Abstract
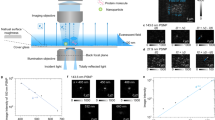
Measuring the binding kinetics of single proteins represents one of the most important and challenging tasks in protein analysis. Here we show that this is possible using a surface plasmon resonance (SPR) scattering technique. SPR is a popular label-free detection technology because of its extraordinary sensitivity, but it has never been used for imaging single proteins. We overcome this limitation by imaging scattering of surface plasmonic waves by proteins. This allows us to image single proteins, measure their sizes and identify them based on their specific binding to antibodies. We further show that it is possible to quantify protein binding kinetics by counting the binding of individual molecules, providing a digital method to measure binding kinetics and analyze heterogeneity of protein behavior. We anticipate that this imaging method will become an important tool for single protein analysis, especially for low volume samples, such as single cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. Source data are provided with this paper.
References
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Polanski, M. & Anderson, N. L. A list of candidate cancer biomarkers for targeted proteomics. Biomark. Insights 1, 1–48 (2007).
Homola, J. Present and future of surface plasmon resonance biosensors. Anal. Bioanal. Chem. 377, 528–539 (2003).
Phillips, K. S. & Cheng, Q. Recent advances in surface plasmon resonance based techniques for bioanalysis. Anal. Bioanal. Chem. 387, 1831–1840 (2007).
Masson, J.-F. Surface plasmon resonance clinical biosensors for medical diagnostics. ACS Sens. 2, 16–30 (2017).
Fang, Y. et al. Plasmonic imaging of electrochemical reactions of single nanoparticles. Acc. Chem. Res. 49, 2614–2624 (2016).
Huang, B., Yu, F. & Zare, R. N. Surface plasmon resonance imaging using a high numerical aperture microscope objective. Anal. Chem. 79, 2979–2983 (2007).
Wang, W. et al. Single cells and intracellular processes studied by a plasmonic-based electrochemical impedance microscopy. Nat. Chem. 3, 249–255 (2011).
Wang, W. et al. Label-free measuring and mapping of binding kinetics of membrane proteins in single living cells. Nat. Chem. 4, 846–853 (2012).
Yang, Y. Z. et al. Label-free tracking of single organelle transportation in cells with nanometer precision using a plasmonic imaging technique. Small 24, 2878–2884 (2015).
Wang, S. P. et al. Label-free imaging, detection, and mass measurement of single viruses by surface plasmon resonance. Proc. Natl Acad. Sci. USA 107, 16028–16032 (2010).
Shan, X. N. et al. Imaging the electrocatalytic activity of single nanoparticles. Nat. Nanotechnol. 7, 668–672 (2012).
Fang, Y. M. et al. Intermittent photocatalytic activity of single CdS nanoparticles. Proc. Natl Acad. Sci. USA 114, 10566–10571 (2017).
Yang, Y. T. et al. Interferometric plasmonic imaging and detection of single exosomes. Proc. Natl Acad. Sci. USA 115, 10275–10280 (2018).
Vollmer, F. & Arnold, S. Whispering-gallery-mode biosensing: label-free detection down to single molecules. Nat. Methods 5, 591–596 (2008).
Zijlstra, P., Paulo, P. M. R. & Orrit, M. Optical detection of single non-absorbing molecules using the surface plasmon resonance of a gold nanorod. Nat. Nanotechnol. 7, 379–382 (2012).
Baaske, M. D., Foreman, M. R. & Vollmer, F. Single-molecule nucleic acid interactions monitored on a label-free microcavity biosensor platform. Nat. Nanotechnol. 9, 933–939 (2014).
Mauranyapin, N. P., Madsen, L. S., Taylor, M. A., Waleed, M. & Bowen, W. P. Evanescent single-molecule biosensing with quantum-limited precision. Nat. Photonics 11, 477–481 (2017).
Zheng, Y. H. et al. Reversible gating of smart plasmonic molecular traps using thermoresponsive polymers for single-molecule detection. Nat. Commun. 6, 8797 (2015).
Gaiduk, A., Yorulmaz, M., Ruijgrok, P. V. & Orrit, M. Room-temperature detection of a single molecule’s absorption by photothermal contrast. Science 330, 353–356 (2010).
Arroyo, J. O., Cole, D. & Kukura, P. Interferometric scattering microscopy and its combination with single-molecule fluorescence imaging. Nat. Protoc. 11, 617–633 (2016).
Young, G. et al. Quantitative mass imaging of single biological macromolecules. Science 360, 423–427 (2018).
Liu, X. W. et al. Plasmonic-based electrochemical impedance imaging of electrical activities in single cells. Angew. Chem. 56, 8855–8859 (2017).
Lu, J. & Li, J. H. Label-free imaging of dynamic and transient calcium signaling in single cells. Angew. Chem. 54, 13576–13580 (2015).
Shan, X. N., Patel, U., Wang, S. P., Iglesias, R. & Tao, N. J. Imaging local electrochemical current via surface plasmon resonance. Science 327, 1363–1366 (2010).
Yu, H., Shan, X. N., Wang, S. P., Chen, H. Y. & Tao, N. J. Molecular scale origin of surface plasmon resonance biosensors. Anal. Chem. 86, 8992–8997 (2014).
