Abstract
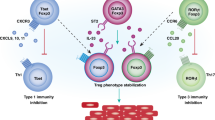
Tissue-resident memory T (TRM) cells are non-recirculating cells that exist throughout the body. Although TRM cells in various organs rely on common transcriptional networks to establish tissue residency, location-specific factors adapt these cells to their tissue of lodgment. Here we analyze TRM cell heterogeneity between organs and find that the different environments in which these cells differentiate dictate TRM cell function, durability and malleability. We find that unequal responsiveness to TGFβ is a major driver of this diversity. Notably, dampened TGFβ signaling results in CD103− TRM cells with increased proliferative potential, enhanced function and reduced longevity compared with their TGFβ-responsive CD103+ TRM counterparts. Furthermore, whereas CD103− TRM cells readily modified their phenotype upon relocation, CD103+ TRM cells were comparatively resistant to transdifferentiation. Thus, despite common requirements for TRM cell development, tissue adaptation of these cells confers discrete functional properties such that TRM cells exist along a spectrum of differentiation potential that is governed by their local tissue microenvironment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The RNA-sequencing data generated from this study have been deposited in the GEO under accession code GSE178769. The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request. Source data are provided with this paper.
Code availability
The code generated and used for the analysis of RNA-sequencing data are available from the corresponding author on reasonable request.
References
Jameson, S. C. & Masopust, D. Understanding subset diversity in T cell memory. Immunity 48, 214–226 (2018).
Mackay, L. K. et al. Long-lived epithelial immunity by tissue-resident memory T (TRM) cells in the absence of persisting local antigen presentation. Proc. Natl Acad. Sci. USA 109, 7037–7042 (2012).
Shin, H. & Iwasaki, A. A vaccine strategy that protects against genital herpes by establishing local memory T cells. Nature 491, 463–467 (2012).
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483, 227–231 (2012).
Mackay, L. K. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 352, 459–463 (2016).
Mackay, L. K. et al. The developmental pathway for CD103+CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Wakim, L. M. et al. The molecular signature of tissue resident memory CD8 T cells isolated from the brain. J. Immunol. 189, 3462–3471 (2012).
Kumar, B. V. et al. Human tissue-resident memory T cells are defined by core transcriptional and functional signatures in lymphoid and mucosal sites. Cell Rep. 20, 2921–2934 (2017).
Masopust, D. & Soerens, A. G. Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37, 521–546 (2019).
Omilusik, K. D. & Goldrath, A. W. Remembering to remember: T cell memory maintenance and plasticity. Curr. Opin. Immunol. 58, 89–97 (2019).
Milner, J. J. et al. Heterogenous populations of tissue-resident CD8+ T cells are generated in response to infection and malignancy. Immunity 52, 808–824.e807 (2020).
Kurd, N. S. et al. Early precursors and molecular determinants of tissue-resident memory CD8+ T lymphocytes revealed by single-cell RNA sequencing. Sci. Immunol. 5, eaaz6894 (2020).
Mueller, S. N. & Mackay, L. K. Tissue-resident memory T cells: local specialists in immune defence. Nat. Rev. Immunol. 16, 79–89 (2016).
Sasson, S. C., Gordon, C. L., Christo, S. N., Klenerman, P. & Mackay, L. K. Local heroes or villains: tissue-resident memory T cells in human health and disease. Cell. Mol. Immunol. 17, 113–122 (2020).
Park, C. O. & Kupper, T. S. The emerging role of resident memory T cells in protective immunity and inflammatory disease. Nat. Med. 21, 688–697 (2015).
Mani, V. et al. Migratory DCs activate TGF-β to precondition naïve CD8+ T cells for tissue-resident memory fate. Science 366, eaav5728 (2019).
Iborra, S. et al. Optimal generation of tissue-resident but not circulating memory T cells during viral infection requires crosspriming by DNGR-1+ dendritic cells. Immunity 45, 847–860 (2016).
Kok, L. et al. A committed tissue-resident memory T cell precursor within the circulating CD8+ effector T cell pool. J. Exp. Med. 217, e20191711 (2020).
Skon, C. N. et al. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat. Immunol. 14, 1285–1293 (2013).
