Abstract
False-negative cell-free DNA (cfDNA) screening results involving Down syndrome are rare, but have high clinical impact on patients and their healthcare providers. Understanding the biology behind these results may allow for improved diagnostic follow-up and counseling. In 5 different centers offering cfDNA prenatal screening, 9 false-negative results were documented in 646 confirmed cases of trisomy 21; a false-negative rate of 1.4% (95% CI, 0.7–2.6). False-negative results included 4 cases of classical trisomy 21 and 5 cases with a de novo 21q;21q rearrangement. Two out of five rearrangements had molecular studies and were confirmed as isochromosomes. When combined with reports from the cfDNA screening literature, 8 out of 29 (28%) Down syndrome cases with a false-negative “non-invasive prenatal test” (NIPT) were associated with a 21q;21q rearrangement, compared with 2% reported in live born children with Down syndrome. In our laboratory series, evidence for placental or fetal mosaicism was present in 3 out of 3 true-positive cases involving a 21q;21q rearrangement and was confirmed in one false-negative case where placental material was available for study. Isochromosome 21q rearrangements are thus overrepresented among false-negative cfDNA screening results involving Down syndrome. Postzygotic isochromosome formation leading to placental mosaicism provides a biological cause for the increased prevalence of these rearrangements among false-negative cases. For clinical practice, a low trisomic fraction (z-score or equivalent measure) relative to the fetal fraction suggests placental mosaicism. Care should be taken as these cases may not reflect confined placental mosaicism, but rather full trisomy in the presence of a placenta containing normal cells.
Similar content being viewed by others
Introduction
Cell-free DNA (cfDNA)-based prenatal screening, also known as non-invasive prenatal testing (NIPT), has unparalleled sensitivity and specificity for trisomy 21, when compared to other screening modalities [1, 2]. Detection rates typically exceed 99%, with a false-positive (FP) rate of <0.1% [3]. Rare false-negative (FN) cases have been documented in larger clinical series [4, 5]. Low fetal fraction (FF) and mosaicism involving a euploid cell line have been implicated in some FN results, while other cases remain unexplained [6]. Here, we describe an overrepresentation of chromosome 21q;21q rearrangements among FN cfDNA screening reports and provide evidence that placental mosaicism arising from postzygotic isochromosome 21q formation is the likely biological cause. A low trisomic fraction [7] relative to the FF may suggest confined placental mosaicism (CPM), or, as described in this report, a full trisomy present in the fetus with a placenta containing normal cells.
Materials and methods
Five clinical laboratories (Academic Medical Center Amsterdam, the Netherlands; Victorian Clinical Genetics Services (VCGS), Melbourne, Australia; Genea, Sydney, Australia; Aarhus University Hospital, Denmark and Fleury Medicina & Saude, Sao Paulo, Brazil), reviewed their cfDNA screening outcome records for trisomy 21 true-positive (TP) and FN cases. All prevailing cfDNA screening technologies were utilized: massive parallel shotgun sequencing (MPSS), targeted MPS (tMPS), targeted microarray (tMA), and targeted single nucleotide polymorphism (tSNP)-based methodologies, using a number of bioinformatics algorithms. Clinical samples obtained from consented patients with average or elevated risk and received between 29 July 2013 and 29 September 2017 were included. Laboratory databases were interrogated for cytogenetic summary reports for all trisomy 21 cases, and for chromosome 21 z-scores [8] (or equivalent measure such as NCV_score [7]) and FF estimates [2, 9,10,11] in FN cases. One laboratory also reviewed its cytogenetic records for all trisomy 21 cases where cfDNA screening was done by external commercial providers. Genetic tests were performed on chorionic villi, amniotic fluid, newborn blood and/or placental material when available. The false-negative rate (FNR) was calculated from confirmed FN and TP cases involving trisomy 21 [(FN)/(TP+FN)]. CfDNA screening technologies and methods used to estimate FF for cases with 21q;21q rearrangements are listed in Table 1. CfDNA screening technologies associated with FN results for all known Down syndrome cases with karyotype data, including those described in literature, are listed in Table 2. Placental mosaicism for trisomy 21 was suspected in positive cfDNA samples that demonstrated a low trisomic fraction [7], relative to the FF, as described previously [11]. The trisomic fraction can be calculated directly from sequence tag-count data [7] and is the relative increase in sequence counts above the expected euploid state seen with massive parallel shotgun sequencing (MPSS) based NIPT methods [11].
