Abstract
T cells dynamically interact with multiple, distinct cellular subsets to determine effector and memory differentiation. Here, we developed a platform to quantify cell location in three dimensions to determine the spatial requirements that direct T cell fate. After viral infection, we demonstrated that CD8+ effector T cell differentiation is associated with positioning at the lymph node periphery. This was instructed by CXCR3 signaling since, in its absence, T cells are confined to the lymph node center and alternatively differentiate into stem-like memory cell precursors. By mapping the cellular sources of CXCR3 ligands, we demonstrated that CXCL9 and CXCL10 are expressed by spatially distinct dendritic and stromal cell subsets. Unlike effector cells, retention of stem-like memory precursors in the paracortex is associated with CCR7 expression. Finally, we demonstrated that T cell location can be tuned, through deficiency in CXCL10 or type I interferon signaling, to promote effector or stem-like memory fates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Eickhoff, S. et al. Robust anti-viral immunity requires multiple distinct T cell-dendritic cell interactions. Cell 162, 1322–1337 (2015).
Groom, J. R. Regulators of T-cell fate: integration of cell migration, differentiation and function. Immunol. Rev. 289, 101–114 (2019).
Hor, J. L. et al. Spatiotemporally distinct interactions with dendritic cell subsets facilitates CD4+ and CD8+ T cell activation to localized viral infection. Immunity 43, 554–565 (2015).
Joshi, N. S. et al. Inflammation directs memory precursor and short-lived effector CD8+ T cell fates via the graded expression of T-bet transcription factor. Immunity 27, 281–295 (2007).
Kaech, S. M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Gautam, S. et al. The transcription factor c-Myb regulates CD8+ T cell stemness and antitumor immunity. Nat. Immunol. 20, 337–349 (2019).
Jeannet, G. et al. Essential role of the Wnt pathway effector Tcf-1 for the establishment of functional CD8 T cell memory. Proc. Natl Acad. Sci. USA 107, 9777–9782 (2010).
Zhou, X. et al. Differentiation and persistence of memory CD8+ T cells depend on T cell factor 1. Immunity 33, 229–240 (2010).
Gattinoni, L., Speiser, D. E., Lichterfeld, M. & Bonini, C. T memory stem cells in health and disease. Nat. Med. 23, 18–27 (2017).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Siddiqui, I. et al. Intratumoral Tcf1+PD-1+CD8+ T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e10 (2019).
Raghu, D., Xue, H.-H. & Mielke, L. A. Control of lymphocyte fate, infection, and tumor immunity by TCF-1. Trends Immunol. 40, 1149–1162 (2019).
Jansen, C. S. et al. An intra-tumoral niche maintains and differentiates stem-like CD8 T cells. Nature 576, 465–470 (2019).
Blank, C. U. et al. Defining ‘T cell exhaustion’. Nat. Rev. Immunol. 19, 665–674 (2019).
Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8+ T cell persistence in chronic infection. Nat. Immunol. 20, 890–901 (2019).
Guilliams, M. et al. Dendritic cells, monocytes and macrophages: a unified nomenclature based on ontogeny. Nat. Rev. Immunol. 14, 571–578 (2014).
Gerner, M. Y., Torabi-Parizi, P. & Germain, R. N. Strategically localized dendritic cells promote rapid T cell responses to lymph-borne particulate antigens. Immunity 42, 172–185 (2015).
Hickman, H. D. et al. Direct priming of antiviral CD8+ T cells in the peripheral interfollicular region of lymph nodes. Nat. Immunol. 9, 155–165 (2008).
De Simone, G et al. CXCR3 identifies human naive CD8+ T cells with enhanced effector differentiation potential. J. Immunol. 203, 3179–3189 (2019).
Hu, J. K., Kagari, T., Clingan, J. M. & Matloubian, M. Expression of chemokine receptor CXCR3 on T cells affects the balance between effector and memory CD8 T-cell generation. Proc. Natl Acad. Sci. USA 108, E118–E127 (2011).
Kohlmeier, J. E. et al. Inflammatory chemokine receptors regulate CD8+ T cell contraction and memory generation following infection. J. Exp. Med. 208, 1621–1634 (2011).
Kurachi, M. et al. Chemokine receptor CXCR3 facilitates CD8+ T cell differentiation into short-lived effector cells leading to memory degeneration. J. Exp. Med. 208, 1605–1620 (2011).
