.
Yan X, Yin J, Yao H, Mao N, Yang Y, Pan L. Increased expression of annexin A3 is a mechanism of platinum resistance in ovarian cancer. Cancer Res. 2010;70:1616–24.
Song J, Shih IeM, Salani R, Chan DW, Zhang Z. Annexin XI is associated with cisplatin resistance and related to tumor recurrence in ovarian cancer patients. Clin Cancer Res. 2007;13:6842–9.
Li J, Han L, Roebuck P, Diao L, Liu L, Yuan Y, et al. TANRIC: an interactive open platform to explore the function of lncRNAs in cancer. Cancer Res. 2015;75:3728–37.
Acknowledgements
The authors thank NovelBioinformatics Ltd., Co. for the support of bioinformatics analysis with their NovelBrain Cloud Analysis Platform (www.novelbrain.com).
Funding
This work was supported by the National Natural Science Foundation of China [82172992, 81972188 and 82103133], Natural Science Foundation of Jiangsu Province [BK20211253 and BK20210973], China Postdoctoral Science Foundation (2020M671395), the Medical Important Talents [ZDRCA2016024], the Program for Innovative Research Team of Nanjing Medical University [JX102GSP201727], and the Wu Jie-ping Foundation [320.6799.15032].
Author information
Authors and Affiliations
Contributions
EZ and RG conceived the project and designed the experiments. XL, JW and CL started and performed majority of the experiments. WW, TQ and XL collected the NSCLC samples. XH provided technical and administrative support. XL analyzed results and wrote the manuscript. RG and EZ contributed to manuscript revision and supervised all experiments. All authors read and provided suggestions during manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Lu, X., Wang, J., Wang, W. et al. Copy number amplification and SP1-activated lncRNA MELTF-AS1 regulates tumorigenesis by driving phase separation of YBX1 to activate ANXA8 in non-small cell lung cancer. Oncogene 41, 3222–3238 (2022). https://doi.org/10.1038/s41388-022-02292-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02292-z
This article is cited by
-
The m6A modification mediated-lncRNA POU6F2-AS1 reprograms fatty acid metabolism and facilitates the growth of colorectal cancer via upregulation of FASN
Molecular Cancer (2024)
-
High-throughput and proteome-wide discovery of endogenous biomolecular condensates
Nature Chemistry (2024)
-
Downregulation of UBB potentiates SP1/VEGFA-dependent angiogenesis in clear cell renal cell carcinoma
Oncogene (2024)
-
Omics-based molecular classifications empowering in precision oncology
Cellular Oncology (2024)
-
Gene amplification-driven lncRNA SNHG6 promotes tumorigenesis via epigenetically suppressing p27 expression and regulating cell cycle in non–small cell lung cancer
Cell Death Discovery (2022)