Abstract
E. coli purine nucleoside phosphorylase is a homohexamer, which structure, in the apo form, can be described as a trimer of dimers. Earlier studies suggested that ligand binding and kinetic properties are well described by two binding constants and two sets of kinetic constants. However, most of the crystal structures of this enzyme complexes with ligands do not hold the three-fold symmetry, but only two-fold symmetry, as one of the three dimers is different (both active sites in the open conformation) from the other two (one active site in the open and one in the closed conformation). Our recent detailed studies conducted over broad ligand concentration range suggest that protein–ligand complex formation in solution actually deviates from the two-binding-site model. To reveal the details of interactions present in the hexameric molecule we have engineered a single tryptophan Y160W mutant, responding with substantial intrinsic fluorescence change upon ligand binding. By observing various physical properties of the protein and its various complexes with substrate and substrate analogues we have shown that indeed three-binding-site model is necessary to properly describe binding of ligands by both the wild type enzyme and the Y160W mutant. Thus we have pointed out that a symmetrical dimer with both active sites in the open conformation is not forced to adopt this conformation by interactions in the crystal, but most probably the dimers forming the hexamer in solution are not equivalent as well. This, in turn, implies that an allosteric cooperation occurs not only within a dimer, but also among all three dimers forming a hexameric molecule.
Similar content being viewed by others
Introduction
Purine nucleoside phosphorylases (PNP, E.C. 2.4.2.1.) are ubiquitous enzymes of the purine salvage pathway enabling the cell to recover purine bases and ribose-1-phosphate from used purine nucleotides1. PNPs catalyse the following reaction: β-purine nucleoside + orthophosphate ↔ purine base + α-D-pentose-1-phosphate. Trimeric sub-family contains PNPs specific vs. 6-oxopurine nucleosides, and such enzymes are found in mammals and in some microorganisms, while hexameric PNPs present mainly in prokaryotes are less specific and accept many substituents at position 6 of the purine ring1. Both subfamilies are potential targets for antitumour, immunosuppressive, and/or antiparasitic agents1,2,3. Differences in specificity between trimeric mammalian PNPs and hexameric, less specific PNPs offer a basis for enzyme-activating prodrug gene therapy by empowering selective killing of tumour cells expressing the E. coli PNP gene4,5,6,7. These potential medical applications of PNPs encourage researchers to study their molecular mechanism and to search for their potent, selective and membrane permeable inhibitors.
Nevertheless, PNPs are also remarkable molecular machines and many molecular features that make them such efficient catalysts are still unclear. The biologically active form of the E. coli enzyme is a homohexamer, consisting of six monomers with an identical aminoacid sequence and, in the apo form, exhibiting also identical three-dimensional structure. However, the whole hexameric molecule has a three-fold symmetry, and can be described as a trimer of dimers. Moreover, each of the dimers has a two-fold symmetry, with the symmetry axis perpendicular to three-fold symmetry axis, so the apo enzyme may be regarded as a dimer of trimers as well (Fig. 1, left panel)8. Dimers, but not monomers, seem to have all the necessary features to conduct catalysis, because monomers within a dimer mutually donate two amino acids to complete the active site of the neighbour9,10. The earlier models describing enzyme kinetics and ligand binding have included two sets of kinetic and two binding constants, respectively, reflecting allosteric cooperation between subunits in a dimer9,11,12. Therefore, the hexameric (not simply dimeric) form adopted by this enzyme seems to be necessary solely in order to ensure stability of the enzyme molecule. This reason for homooligomerisation is observed for many other enzymes14 and we have already shown that it is important for this particular enzyme as well15.
Left panel: Schematic view of the E. coli PNP in the apo form (PDB 1ECP8). All dimers are symmetrical, as all subunits have the same, elongated structure of the helix H8, resulting in the open conformation of the active site. The overall structure of the enzyme may be therefore regarded as a trimer of dimers as well as a dimer of trimers since the molecule has 32 point group symmetry. Middle panel: Schematic view of the E. coli PNP complexed with phosphate (PDB 4TS319). In two subunits (here shown in cyan) binding of phosphate leads to the segmentation of the H8 helix and the partial closing of the active site pocket. Two dimers become unsymmetrical (shown here with cyan and green subunits), as they differ in the conformation of the N-terminal part of the H8 helix (shown in red). The third dimer (with green subunits) has both active sites in the open conformation, hence is symmetrical. The overall structure of the enzyme in this case is therefore a dimer of trimers. This kind of architecture is observed in the most of the crystal structures of E. coli PNP complexes available up to now, binary (with phosphate) and ternary (with phosphate and various nucleosides or purine bases). Right panel: Subunits in the open and closed conformation of the active site from the structure depicted in the middle panel, were overlaid to show differences in these two active site conformations: the segmentation of the helix H8 and the movement of its N-terminal part (shown in darker shades of green and cyan, respectively) and the loop towards the active site pocket (indicated by the red arrow).