Groom, J. R. & Luster, A. D. CXCR3 ligands: redundant, collaborative and antagonistic functions. Immunol. Cell Biol. 89, 207–215 (2011).
Groom, J. R. et al. CXCR3 chemokine receptor-ligand interactions in the lymph node optimize CD4+ T helper 1 cell differentiation. Immunity 37, 1091–1103 (2012).
Sung, J. H. et al. Chemokine guidance of central memory T cells is critical for antiviral recall responses in lymph nodes. Cell 150, 1249–1263 (2012).
Kastenmuller, W., Torabi-Parizi, P., Subramanian, N., Lämmermann, T. & Germain, R. N. A spatially-organized multicellular innate immune response in lymph nodes limits systemic pathogen spread. Cell 150, 1235–1248 (2012).
Gregory, J. L. et al. Infection programs sustained lymphoid stromal cell responses and shapes lymph node remodeling upon secondary challenge. Cell Rep. 18, 406–418 (2017).
Rodda, L. B. et al. Single-cell RNA sequencing of lymph node stromal cells reveals niche-associated heterogeneity. Immunity 48, 1014–1028.e6 (2018).
Alexandre, Y. O. & Mueller, S. N. Stromal cell networks coordinate immune response generation and maintenance. Immunol. Rev. 283, 77–85 (2018).
Brown, F. D. et al. Fibroblastic reticular cells enhance T cell metabolism and survival via epigenetic remodeling. Nat. Immunol. 20, 1668–1680 (2019).
Groom, J. R. Moving to the suburbs: T-cell positioning within lymph nodes during activation and memory. Immunol. Cell Biol. 93, 330–336 (2015).
Intlekofer, A. M. et al. Requirement for T-bet in the aberrant differentiation of unhelped memory CD8+ T cells. J. Exp. Med. 204, 2015–2021 (2007).
Ollion, J., Cochennec, J., Loll, F., Escudé, C. & Boudier, T. TANGO: a generic tool for high-throughput 3D image analysis for studying nuclear organization. Bioinformatics 29, 1840–1841 (2013).
Ozga, A. J. et al. pMHC affinity controls duration of CD8+ T cell–DC interactions and imprints timing of effector differentiation versus expansion. J. Exp. Med. 213, 2811–2829 (2016).
Dose, M. et al. β-Catenin induces T-cell transformation by promoting genomic instability. Proc. Natl Acad. Sci. USA 111, 391–396 (2014).
Mielke, L. A. et al. TCF-1 limits the formation of Tc17 cells via repression of the MAF–RORγt axis. J. Exp. Med. 216, 1682–1699 (2019).
Wu, T. et al. The TCF1-Bcl6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Sci. Immunol. 1, eaai8593 (2016).
Jadhav, R. R. et al. Epigenetic signature of PD-1+ TCF1+ CD8 T cells that act as resource cells during chronic viral infection and respond to PD-1 blockade. Proc. Natl Acad. Sci. USA 116, 14113–14118 (2019).
Danilo, M., Chennupati, V., Silva, J. G., Siegert, S. & Held, W. Suppression of Tcf1 by inflammatory cytokines facilitates effector CD8 T cell differentiation. Cell Rep. 22, 2107–2117 (2018).
Huang, Z. et al. IL-27 promotes the expansion of self-renewing CD8+ T cells in persistent viral infection. J. Exp. Med. 216, 1791–1808 (2019).
Marsman, C. et al. Plasmacytoid dendritic cell heterogeneity is defined by CXCL10 expression following TLR7 stimulation. Immunol. Cell Biol. 96, 1083–1094 (2018).
Chow, M. T. et al. Intratumoral activity of the CXCR3 chemokine system is required for the efficacy of anti-PD-1 therapy. Immunity 50, 1498–1512.e5 (2019).
Jung, Y. W., Rutishauser, R. L., Joshi, N. S., Haberman, A. M. & Kaech, S. M. Differential localization of effector and memory CD8 T cell subsets in lymphoid organs during acute viral infection. J. Immunol. 185, 5315–5325 (2010).
Dyer, D. Understanding the mechanisms that facilitate specificity, not redundancy, of chemokine-mediated leukocyte recruitment. Immunology 160, 336–344 (2020).
Metzemaekers, M., Vanheule, V., Janssens, R., Struyf, S. & Proost, P. Overview of the mechanisms that may contribute to the non-redundant activities of interferon-inducible CXC chemokine receptor 3 ligands. Front. Immunol. 8, 1970 (2018).