Casey, K. A. et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 188, 4866–4875 (2012).
Steinert, E. M. et al. Quantifying memory CD8 T cells reveals regionalization of immunosurveillance. Cell 161, 737–749 (2015).
Zhang, N. & Bevan, M. J. Transforming growth factor-β signaling controls the formation and maintenance of gut-resident memory T cells by regulating migration and retention. Immunity 39, 687–696 (2013).
Sheridan, B. S. et al. Oral infection drives a distinct population of intestinal resident memory CD8+ T cells with enhanced protective function. Immunity 40, 747–757 (2014).
Mohammed, J. et al. Stromal cells control the epithelial residence of DCs and memory T cells by regulated activation of TGF-β. Nat. Immunol. 17, 414–421 (2016).
Thom, J. T., Weber, T. C., Walton, S. M., Torti, N. & Oxenius, A. The salivary gland acts as a sink for tissue-resident memory CD8+ T cells, facilitating protection from local cytomegalovirus infection. Cell Rep. 13, 1125–1136 (2015).
Hu, Y., Lee, Y. T., Kaech, S. M., Garvy, B. & Cauley, L. S. Smad4 promotes differentiation of effector and circulating memory CD8 T cells but is dispensable for tissue-resident memory CD8 T cells. J. Immunol. 194, 2407–2414 (2015).
Wakim, L. M., Woodward-Davis, A. & Bevan, M. J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. Proc. Natl Acad. Sci. USA 107, 17872–17879 (2010).
Bergsbaken, T. & Bevan, M. J. Proinflammatory microenvironments within the intestine regulate the differentiation of tissue-resident CD8+ T cells responding to infection. Nat. Immunol. 16, 406–414 (2015).
Barber, D. L. et al. Restoring function in exhausted CD8 T cells during chronic viral infection. Nature 439, 682–687 (2006).
Mackay, L. K. et al. T-box transcription factors combine with the cytokines TGF-β and IL-15 to control tissue-resident memory T cell fate. Immunity 43, 1101–1111 (2015).
Laidlaw, B. J. et al. CD4+ T cell help guides formation of CD103+ lung-resident memory CD8+ T cells during influenza viral infection. Immunity 41, 633–645 (2014).
Klose, C. S. et al. The transcription factor T-bet is induced by IL-15 and thymic agonist selection and controls CD8αα+ intraepithelial lymphocyte development. Immunity 41, 230–243 (2014).
McNamara, H. A. et al. Up-regulation of LFA-1 allows liver-resident memory T cells to patrol and remain in the hepatic sinusoids. Sci. Immunol. 2, eaaj1996 (2017).
Bartholin, L. et al. Generation of mice with conditionally activated transforming growth factor β signaling through the TβRI/ALK5 receptor. Genesis 46, 724–731 (2008).
Fonseca, R. et al. Developmental plasticity allows outside-in immune responses by resident memory T cells. Nat. Immunol. 21, 412–421 (2020).
Behr, F. M. et al. Tissue-resident memory CD8+ T cells shape local and systemic secondary T cell responses. Nat. Immunol. 21, 1070–1081 (2020).
Klicznik, M. M. et al. Human CD4+CD103+ cutaneous resident memory T cells are found in the circulation of healthy individuals. Sci. Immunol. 4, eaav8995 (2019).
Mackay, L. K. & Kallies, A. Transcriptional regulation of tissue-resident lymphocytes. Trends Immunol. 38, 94–103 (2017).
Milner, J. J. et al. Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumours. Nature 552, 253–257 (2017).
Frizzell, H. et al. Organ-specific isoform selection of fatty acid-binding proteins in tissue-resident lymphocytes. Sci. Immunol. 5, eaay9283 (2020).
Walsh, D. A. et al. The functional requirement for CD69 in establishment of resident memory CD8+ T cells varies with tissue location. J. Immunol. 203, 946–955 (2019).
Schenkel, J. M. et al. IL-15-independent maintenance of tissue-resident and boosted effector memory CD8 T cells. J. Immunol. 196, 3920–3926 (2016).