For the identification of cytogenetically documented FN cfDNA screening cases, we relied on a systematic review published in July 2017 [3]. We included all publications used in this meta-analysis and listed in supplementary files, except for proof-of-concept studies, which we excluded. Additional records were found by cross-referencing.
Results
Five participating laboratories tested 58,504 cfDNA clinical samples and recorded 6 FN results from 500 confirmed cases of trisomy 21. Another 3 FN cases were documented in 146 diagnostic samples with trisomy 21 karyotype where cfDNA screening was done by external commercial providers. In total, 9 FN cases were documented from 646 confirmed cases of trisomy 21. The FNR for the combined series was 1.4% (95% CI, 0.7–2.6). FN results were recorded using targeted sequencing, whole genome sequencing, array-based and SNP-based techniques.
Five out of 9 FN samples involved a chromosome 21q;21q rearrangement; the other 4 cases had classical trisomy 21 karyotypes. All 5 cases with rearrangements resulted in Down syndrome live births (Table 1). In cases 1 and 3 the rearrangement was an isochromosome 21q based on the results of quantitative fluorescence PCR (QF-PCR) analysis. In cases 2, 4, and 5 no molecular studies were done. All 5 rearrangements were de novo based on normal parental karyotypes. The average fetal fraction at first analysis in FN cases with 21q;21q rearrangements was 10.1% (range 3–17.2%).
In case 1, ultrasound findings after low risk cfDNA screening led to amniocentesis and a non-mosaic isochromosome 21q was found. After birth, mosaicism involving a disomic cell-line was seen in chorionic villi from the placenta [12]. In cases 2–5, the newborn had a non-mosaic karyotype. The placenta was not available for analysis to confirm or exclude placental mosaicism in these cases (Table 1).
Three TP cases of trisomy 21 involving 21q;21q rearrangements (cases 6–8) were recorded in the laboratory series (Table 1). In case 6, true fetal mosaicism for trisomy 21 was reported after amniocentesis and a low-grade mosaicism involving an isochromosome 21q was confirmed in newborn blood. The placenta was not available. The two remaining TP cases with 21q;21q rearrangements demonstrated placental mosaicism based on cytogenetic analysis of chorionic villi (case 7) and/or bioinformatics analysis of cfDNA (cases 7 and 8) (Table 1 and Fig. 1).
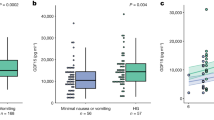
Singleton pregnancies (n = 132) at increased risk for trisomy 21 after cfDNA screening and their cytogenetic outcomes. For each sample, the x-axis indicates the fetal fraction (FF) and the y-axis shows the trisomic fraction. Non-mosaic true positive samples with trisomy 21 (black circles) lie close to the unity line (y = x). Samples that plot below the unity line have a trisomic fraction that is substantially lower than the FF. These samples were associated with true fetal mosaicism (TFM) for trisomy 21 (yellow circles) or mosaicism confined to the placenta (CPM) (blue circles). Trisomy 21 samples with TFM and CPM may occasionally plot on the unity line. Three true positive trisomy 21 samples with 21q;21q rearrangements are represented by red circles. Two plot well below the unity line, indicating placental mosaicism, but were associated with non-mosaic trisomy 21 in the fetus (cases 7 and 8 in main text). The third case plots on the unity line and was associated with TFM for trisomy 21 after amniocentesis, with a low proportion of trisomic cells (3/85) confirmed in the newborn (case 6 in main text)
In total 77 papers were reviewed for FN reports; 70 from the published systematic review [3] and another 7 via cross-referencing [13,14,15,16,17,18,19]. In total 16 papers mentioned FN cfDNA screening results for Down syndrome and 7 specified the karyotype. After combining these data with the results from the laboratory series, 8 out of 29 FN results for Down syndrome (28%) involved a 21q;21q rearrangement (Table 2).