DiToro, D. et al. Differential IL-2 expression defines developmental fates of follicular versus nonfollicular helper T cells. Science 361, eaao2933 (2018).
Sheikh, A. A. et al. Context-dependent role for T-bet in T follicular helper differentiation and germinal center function following viral infection. Cell Rep. 28, 1758–1772.e4 (2019).
Duckworth, B. C. & Groom, J. R. Conversations that count: cellular interactions that drive T cell fate. Immunol. Rev. https://doi.org/10.1111/imr.12945 (2021).
Hancock, W. W. et al. Requirement of the chemokine receptor CXCR3 for acute allograft rejection. J. Exp. Med. 192, 1515–1520 (2000).
Pircher, H., Bürki, K., Lang, R., Hengartner, H. & Zinkernagel, R. M. Tolerance induction in double specific T-cell receptor transgenic mice varies with antigen. Nature 342, 559–561 (1989).
Park, M. K. et al. The CXC chemokine murine monokine induced by IFN-γ (CXC chemokine ligand 9) is made by APCs, targets lymphocytes including activated B cells, and supports antibody responses to a bacterial pathogen in vivo. J. Immunol. 169, 1433–1443 (2002).
Dufour, J. H. et al. IFN-γ-inducible protein 10 (IP-10; CXCL10)-deficient mice reveal a role for IP-10 in effector T cell generation and trafficking. J. Immunol. 168, 3195–3204 (2002).
Hwang, S. Y. et al. A null mutation in the gene encoding a type I interferon receptor component eliminates antiproliferative and antiviral responses to interferons alpha and beta and alters macrophage responses. Proc. Natl Acad. Sci. USA 92, 11284–11288 (1995).
Mombaerts, P. et al. RAG-1-deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Li, W., Germain, R. N. & Gerner, M. Y. Multiplex, quantitative cellular analysis in large tissue volumes with clearing-enhanced 3D microscopy (Ce3D). Proc. Natl Acad. Sci. USA 114, E7321–E7330 (2017).
Preibisch, S. et al. Efficient Bayesian-based multiview deconvolution. Nat. Methods 11, 645–648 (2014).
Preibisch, S., Saalfeld, S., Schindelin, J. & Tomancak, P. Software for bead-based registration of selective plane illumination microscopy data. Nat. Methods 7, 418–419 (2010).
Ballester, M. et al. The nuclear localization of WAP and CSN genes is modified by lactogenic hormones in HC11 cells. J. Cell Biochem. 105, 262–270 (2008).
Acknowledgements
We thank S. L. Nutt, K. L. Good-Jacobson and K. D. Shortman for helpful discussions and/or critical reading of the manuscript. This work was supported by National Health and Medical Research Council (NHMRC) Ideas (no. GNT1182649) and Project (no. GNT1137989) grants to J.R.G.; grant nos. GNT1006592, GNT1045549 and GNT1065626 to M.P.; and grant no. GNT1185513 to L.A.M. B.C.D. is supported by a WEHI Academic Excellence scholarship. A.A.S. and L.D. are supported by Melbourne research scholarships. J.R.G. was supported by an Australian Research Council Future Fellowship (no. FT130100708) and is supported by a Walter and Eliza Hall Centenary Fellowship sponsored by CSL Behring. M.P. is supported by an NHMRC Investigator (no. GNT1175011) and Sylvia & Charles Viertel Senior Medical Research Fellowships. L.A.M. is supported by a Victorian Cancer Agency mid-career fellowship. The experimental scheme figures were generated with BioRender. This work was made possible through the Victorian State Government Operational Infrastructure Support and Australian Government NHMRC Independent Research Institutes Infrastructure Support Scheme.
Author information
Authors and Affiliations
Contributions
J.R.G. conceptualized the study. V.C.W., F.L., B.C.D., T.B., K.L.R. and S.N.M. devised the methodology. B.C.D., V.C.W., B.J.B., L.D., Y.O.A., C.A., A.A.S., R.Z.Q. and F.L. carried out the investigation. T.B., B.C.D., L.A.M. and F.L. carried out the software and formal analysis. V.C.W., B.C.D., F.L. and J.R.G. carried out the data vizualization. M.P. obtained the resources. J.R.G. and B.C.D. wrote the original draft and resubmission. J.R.G. acquired the funding and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quantification of cells within 3D lymph nodes.