Bommireddy, R. et al. TGF-β1 regulates lymphocyte homeostasis by preventing activation and subsequent apoptosis of peripheral lymphocytes. J. Immunol. 170, 4612–4622 (2003).
Verma, N. K. et al. Leukocyte function-associated antigen-1/intercellular adhesion molecule-1 interaction induces a novel genetic signature resulting in T-cells refractory to transforming growth factor-β signaling. J. Biol. Chem. 287, 27204–27216 (2012).
Daussy, C. et al. T-bet and Eomes instruct the development of two distinct natural killer cell lineages in the liver and in the bone marrow. J. Exp. Med. 211, 563–577 (2014).
Robinette, M. L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015).
Clark, R. A. Resident memory T cells in human health and disease. Sci. Transl. Med. 7, 269rv1 (2015).
Beura, L. K. et al. T cells in nonlymphoid tissues give rise to lymph-node-resident memory T cells. Immunity 48, 327–338.e5 (2018).
Park, S. L. et al. Tissue-resident memory CD8+ T cells promote melanoma–immune equilibrium in skin. Nature 565, 366–371 (2019).
Acknowledgements
We thank the Flow Cytometry Unit and Bioresources Facility at Peter Doherty Institute (University of Melbourne) for technical assistance. This work was supported by a Howard Hughes Medical Institute and Bill & Melinda Gates International Research Scholarship OPP1175796 to L.K.M. and National Health and Medical Research Council (NHMRC) APP1129711 to L.K.M. S.L.P. was supported by a Cancer Council Victoria Postdoctoral Fellowship and an NHMRC Emerging Leadership Investigator Grant. F.S.-F.-G. was supported by a project grant from the NHMRC (no. 1140406), and a grant (no. 1158085) awarded through the Priority driven Collaborative Cancer Research Scheme and funded by Cure Cancer Australia with the assistance of Cancer Australia. L.K.M. is a Senior Medical Research Fellow supported by the Sylvia and Charles Viertel Charitable Foundation.
Author information
Authors and Affiliations
Contributions
S.N.C., S.L.P., R.F., L.C.G., M.E., D.C., T.N.B., D.M.N., Y.O.A. and N.C. performed experiments and analyzed data; W.S. and T.P.S. provided supervision; D.G.P., F.S.F.G., G.T.B., L.B., N.D.H., A.L. and M.L. generated and/or provided reagents; S.N.C., N.M.Z., S.N.M., W.R.H., F.G., F.R.C., A.K. and L.K.M. contributed to experimental design. S.N.C., S.L.P., F.R.C. and L.K.M. prepared the manuscript. L.K.M. provided funding and led the research program.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Immunology thanks Ian Cockburn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Phenotypic and functional heterogeneity between skin and liver TRM.
a,b, C57BL/6 mice received naïve gBT-I.CD45.1 or P14.CD45.1 transgenic CD8+ T cells (Vα2+CD45.1+) prior to infection with HSV or LCMV, respectively. (a) Experimental schematic. (b) Expression of indicated proteins by CD69+CD62L− gBT-I or P14 TRM from skin (red) and liver (dark grey), respectively, and splenic CD69−CD62L− P14 TEM (light grey). Dashed line represents the MFI of splenic TEM. c,d, Mice received in vitro activated effector OT-I.CD45.1 T cells (Vα2+CD45.1+) and were treated on the flank with DNFB. (c) Experimental schematic. (d) Expression of indicated molecules by CD69+CD62L− OT-I TRM isolated from skin (red) or the liver (dark grey), and splenic CD69−CD62L− OT-I TEM (light grey) >30 d post transfer. Dashed line represents the MFI of splenic TEM. e, f, Mice received effector OT-I.CD45.1 T cells and were treated on the flank with DNFB. CD69+ OT-I TRM were isolated from the skin and liver of TRM-bearing mice after 30 d and cells co-cultured with OVA peptide or gB control peptide ex vivo for 4 h. (e) MFI of IFNγ, TNF and IL-2 by skin and liver CD69+ OT-I TRM. Expression is relative to gB control peptide (Ctrl; normalized to 1). (f) Expression of IFNγ, TNF and IL-2 expression by CD69+ OT-I TRM in the skin (red) and liver (dark grey) or the corresponding gB control peptide (Ctrl; light grey) in each organ. Shaded area represents frequency of cytokine expression. g,h, Mice received naïve OT-I.CD45.1 or P14.CD45.1 T cells (Vα2+CD45.1+) prior to infection with HSV-OVA or LCMV, respectively. (g) Experimental schematic. (h) Enumeration of CD69+ CD45.1+ OT-I or P14 TRM isolated from the skin and liver, respectively, at 14, 30 and >60 d p.i. t1/2 denotes half-life. Data are pooled from 3 (b), 2-3 (d-f) or 1-5 (h) independent experiments with n = 9 (b), n = 11 (d), n = 9 (e-f) or n = 10-18 (h) mice. Two-tailed Wilcoxon paired rank test. Bars represent the mean; error bars represent mean ± SEM; symbols represent individual mice.