Discussion
Down syndrome caused by 21q;21q rearrangements occurs in ~2% of cases [20]. Some 21q;21q rearrangements are Robertsonian translocations between two different chromosome arms, but most represent true isochromosomes [21]. Isochromosomes have two identical chromosome arms, and arise de novo from a post-fertilization event involving centromere misdivision or U-type exchange between sister chromatids [21, 22]. Depending on the timing of this event, mosaicism for the initial euploid cell line may persist in the placenta. Chorionic villus sampling (CVS) literature describes FN cases of trisomy 21 associated with de novo isochromosome 21q. The chromosome aberration is present in the fetus and usually the mesenchymal core of the chorionic villi (long-term CVS culture), but is absent from the villi’s outer cytotrophoblast layer (short-term CVS culture) [23,24,25,26]. Placental cytotrophoblast cells are more distantly related to the embryo than are cells from the mesenchymal core, which derive from the inner cell mass, along with the embryo [27]. If de novo isochromosome 21q formation occurs in inner cell mass precursors, then the mesenchymal core and fetus would have trisomy 21, while the cytotrophoblast cells would remain predominantly euploid. As the cytotrophoblast is the primary source of the “fetal” cfDNA obtained from maternal plasma [28], isochromosome 21q might also be seen among FN cfDNA screening results, regardless of the cfDNA technology used.
Our data show that 8 of 29 (28%) FN results with karyotype information involved 21q;21q rearrangements; a 14-fold increase over the 2% of cases reported in live born children with Down syndrome. In 21q;21q rearrangement cases, placental mosaicism can lead to a cfDNA “trisomic” fraction that is reduced or absent relative to the FF (Fig. 1). The likelihood of a FN result will be influenced by variables such as the proportion of abnormal cells in the placental cytotrophoblast, the FF, and the assay’s limit of detection. Furthermore, a low trisomic fraction relative to FF may be seen in cases of true fetal mosaicism for trisomy 21 and in cases of CPM with normal fetal karyotype (Fig. 1). Only an invasive diagnostic test can distinguish these mosaic outcomes, with amniocentesis providing the most definitive prenatal result. It should be noted that FFs estimated by different methods are not directly comparable [29] and each cfDNA-based screening assay will need to establish its own method for assessing possible mosaicism.
In case 1, repeated cfDNA testing showed an increase in chromosome 21 z-scores as the FF increased with gestational age and placental mass. The pregnancy was eventually called at high risk at 32 weeks of gestation; the FF was 14.3% (Table 1). Placental mosaicism was proven after birth with disomic cells being confined to the cytotrophoblast. For TP cases 7 and 8, the trisomic fraction was calculated [7, 11] and was substantially lower than the estimated FF, consistent with placental mosaicism involving cytotrophoblast (Table 1 and Fig. 1). The mosaicism was confirmed in case 7 but placental tissue was not available in case 8.
In case 1, ultrasound anomalies prompted amniocentesis and the diagnosis was made prenatally. In the 4 other cases either ultrasound was normal or “soft markers” or an enlarged nuchal translucency were observed and no invasive testing was done, consistent with current guidelines [30], and parental choice.
A limitation of our study design is the selection bias in favor of centers confronted with FN cases of Down syndrome involving 21q;21q rearrangements. We therefore sought to confirm the data by reviewing published cases of FN cfDNA screening results. Although only about half of the studies reporting FN results specified the trisomy 21 karyotype, our findings were confirmed, with 3 out of 20 (15%) FN results recording 21q;21q rearrangements. We acknowledge that reports of FN cases in the literature might also be influenced by publication bias. Thus, our total calculated prevalence of 28% for FN 21q;21q rearrangements is likely to be an upper estimate.
We conclude that Down syndrome due to de novo isochromosome 21q is more likely to record a FN result than standard trisomy 21 during cfDNA screening. This can be explained by placental mosaicism. For centers reporting cfDNA screening outcome data we recommend recording the karyotype of all discrepant results. For clinical practice, a low trisomic fraction relative to the FF may suggest placental mosaicism. Identification and interpretation of potential mosaicism helps guide further patient counseling and management, such as recommending amniocentesis rather than CVS for follow-up studies. Care should be taken as these cases may not reflect confined placental mosaicism, but rather full trisomy in the presence of a placenta containing normal cells. Finally, the results of our study will help clinicians and patients understand that the biology of cfDNA screening is complex. False-negative results can occur through mechanisms other than poor quality, technical error or negligence.
References
Bianchi DW, Parker RL, Wentworth J, et al. DNA sequencing versus standard prenatal aneuploidy screening. N Engl J Med. 2014;370:799–808.