Cell position within lymph nodes were quantified using Eroded Volume Fraction analysis. (a,b) To determine the 3D structure, images were smoothed and global thresholding was set. (b) 3D distance map is computed inside the lymph nodes where EVF values are divided into 100 layers of equal volume from 0, near the lymph node periphery, to 1, the lymph node center. (c) To position cells within the lymph nodes cell images were top-hat filtered and cell signal threshold was set manually. (d) To determine the B cell follicle signal, images were smoothed and global thresholding was set. (e) Density of B220 B cell follicles (cyan), naïve (grey) and day 6 post LCMV infection (purple) P14 cells within intact lymph nodes. Lymph node periphery (EVF = 0) to lymph node center (EVF = 1). Data are pooled from 10 mice from 3 independent experiments. Data are mean ± SEM.
Extended Data Fig. 2 Uninfected P14 cell location and expression of ZsGreen_T-bet and CXCR3 during LCMV infection.
DsRED_ZsGreen_T-bet reporter P14 cells in LCMV infected WT hosts with control or αCD62L and FTY720 blocking treatments. (a) LSFM micrographs of uninfected inguinal lymph node, showing dsRED P14 (magenta) and ZsGreen_T-bet reporter (cyan) P14 cells. (b) Frequency and number of total P14 cells in lymph nodes in control or αCD62L/FTY720 treated mice at indicated days post LCMV infection. (c) Frequency and number of ZsTbet+CXCR3+ P14 cells in peripheral lymph nodes in control or αCD62L/FTY720 treated mice at indicated days post LCMV infection. (b,c) Data 3 mice per group and represent three independent experiments, total n = 10. Data are mean ± SEM.
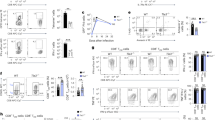
Extended Data Fig. 3 Loss of CXCR3 promotes the formation of stem-like memory precursors.
GFP_WT or GFP_Cxcr3–/– P14 cells in LCMV infected WT hosts day 6 post infection (a,b) Representative histograms of (a) SLAMF6 and (b) SCA1 expression in naïve (CD44-CD8+) host, TSCM (CD44+TCF1+CD62L+) P14 or TSLEC (CD44+KLRG1+CD62L−) populations in WT and Cxcr3–/– P14 cells, gMFI for each population is shown. (c) Frequency of TSLEC (CD44+KLRG1+CD62L−) and (CD44+KLRG1+TCF1−), and TSCM (CD44+KLRG1−CD62L+), (CD44+KLRG1−TCF1+) and (CD44+TCF1+CD62L+) adoptively transferred GFP_WT or GFP_Cxcr3–/– P14 cells and WT polyclonal (GFP−CD44+CD8+) host T cells. Data are from 4 mice, and represent 3 independent experiments, total n = 10. Data are mean ± SEM. (d,f) Plots and (e,g) frequency of TSLEC (CD44+KLRG1+CD62L−) and TSCM (CD44+KLRG1−CD62L+) (d,e) GP33 and (f,g) NP396 tetramer+CD44+ cells in WT or Cxcr3–/– cells. p value indicates t-test between WT and Cxcr3–/– cells. (e,g) Data are from 5 mice, and represent 2 independent experiments, total n = 10. Data are mean ± SEM. (h) Plots and (i) time course of emergence of TSLEC (CD44+KLRG1+TCF1−) and TSCM (CD44+KLRG1−TCF1+) at indicated days post infection (h,i) Data are 4 mice per group and represent 3 independent experiments, total n = 10. Data are mean ± SEM.
Extended Data Fig. 4 Uninfected REX3 expression and DC gating strategy.
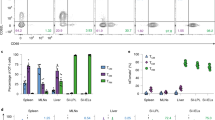
(a) LSFM micrographs of uninfected, REX3 inguinal lymph node, CXCL9-RFP (magenta), CXCL10-BFP (cyan), B220 (blue), CD31 (yellow). Lower panels show longitudinal slice through lymph node center. Data represent 6 independent replicates. (b) Gating strategy to define dendritic cell subsets. Digested lymph node cells were stained to identify CD11c+MHCII+ total DCs and subsets were defined within this gate, cDC1 (XCR1+CD11b+CD103+), cDC2 (Ly6C−SIRP1α+,CD11b+,CD103−) and moDC (CD64+Ly6C+SIRP1α+,CD11b+,CD103−).
Extended Data Fig. 5 Host deficiency in IFN-I signaling promotes stem-ness phenotype.