Extended Data Fig. 2 Skin and liver TRM exhibit common transcriptional and cytokine requirements.
a, Mice were co-transferred with naïve OT-I.CD45.1×CD45.2 (OT-I.WT) and OT-I.CD45.1.Tbx21−/− (OT-I.Tbx21−/−) T cells prior to infection with HSV-OVA or LmOVA. Enumeration of CD69+ OT-I.WT and OT-I.Tbx21−/− TRM in the skin (HSV-OVA) and liver (LmOVA) >60 d p.i (left panel) and the ratio of OT-I.Tbx21−/− T cells relative to OT-I.WT in the indicated organs (right panel). b, Mice were co-transferred with naïve OT-I.WT and OT-I.CD45.1.Eomes−/− (OT-I.Eomes−/−) T cells prior to infection with HSV-OVA or LmOVA. Enumeration of CD69+ OT-I.WT and OT-I.Eomes−/− TRM in the skin (HSV-OVA) and liver (LmOVA) >60 d p.i (left panels) and the ratio of OT-I.Eomes−/− T cells relative to OT-I.WT in the indicated organs (right panel). c, C57BL/6 (WT) or Il-15−/− mice received effector OT-I.WT T cells and were treated on the flank with DNFB. Enumeration of CD69+ OT-I TRM in the skin and liver of WT or Il-15−/− mice >30 d post transfer (left panels) and the ratio of OT-I T cells from Il-15−/− mice relative to WT mice in the indicated organs (right panel). Data are pooled from 2 independent experiments with n = 10 mice. NS, p > 0.05, two-tailed Wilcoxon paired rank test (a,b) or two-tailed Mann Whitney test (c). Bars represent the mean; error bars represent mean ± SEM; symbols represent individual mice.
Extended Data Fig. 3 Skin but not liver TRM require TGF-β for development.
a,b, Mice received effector OT-I.CD45.1×CD45.2 (WT) and OT-I.CD45.1.Tgfbr2−/− (Tgfbr2−/−) T cells and were treated on the flank with DNFB. (a) CD69 and CD103 expression by WT and Tgfbr2−/− T cells in the skin and liver at the indicated times. (b) Ratio of Tgfbr2−/− to WT T cells in the indicated organs 14 and >30 d post transfer. c-e Effector WT and Tgfbr2−/− T cells were transferred into mice infected with (c) HSV-OVA or (d) Lm-OVA. Enumeration of WT and Tgfbr2−/− T cells in the spleen and skin >30 d post- infection. (e) Representative histograms and MFI of Eomes and T-bet by CD69+ WT (red) and Tgfbr2−/− (dark grey) T cells in the skin of HSV-OVA infected mice and liver of Lm-OVA infected mice and their respective splenic WT T cells (light grey) 10 d p.i. f,g, Effector CD8+ T cells were transduced with Eomes-GFP (Eomes RV+) or Tbx21-GFP (T-bet RV+) and control-GFP (Ctrl RV+) retroviruses (RV). Equal numbers of GFP-expressing cells were transferred into mice that were treated on the flank with DNFB. (f) Enumeration of GFP-expressing Eomes RV+ or Ctrl RV+ CD8+CD69+ TRM isolated from skin and the liver (left panels) and ratio of Eomes RV+ T cells relative to Ctrl RV+ T cells in the indicated organs (right panel) 14 d post transfer. (g) Ratio of T-bet RV+ T cells relative to Ctrl RV+ T cells in the indicated organs 14 d post transfer. Data are pooled from 3 (a-b, d, f, g), 1-2 (c) or 4 (e) independent experiments with n = 13−15 (a-b), n = 5-10 (c-d), n = 12 (e) or n = 18 (f-g) mice. NS, p > 0.05, two-tailed Wilcoxon paired rank test (b, d, f; right panel) or two-tailed Mann Whitney test (c,f; left panel). Bars represent the mean; error bars represent mean ± SEM; symbols represent individual mice.