However, most of the crystal structures of binary (with phosphate) and ternary (with phosphate and various nucleosides or purine bases) complexes of this enzyme available so far do not hold this three-fold symmetry13,16,17,18,19. Binding of a phosphate substrate, induces the conformational change leading to the closure of the active site pocket (Fig. 1, right panel). But this occurs only in some subunits. Two of the dimers are unsymmetrical, with one active site in the open and the other in the closed active site conformation (“open-closed” dimer), whereas the third dimer has both active sites in the open conformation (Fig. 1, middle panel). Since the closure of the active site is essential for catalysis9,13, only two “open-closed” dimers seem to be active.
Our earlier X-ray studies13,19 suggest that the open sites of the “open-open” dimer differ from the open sites of the “open-closed” dimer. In this study, we present the results of our research addressing the question whether the “open-open” dimer is forced to be symmetric by some interactions present only in the crystalline form, or it also differs from its neighbours in solution. In the latter case, its presence should be taken into account in the study of the molecular mechanism of catalysis. Therefore, we have examined in detail the properties in solution of the binary complex of E. coli PNP with phosphate, and of the ternary complex with phosphate and the nucleoside, formycin A, the structural analogue of the substrate, adenosine, and the potent inhibitor (formycin A, see Fig. 2, inset).
Comparison of active sites of the Y160W mutant (PDB 6XZ2, violet) and the WT PNP (PDB 4TS319, green) complexed with formycin A and sulfate. Aminoacids forming the active site and the network of hydrogen bonds are shown. The residues His4 and Arg43 belonging to the neighbouring subunit in a dimer are marked with *. Inset: Formycin A, a structural analogue of adenosine, the natural E. coli PNP substrate. Due to the C–C glycosidic bond, that links the base and the pentose, formycin A is not a substrate for PNP, but an inhibitor, competitive vs. the nucleoside substrate. Note the difference in the base ring numbering when compared with the purine ring numbering.
Analysis of the fluorescence titration data for complex formation of PNP with phosphate was the first indication that two types of dimers observed in the crystallographic structures were not just a crystallographic artefact. As the wild type PNP exhibits uniform changes of fluorescence upon phosphate binding, all titration curves, irrespectively of excitation and observation wavelength, have the same shape and are not visibly multiphase, two approaches were employed to further investigate the observed phenomena. First, a single-tryptophan Y160W mutant was engineered, with the mutation in a close proximity to the active site, but without significant interactions with ligands (Fig. 2). Since each subunit of the wild type PNP contains six tyrosine residues and no tryptophan1, this approach resulted in a protein with a significantly enhanced fluorescence response to ligand binding, and with retained catalytic and kinetic properties of the WT enzyme. Second, several experimental techniques were used to study the formation of binary and ternary complexes in order to explore the physical properties of the enzyme other than the intrinsic fluorescence of the protein.
Results
Y160W mutant shares typical features with the WT E. coli PNP
Analytical centrifugation confirmed that Y160W mutation did not affect the oligomeric state of the protein in solution. Virtually all protein molecules form a single band profile (Figure S1) with a sedimentation coefficient s20,w = 7.80 which corresponds to the molecular weight of 150 kDa, in agreement with the hexameric assembly (6 × 25.970 = 155.8 kDa).