Kretschmann, M. Decay of non radiative surface plasmons into light on rough silver films. Comparison of experimental and theoretical results. Opt. Commun. 6, 185–187 (1972).
Bozhevolnyi, S. I. & Coello, V. Elastic scattering of surface plasmon polaritons: modeling and experiment. Phys. Rev. B 58, 10899–10910 (1998).
Shchegrov, A. V., Novikov, I. V. & Maradudin, A. A. Scattering of surface plasmon polaritions by a circularly symmetric surface defect. Phys. Rev. Lett. 78, 4269–4272 (1997).
Sirbuly, D. J., Tao, A., Law, M., Fan, R. & Yang, P. D. Multifunctional nanowire evanescent wave optical sensors. Adv. Mater. 19, 61–66 (2007).
Agnarsson, B. et al. Evanescent light-scattering microscopy for label-free interfacial imaging: from single sub-100 nm vesicles to live cells. ACS Nano 9, 11849–11862 (2015).
Cole, D., Young, G., Weigel, A., Sebesta, A. & Kukura, P. Label-free single-molecule imaging with numerical-aperture-shaped interferometric scattering microscopy. ACS Photonics 4, 211–216 (2017).
Liebel, M., Hugall, J. T. & Hulst, N. F. Ultrasensitive label-free nanosensing and high-speed tracking of single proteins. Nano Lett. 17, 1277–1281 (2017).
Rita, S. Weighing single proteins with light. Nat. Methods 15, 477 (2018).
Simpson, W. D. & Volkmar, H. Modulation of the drag forceexerted by microfluidic flow on laser-trapped particles: A new method to assess surface-binding kinetics, analyte size, and solution viscosity. Biophys. J. 114, 692a (2018).
Huang, Y. et al. Coherent brightfield microscopy provides the spatiotemporal resolution to study early stage viral infection in live cells. ACS Nano 11, 2575–2585 (2017).
Ignatovich, F. V. & Novotny, L. Real-time and background-free detection of nanoscale particles. Phys. Rev. Lett. 96, 013901 (2006).
Deutsch, B., Beams, R. & Novotny, L. Nanoparticle detection using dual-phase interferometry. Appl. Opt. 49, 4921–4925 (2010).
Özkumur, E. et al. Label-free and dynamic detection of biomolecular interactions for high-throughput microarray applications. Proc. Natl Acad. Sci. USA 105, 7988–7992 (2008).
Piliarik, M. & Sandoghdar, V. Direct optical sensing of single unlabelled proteins and super-resolution imaging of their binding sites. Nat. Commun. 5, 4495 (2014).
S Abramoff, M. D., Magalhaes, P. J. & Ram, S. J. Image processing with imageJ. Biophotonics Int. 11, 36–42 (2004).
Tinevez, J.-Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
Acknowledgements
We are grateful for financial support from the Gordon and Betty Moore Foundation (N.T.) and the National Institute of General Medical Sciences of the National Institutes of Health grant R01GM107165 (S.W.). We acknowledge the use of facilities within the ASU NanoFab supported in part by NSF program NNCI-ECCS-1542160. The content is solely the responsibility of the authors and does not necessarily represent the official views of the sponsors.
Author information
Authors and Affiliations
Contributions
P.Z. performed the experiments and data analysis. G.M. contributed to the protein studies. W.D. performed some of the preliminary experiments with nanoparticles. Z.W. prepared gold-coated glass slides and performed atomic force microscopy measurements. S.W. contributed to the design and construction of the optical setup. N.T. and S.W. conceived and supervised the project. P.Z., G.M., S.W. and N.T. wrote the manuscript. All authors reviewed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
A US provisional patent application (62/975,473) has been filed by Arizona Board of Regents on behalf of Arizona State University for single-molecule imaging based on an early draft of this article. Inventors are N.T., S.W. and P.Z.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Tables 1 and 2 and Notes 1–19.
Supplementary Video 1
Dynamic binding of single 26-nm nanoparticles on bare gold.
Supplementary Video 2
Dynamic binding of single IgA and IgM proteins on bare gold.
Supplementary Video 3
Identification of single IgA and IgM proteins on anti-IgA antibody coated gold, and identification of single anti-CaM antibody and IgA proteins on CaM-coated gold.
Supplementary Video 4
Differential video showing the on-off process of one IgA protein.
Supplementary Video 5
Three different behaviors of binding of individual IgA molecules.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Rights and permissions
About this article
Cite this article
Zhang, P., Ma, G., Dong, W. et al. Plasmonic scattering imaging of single proteins and binding kinetics. Nat Methods 17, 1010–1017 (2020). https://doi.org/10.1038/s41592-020-0947-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0947-0
This article is cited by
-
Accessible hotspots for single-protein SERS in DNA-origami assembled gold nanorod dimers with tip-to-tip alignment
Nature Communications (2023)
-
Evanescent scattering imaging of single protein binding kinetics and DNA conformation changes
Nature Communications (2022)
-
Digital plasmonic nanobubble detection for rapid and ultrasensitive virus diagnostics
Nature Communications (2022)
-
Label-free nanofluidic scattering microscopy of size and mass of single diffusing molecules and nanoparticles
Nature Methods (2022)
-
Operando monitoring of ion activities in aqueous batteries with plasmonic fiber-optic sensors
Nature Communications (2022)