Norton ME, Jacobsson B, Swamy GK, et al. Cell-free DNA analysis for noninvasive examination of trisomy. N Engl J Med. 2015;372:1589–97.
Gil MM, Accurti V, Santacruz B, Plana MN, Nicolaides KH. Analysis of cell-free DNA in maternal blood in screening for aneuploidies: updated meta-analysis. Ultrasound Obstet Gynecol. 2017;50:302–14.
Taneja PA, Snyder HL, de Feo E, et al. Noninvasive prenatal testing in the general obstetric population: clinical performance and counseling considerations in over 85000 cases. Prenat Diagn. 2016;36:237–43.
Zhang H, Gao Y, Jiang F, et al. Non-invasive prenatal testing for trisomies 21, 18 and 13: clinical experience from 146,958 pregnancies. Ultrasound Obstet Gynecol. 2015;45:530–8.
Hartwig TS, Ambye L, Sorensen S, Jorgensen FS. Discordant non-invasive prenatal testing (NIPT)–a systematic review. Prenat Diagn. 2017;37:527–39.
Rava RP, Srinivasan A, Sehnert AJ, Bianchi DW. Circulating fetal cell-free DNA fractions differ in autosomal aneuploidies and monosomy X. Clin Chem. 2014;60:243–50.
Straver R, Sistermans EA, Holstege H, Visser A, Oudejans CB, Reinders MJ. WISECONDOR: detection of fetal aberrations from shallow sequencing maternal plasma based on a within-sample comparison scheme. Nucleic Acids Res. 2014;42:e31.
Kim SK, Hannum G, Geis J, et al. Determination of fetal DNA fraction from the plasma of pregnant women using sequence read counts. Prenat Diagn. 2015;35:810–5.
Straver R, Oudejans CB, Sistermans EA, Reinders MJ. Calculating the fetal fraction for noninvasive prenatal testing based on genome-wide nucleosome profiles. Prenat Diagn. 2016;36:614–21.
Pertile MD, Halks-Miller M, Flowers N et al. Rare autosomal trisomies, revealed by maternal plasma DNA sequencing, suggest increased risk of feto-placental disease. Sci Transl Med. 2017; 30;9. pii: eaan1240. doi: 10.1126/scitranslmed.aan1240.
Oepkes D, Page-Christiaens GC, Bax CJ, et al. Trial by Dutch laboratories for evaluation of non-invasive prenatal testing. Part I-clinical impact. Prenat Diagn. 2016;36:1083–90.
Nicolaides KH, Syngelaki A, Ashoor G, Birdir C, Touzet G. Noninvasive prenatal testing for fetal trisomies in a routinely screened first-trimester population. Am J Obstet Gynecol. 2012;207:374. e371-6
Nicolaides KH, Syngelaki A, Gil M, Atanasova V, Markova D. Validation of targeted sequencing of single-nucleotide polymorphisms for non-invasive prenatal detection of aneuploidy of chromosomes 13, 18, 21, X, and Y. Prenat Diagn. 2013;33:575–9.
Willems PJ, Dierickx H, Segers N et al. High positive predictive value (PPV) of cell-free DNA (cfDNA) testing in a clinical study of 10,000 consecutive pregnancies. J Mol Biomark Diagn. 2016, 7: 285
Quezada MS, Gil MM, Francisco C, Orosz G, Nicolaides KH. Screening for trisomies 21, 18 and 13 by cell-free DNA analysis of maternal blood at 10-11 weeks’ gestation and the combined test at 11–13 weeks. Ultrasound Obstet Gynecol. 2015;45:36–41.
Song Y, Liu C, Qi H, Zhang Y, Bian X, Liu J. Noninvasive prenatal testing of fetal aneuploidies by massively parallel sequencing in a prospective Chinese population. Prenat Diagn. 2013;33:700–6.
Wang Y, Zhu J, Chen Y, et al. Two cases of placental T21 mosaicism: challenging the detection limits of non-invasive prenatal testing. Prenat Diagn. 2013;33:1207–10.
Zimmermann B, Hill M, Gemelos G, et al. Noninvasive prenatal aneuploidy testing of chromosomes 13, 18, 21, X, and Y, using targeted sequencing of polymorphic loci. Prenat Diagn. 2012;32:1233–41.