Crystallographic structure of the ternary complex of the Y160W mutant with two inhibitors, substrate analogues, sulfate ion and formycin A (PDB 6XZ2) shows that mutation has no significant influence on the secondary and ternary structure of the protein as well. Also, interactions of formycin and sulfate in the active site of the enzyme are comparable to those observed for the WT enzyme (Fig. 2). Complex crystallised in P 6122 space group with the half of the hexamer in the asymmetric unit. All active sites are in the open conformation, consistent with the data on protein crystallisation in the absence of phosphate conditions13. Both ligands are in the same positions as in the active site of the WT PNP (PDB 4TS3, 4TS919). Sulfate ion is bound in the hydrogen bond distance from Arg87, Arg43* (from the neighbouring subunit), Ser90, Gly20. The side of chain Arg24 points away from the active site, what is typical for the open conformation of the active site9,13. Ribose of formycin A is involved in hydrogen bonds with sulfate ion, Glu181 and His4* (from the neighbouring subunit), whereas aglycon—with Asp204 and Ser90.
The effect of the Y160W mutation on catalytic properties of the enzyme was checked by measuring the specific activity of the mutant vs. typical PNP substrates (Table 1), by collecting full kinetic data for guanosine, 7-methylguanosine and phosphate as variable substrates (Table 2), and by measuring the effect on the catalysis of the typical inhibitor, formycin A, competitive vs. nucleoside. Activity of the WT PNP and the mutant towards natural and modified substrates is very similar, but kinetic constants tend to be slightly higher for the mutant than those observed for the WT enzyme. However, all reaction models are retained: Michaelis–Menten for the nucleoside substrate, two allosterically interacting active sites for phosphate as a variable substrate, and competitive inhibition of formycin A vs. nucleoside substrate. All these results justify the use of the Y160W mutant to study formation of E. coli PNP complexes with ligands.
Effects of enzyme-phosphate interaction on enzyme fluorescence
For the WT protein, as it has no tryptophan in the structure, increasing concentration of phosphate (Pi) in solution leads to only small quenching of the fluorescence without any significant changes in the shape of the protein fluorescence spectra, regardless of the excitation wavelength (Fig. 3). However, for the Y160W mutant changes in the protein fluorescence upon phosphate binding are much more diverse. Fluorescence spectra were collected with excitation at the maximum of the tyrosine (274 nm) and tryptophan (278 nm) absorption spectra, and at 290 nm where only tryptophan absorbs. Increasing the concentration of phosphate in the solution quenches or enhances the intrinsic fluorescence of the Y160W mutant (Fig. 3), and the effects strongly depend on the excitation wavelength and the region of the fluorescence spectrum. There are no isosbestic points, indicating the presence of more than two species (Fig. 3).
Differential fluorescence spectra of the E. coli PNP complexes with phosphate, left upper panel for the WT PNP (red), remaining graphs for the Y160W mutant (green), obtained at 25 °C, in 50 mM Tris/HCl buffer pH 7.6. The excitation wavelengths at which the spectra were measured are denoted on each graph. Phosphate concentrations were as follows: 10 μM, 100 μM, 1000 μM, 10 000 μM. The increase in phosphate concentration is marked with the gradient of the colour, black for the lowest, red (for the WT) or green (for the Y160W mutant) for the highest phosphate concentration. In the graphs for the Y160W mutant, the wavelengths chosen for the observation for each excitation wavelengths are marked by arrows. These data, together with the excitation wavelength shown on the graph, define conditions, in which titrations presented on Fig. 4, were performed. Direction of the arrows indicates only the direction of the phosphate concentration growth.
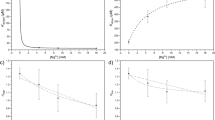
Based on the analysis of the differential spectra obtained for the Y160W mutant (Fig. 3) eight combinations of excitation and observation wavelengths were selected for titration measurements, in order to obtain curves with the greatest possible total change in fluorescence, and as many various titration curve shapes as possible. The typical series of titrations for both WT protein and Y160W mutant are shown in Figure 4. Even visual analysis of the curves obtained for the mutant suggests that binding is at least a three-step process, as some of the curves have more than one extremum (in the case of 290 nm/360 nm and 274 nm/365 nm excitation/observation wavelength, respectively). Curves were analysed globally as described in “Methods” section. Details of a discrimination analysis of the tested models are depicted in Table S1 in the Supplementary Information. For both WT protein and Y160W mutant three-binding-site model describes the data best, and values of dissociation constants obtained are shown in Table 3. As with kinetic parameters (Tables 1 and 2), constants for the Y160W mutant are slightly higher than for the WT protein, although within the 3σ test, Kd1 and Kd3 constants for both proteins are consistent, only the difference between constants Kd2 exceeds the condition of compliance of this test. This may mean that introducing of tryptophan at position 160 has a not negligible effect on the ligand binding process described by the Kd2 constant. Most importantly, however, the model describing the complex formation process for both proteins is the same, i.e. three binding sites with different affinity for the ligand are observed.