Extended Data Fig. 4 TGF-β regulates the transcriptional profile of skin TRM.
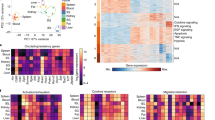
(a) Expression of TRM-associated genes in TGF-β-treated effector OT-I T cells relative to untreated cells determined by qPCR. b-d, Effector OT-I.WT or OT-I.Tgfbr2−/− T cells were transferred into mice infected with HSV-OVA. WT and Tgfbr2−/− T cells were sorted from the skin 14 d p.i. for RNAseq. (b) Heatmap of standardized log expression (gene-wise Z-score) of top 100 DEG between WT versus Tgfbr2−/−. (c) Barcode enrichment plots showing std.log-fc established signatures5: skin TRM versus spleen TEM comparison (upper panel) and liver TRM versus spleen TEM comparison (lower panel) both for the top 100 TGF-β-regulated genes. (d) Barcode enrichment plots showing standardized log fold changes (std.log-fc) from established signatures5: skin TRM v spleen TEM comparison (upper panel) and liver TRM v spleen TEM comparison (lower panel) for the top 100 upregulated genes in the skin WT versus Tgfbr2−/−. Data are pooled from n = 3 (a) or n = 2 (b) biological replicates.
Extended Data Fig. 5 Differential expression of LFA-1 on skin and liver TRM.
(a) DNFB-treated mice infected with LCMV and >30 d p.i, expression of CD11a by CD8+CD44hi T cells in the spleen and CD8+CD44hiCD69+ TRM in the skin and liver was assessed. Shown is a representative histogram (left plot) and quantified mean fluorescence intensity (MFI; right graphs) of CD11a. (b) Mixed bone chimeras reconstituted with WT and Tgfbr1CA cells were infected with LCMV and were treated with DNFB on the flank. Shown is the MFI of CD11a by WT and Tgfbr1CA T cells in the spleen and TRM in the skin and liver >30 d p.i. Data are pooled from 2 independent experiments with n = 6 mice. NS, p > 0.05, one-way ANOVA with Tukey’s multiple comparisons test (a) or two-tailed Wilcoxon paired rank test (b). Bars represent the mean; error bars represent mean ± SEM; symbols represent individual mice.
Extended Data Fig. 6 Contribution of ex-TRM to secondary memory responses.
a-d Mice received effector OT-I.CD45.1 T cells and were treated on the flank with DNFB. Liver CD69+ TRM or CD69− TCIRC from the liver and spleen were sorted >30 d and transferred into separate recipient mice infected with HSV-OVA. (a) Experimental schematic. (b) Recovery of ex-liver TRM, ex-liver and ex-spleen OT-I TCIRC (Vα2+ CD45.1+) from the liver of mice 14 d p.i. (c) Enumeration and (d) expression of CD69 and CD103 on ex-liver TRM and ex-liver and ex-spleen OT-I TCIRC 14 d p.i. e, Mice received effector OT-I.CD45.1 T cells and >21 d later, were treated with anti-Gr-1 +/− anti-CXCR3 Ab and assessed after 7 d. Shown is the experimental schematic (left panel) and Ly6C and CXCR3 expression in the liver of control (+TCIRC), anti-Gr-1-treated (-TCIRC) or anti-Gr-1 + anti-CXCR3 (-TCIRC -TRM) groups (right panels). f-i, Mice received effector OT-I.CD45.1 T cells and >21 d later, were treated with anti-Gr-1 +/− anti-CXCR3 Ab before being surgically joined to naïve syngeneic partners 7 d after treatment. Conjoined mice received OVA peptide and adjuvant (PolyI:C and anti-CD40) i.v. (as indicated +/−OVA) and tissues harvested 24-27 d post-surgery. (f) Recovery of ex-TRM OT-I cells (Vα2+CD45.1+) in the spleen of host and partner parabionts. (g) Enumeration of ex-TRM OT-I T cells in the partner parabiont. Proportion of ex-TRM OT-I T cells expressing (h) CD69 or (i) CD69 and CD103 (left) and representative plot (right) in the partner parabiont. Data are pooled from 2 (b-d) or 5 (e) independent experiments with n = 7-9 (b-d) or n = 9−13 (e) mice. Data are pooled from 1-2 independent experiments with n = 2 or n = 12 mice from 1 or 6 pairs, respectively (f-i). NS, p > 0.05, one-way ANOVA Kruskal-Wallis with Dunn’s multiple comparison’s test. Bars represent the mean; error bars represent mean ± SEM; symbols represent individual mice.