Zhao W, Chen F, Wu M, et al. Postnatal identification of trisomy 21: an overview of 7,133 postnatal trisomy 21 cases identified in a diagnostic reference laboratory in China. PLoS ONE. 2015;10:e0133151.
Shaffer LG, McCaskill C, Haller V, Brown JA, Jackson-Cook CK. Further characterization of 19 cases of rea(21q21q) and delineation as isochromosomes or Robertsonian translocations in Down syndrome. Am J Med Genet. 1993;47:1218–22.
Robinson WP, Bernasconi F, Basaran S, et al. A somatic origin of homologous Robertsonian translocations and isochromosomes. Am J Hum Genet. 1994;54:290–302.
Brisset S, Aboura A, Audibert F, et al. Discordant prenatal diagnosis of trisomy 21 due to mosaic structural rearrangements of chromosome 21. Prenat Diagn. 2003;23:461–9.
Gilardi JL, Perrotin F, Paillet C, et al. Prenatal diagnosis of trisomy 21 by i(21q): a rare case of fetoplacental chromosomal discrepancy. Prenat Diagn. 2002;22:856–8.
Riegel M, Wisser J, Baumer A, Schinzel A. Postzygotic isochromosome formation as a cause for false-negative results from chorionic villus chromosome examinations. Prenat Diagn. 2006;26:221–5.
Caloone J, Sanlaville D, Fichez A, Abel C, Huissoud C, Rudigoz RC. Trisomy 21 by isochromosome: a case report of true false negative of chorionic villi sampling. Gynecol Obstet Fertil. 2011;39:e77–80.
Bianchi DW, Wilkins-Haug LE, Enders AC, Hay ED. Origin of extraembryonic mesoderm in experimental animals: relevance to chorionic mosaicism in humans. Am J Med Genet. 1993;46:542–50.
Flori E, Doray B, Gautier E, et al. Circulating cell-free fetal DNA in maternal serum appears to originate from cyto- and syncytio-trophoblastic cells. Case report. Hum Reprod. 2004;19:723–4.
van Beek DM, Straver R, Weiss MM, et al. Comparing methods for fetal fraction determination and quality control of NIPT samples. Prenat Diagn. 2017;37:769–73.
Salomon LJ, Alfirevic Z, Audibert F, et al. ISUOG updated consensus statement on the impact of cfDNA aneuploidy testing on screening policies and prenatal ultrasound practice. Ultrasound Obstet Gynecol. 2017;49:815–6.
Futch T, Spinosa J, Bhatt S, de Feo E, Rava RP, Sehnert AJ. Initial clinical laboratory experience in noninvasive prenatal testing for fetal aneuploidy from maternal plasma DNA samples. Prenat Diagn. 2013;33:569–74.
Stumm M, Entezami M, Haug K, et al. Diagnostic accuracy of random massively parallel sequencing for non-invasive prenatal detection of common autosomal aneuploidies: a collaborative study in Europe. Prenat Diagn. 2014;34:185–91.
Acknowledgements
We thank Dr. D. van Beek, University Medical Center Utrecht, and Dr. M. C. van Maarle, Academic Medical Center Amsterdam, for their contributions to data for Case 1. Research conducted at the Murdoch Children’s Research Institute was supported by the Victorian Government’s Operational Infrastructure Support Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Dr. Lieve Page-Christiaens is Associate Medical Director at Illumina, Inc. San Diego and holds equity in Illumina. The remaining authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Huijsdens-van Amsterdam, K., Page-Christiaens, L., Flowers, N. et al. Isochromosome 21q is overrepresented among false-negative cell-free DNA prenatal screening results involving Down syndrome. Eur J Hum Genet 26, 1490–1496 (2018). https://doi.org/10.1038/s41431-018-0188-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-018-0188-1
This article is cited by
-
Two cases of placental trisomy 21 mosaicism causing false-negative NIPT results
Molecular Cytogenetics (2023)
-
Fission yeast Srr1 and Skb1 promote isochromosome formation at the centromere
Communications Biology (2023)
-
Performance of cell-free DNA sequencing-based non-invasive prenatal testing: experience on 36,456 singleton and multiple pregnancies
BMC Medical Genomics (2021)
-
A rare Down syndrome foetus with de novo 21q;21q rearrangements causing false negative results in non-invasive prenatal testing: a case report
BMC Medical Genomics (2020)
-
Genetic diagnosis in the fetus
Journal of Perinatology (2020)