Fluorescence titration curves of the enzyme with phosphate: WT (left panel) and Y160W mutant (right panel). Data (points), global best fit model (lines) and residual plots are presented. Titrations were performed at 25 °C, in 50 mM Tris/HCl buffer pH 7.6 and in following excitation/observation wavelengths for the WT enzyme: 270/310 nm (brown), 278/312 nm (orange), 277/312 nm (green), 270/315 nm (violet), 270/320 nm (red), 270/325 nm (grey), 278/325 (blue); and for the Y160W mutant: 278/346 nm (brown), 290/360 nm (orange), 274/365 nm (green), 274/382 nm (violet), 278/334 nm (red), 278/339 nm (grey), 278/363 nm (blue), 290/374 (black).
Binding of phosphate from MST and CD titrations
To confirm that there are three different types of binding sites, a series of thermophoretic (MST), calorimetric (ITC) and circular dichroism (CD) experiments were performed. The shapes of the binding curves obtained from MST titrations (Fig. 5 and Figure S2) are similar for WT PNP and Y160W mutant. Two-binding-site model fits the best to experimental binding curves (see details of the discrimination analysis, Table S1 in the Supplementary Information). The obtained values of the dissociation constants are shown in Table 3. Comparing these values with those obtained by fluorescence titrations, shows that dissociation constants determined by MST reproduce the strongest (Kd1) and the weakest (Kd3) constants obtained from fluorescence titrations analysis. Kd2 constant is either undetectable by MST or is an artefact observed in fluorescence titrations.
Thermophoretic titration curves of the enzyme with phosphate (left panel), CD titration curves of the enzyme with phosphate (middle panel), calorimetric titration curves of enzyme-phosphate complex with formycin A (right panel). On each panel data for the WT enzyme (violet) and the Y160W mutant (green) are presented: data (points), global best fit model (lines) and residual plots below. For the sake of clarity the number of data has been limited to one curve from each titration series. Complete data are shown in the Supplementary Information. Titrations were performed at 25 °C, in 50 mM Tris/HCl buffer pH 7.6. For each method and each protein variant the curve with the largest change of the measured physical property was chosen.
To address this problem, calorimetric titrations (ITC) were employed, but due to the poor quality and low repeatability of results obtained, we decided to change the approach and perform CD titration instead. The near-UV CD spectra (230–300 nm) were collected to follow phosphate binding to the WT PNP and the Y160W mutant (Fig. 5 and Figure S4). The differential spectra of protein-phosphate complex vs. protein have the same shape for the WT protein and Y160W mutant, however introducing Trp at position 160 markedly changes the protein CD spectra, and the overall signal change is larger for the Y160W mutant (Figure S3). Titration curves were obtained by cross-secting the spectra in the wavelengths where the largest changes of the signal occur (see “Methods” section). The titration curves for both protein variants are best described by the three-binding-site model (see “Methods” section, Table S1 in the Supplementary Information for the details of a discrimination analysis). The values of dissociation constants derived from fitting this model are shown in Table 3.
Interactions of E. coli PNP with phosphate
The results of titrations performed with the three techniques described above, i.e., fluorescence, thermophoresis and circular dichroism, show great similarity (Table 3). The values of the smallest dissociation constant (describing the strongest binding) determined by all methods are consistent with each other in the 3σ test. The same is true for the weakest constants. The third constant of 80–200 μM is unequivocally revealed by two methods, fluorescence and CD titrations. This suggests that in MST experiments this constant is simply undetectable. Possible reasons will be addressed in “Discussion” section. The results presented above indicate that the interaction of E. coli PNP with phosphate in solution requires three binding constants for the correct description, which is consistent with the crystallographic structure of the enzyme with phosphate (PDB 3OOH13). In this structure, the three-fold symmetry observed for the apo enzyme is broken, the active site pocket is closed in only two subunits, and as a result of this conformational change, the three dimers that build up the enzyme are no longer structurally equivalent.