(a,d) Experimental schemes. GFP_P14 cells in WT or Ifnar–/– host mice were sorted day 6 post infection and transferred into naïve (a–c) Rag1–/– or (d-f) WT host mice, prior to infection with LCMV and harvest at either (a) day 14 post infection or (d) day 6 post infection (b,e) Plots and (c,f) total GFP_ P14 cells recovered from indicated hosts. Data are 3 and 4 mice per group and represent 2 independent experiments, total n = 8. Data are mean ± SEM.
Supplementary information
Supplementary Video 1
DsRED_ZsGreen_T-bet reporter P14 cells in LCMV-infected WT hosts. Cleared intact inguinal lymph node at day 6 p.i, showing dsRED P14 (magenta) ZsGreen_T-bet reporter (cyan) P14 cells.
Supplementary Video 2
DsRED_ZsGreen_T-bet reporter P14 cells in LCMV-infected WT hosts. LSFM 3D image of cleared intact inguinal lymph node at day 3 p.i, showing dsRED P14 (magenta) ZsGreen_T-bet reporter (cyan) P14 cells.
Supplementary Video 3
GFP_WT or GFP_Cxcr3–/– P14 cells in LCMV-infected WT hosts. Representative LSFM 3D images of cleared intact inguinal lymph nodes containing GFP_WT (left) or GFP_Cxcr3–/– (right) P14 cells at day 6 p.i, showing P14 cells (yellow), B220 (B cell follicles, cyan) and CD31 (lymphatic network, magenta).
Supplementary Video 4
GFP_WT or GFP_Cxcr3–/– P14 cells in LCMV-infected WT hosts with control or FTY720 treatment. LSFM 3D images of cleared intact inguinal lymph nodes containing GFP_WT (top left) or GFP_Cxcr3–/– (top right) FTY720-treated GFP_WT (lower left) or FTY720-treated GFP_Cxcr3–/– (lower right) P14 cells at day 6 p.i, showing P14 cells (yellow), B220 (B cell follicles, cyan) and CD31 (lymphatic network, magenta).
Supplementary Video 5
Intact REX3 reporter mice were infected with LCMV, and lymph nodes were analyzed at the indicated days post-infection. Representative LSFM 3D images of a cleared intact REX3 inguinal lymph node day 6 post-infection, showing CXCL9-RFP (magenta), CXCL10-BFP (cyan) and CD31 (lymphatic network, yellow).
Supplementary Video 6
REX3 bone-marrow-chimeric mice were infected with LCMV, and lymph nodes were analyzed at the indicated days post-infection. Representative LSFM 3D images of cleared REX3 BM chimeric inguinal lymph node day 6 post-infection, showing CXCL9-RFP (magenta), CXCL10-BFP (cyan) and CD31 (lymphatic network, yellow).
Supplementary Video 7
REX3 host chimeric mice were infected with LCMV, and lymph nodes were analyzed at the indicated days post-infection. Representative LSFM 3D images of cleared REX3 host chimeric inguinal lymph node day 6 post-infection, showing CXCL9-RFP (magenta), CXCL10-BFP (cyan) and CD31 (lymphatic network, yellow).
Supplementary Video 8
GFP_P14 cells in LCMV-infected WT and Ifnar–/– hosts. Representative LSFM 3D images of cleared intact popliteal lymph nodes containing WT (left) or Ifnar–/– (right) hosts at day 6 post-infection, showing P14 cells (yellow), B220 (B cell follicles, cyan) and CD31 (lymphatic network, magenta).
Rights and permissions
About this article
Cite this article
Duckworth, B.C., Lafouresse, F., Wimmer, V.C. et al. Effector and stem-like memory cell fates are imprinted in distinct lymph node niches directed by CXCR3 ligands. Nat Immunol 22, 434–448 (2021). https://doi.org/10.1038/s41590-021-00878-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-00878-5
This article is cited by
-
How chemokines organize the tumour microenvironment
Nature Reviews Cancer (2024)
-
Structural insights into the activation and inhibition of CXC chemokine receptor 3
Nature Structural & Molecular Biology (2024)
-
Antigen presenting cells in cancer immunity and mediation of immune checkpoint blockade
Clinical & Experimental Metastasis (2024)
-
Targeting the epigenome to reinvigorate T cells for cancer immunotherapy
Military Medical Research (2023)
-
C5aR+ dendritic cells fine-tune the Peyer’s patch microenvironment to induce antigen-specific CD8+ T cells
npj Vaccines (2023)