Extended Data Fig. 7 Phenotypic heterogeneity between CD103+ and CD103− CD69+ TRM in the salivary gland.
a, Mice received naïve P14.CD45.1 T cells (Vα2+CD45.1+) and were infected with LCMV. SG P14 T cells were analyzed >30 d p.i. MFI of indicated proteins relative to splenic CD69−CD62L− TEM (normalized to 1). b,c, Mice were co-transferred with effector OT-I.WT and OT-I.Tgfbr2−/− T cells. (b) Enumeration of CD69+ WT and Tgfbr2−/− T cells and (c) MFI of indicated proteins in CD69-expressing CD103− and CD103+ TRM from the SG > 30 d post transfer. Data are pooled from 3 independent experiments with n = 9 (a) or n = 15 (b-c) mice. NS, p > 0.05, two-tailed Wilcoxon paired rank test (a) or two-tailed Mann Whitney test (b,c). Violin plots represent the distribution of minimum to maximum values, and dotted line represents the median. Bars represent the mean; error bars represent mean ± SEM; symbols represent individual mice.
Extended Data Fig. 8 Circulating memory T cells exhibit superior expansion and tissue infiltration than TRM.
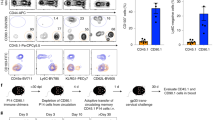
a-d, Mice received naive P14.CD45.1 T cells and were infected with LCMV. SG CD69+CD103 TRM and splenic CD69− TCIRC P14 T cells were sort purified >30 d p.i. and transferred into recipient mice infected with LCMV. (a) Experimental schematic. (b) Enumeration of ex-SG CD103− TRM and ex-spleen TCIRC P14 cells (c) expression of CD69 and CD103 and (d) enumeration of ex-SG CD103− TRM and ex-spleen TCIRC CD69+ P14 T cells in the indicated organs 14 d p.i. Data are pooled from 2 independent experiments with n = 8 mice. Two-tailed Mann Whitney test. Bars represent the mean; error bars represent mean ± SEM; symbols represent individual mice.
Supplementary information
Supplementary Information
Supplementary Methods.
Supplementary Table 1
List of antibodies used in this study.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Christo, S.N., Evrard, M., Park, S.L. et al. Discrete tissue microenvironments instruct diversity in resident memory T cell function and plasticity. Nat Immunol 22, 1140–1151 (2021). https://doi.org/10.1038/s41590-021-01004-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-01004-1
This article is cited by
-
Joint-specific memory, resident memory T cells and the rolling window of opportunity in arthritis
Nature Reviews Rheumatology (2024)
-
Targeting pathogenic CD8+ tissue-resident T cells with chimeric antigen receptor therapy in murine autoimmune cholangitis
Nature Communications (2024)
-
Defective mitochondria remodelling in B cells leads to an aged immune response
Nature Communications (2024)
-
Prostaglandin E2 controls the metabolic adaptation of T cells to the intestinal microenvironment
Nature Communications (2024)
-
Aging modifies endometrial dendritic cell function and unconventional double negative T cells in the human genital mucosa
Immunity & Ageing (2023)