Binding of nucleoside analogue—ITC and MST titration studies
Binding studies of nucleoside by the WT PNP and the Y160W mutant were performed using the inhibitor, formycin A (FA, see Fig. 2, inset), a structural analogue of adenosine. Formycin A and formycin B, the analogues of the natural PNP substrates, adenosine and inosine, respectively, show strong absorption, and their fluorescence spectra overlap with that of the protein. Thus, fluorimetric studies employing natural emission of the protein and/or of the ligand, even at moderate ligand concentration, are impossible. Surprisingly, no changes in the CD signal were observed upon formycin A binding by either the WT or Y160W enzyme forms. Therefore, MST experiments and additionally isothermal calorimetric titrations were performed to characterize the binding of formycin A by PNP.
Calorimetric titrations were performed in series for the WT protein and for the Y160W mutant, and the enzyme concentration range used was 70–1300 μM (Fig. 5 and Figure S5). Global analysis of each titration series, for both proteins, were carried out in the Sedphat program20,21. Discrimination analysis (see “Methods” section and Table S1) in both cases pointed to the model with three binding sites as best describing titration data, and the values of dissociation constants derived from fitting of this equation are given in Table 3. In the case of the WT protein the errors of fitted parameters are relatively high, thus the values should be treated as estimates. In the case of thermophoretic measurements, even under the best conditions, the overall signal changes upon binding of formycin A molecules to the WT protein are relatively small, hence the analysis was not straightforward. The titration curves (Fig. 6) for both protein variants are clearly multiphase, indicating that the one-binding-site model is not consistent with the data. In the case of the WT protein, analysis of residual plots of the two-binding-site model suggests a more complex model (probability from Wald-Wolfowitz run test22 is 0.13, number of runs is below expected value). However, fitting the three-site-model returns the Kd1 ~ 10–8 μM, while Kd2 and Kd3 are similar to those obtained from the two-binding-site model. The probability from the Wald-Wolfowitz run test is also very low (0.002) and the number of runs exceeds the expected value. The Akaike criterion23 indicates the two-binding-site model as the best (with Akaike weight equals 0.904). Thus, the two-binding-site model was chosen as best describing data for the WT protein. The values of dissociation constants are shown in Table 3. The analysis in the case of the Y160W mutant was unambiguous since the overall change in thermophoretic signal was significant (Fig. 6). All criteria point to the three-binding-site model as best describing the data obtained. The values of dissociation constants obtained by fitting this model are shown in Table 3.
Thermophoretic titration curves of PNP-Pi complexes with formycin A: WT (left panel) and Y160W (right panel). Data (points), global best fit model (lines) and residual plots are presented. Titrations were performed at 25 °C, in 50 mM Tris/HCl buffer pH 7.6 and with the relative heating laser power at 80% for both WT protein and Y160W mutant. Seven repetitions of each titration were measured.
Our recently published crystal structures of the WT PNP ternary complexes with phosphate and formycin A (PDB ID 4TS9, 4TS319) show steps in nucleoside binding to the subunits of the enzyme hexamer. Crystallisation of the WT PNP complex with phosphate (at saturating concentration) and increasing concentration of formycin A (structure 4TS3) allowed to establish that in the first step, this inhibitor binds only in two closed active sites, which are located in adjacent subunits, but not belonging to the same dimer (as shown in Fig. 1, middle panel for the binary complex enzyme-phosphate). The position of formycin A in the active site of the Y160W mutant, in the structure presented in this study is virtually the same as in the active site of the WT protein (Fig. 2). In addition, the nucleoside is bound in only two active sites of the hexamer, in adjacent subunits but not belonging to the same dimer, exactly as in the partially occupied by formycin WT PNP structure 4TS3. Moreover, even in these active sites the ligand is not bound in all protein molecules, since the occupancy of the formycin A molecule was found to be 0.71. In the remaining active sites, faint electron density blobs are visible in the place usually occupied by a formycin molecule, suggesting that in a small percentage of the Y160W protein molecules the nucleoside may be bound in all active sites, as in the 4TS9 structure of the WT enzyme.
Discussion
Early studies of hexameric E. coli PNP interactions with ligands, carried out over relatively narrow ligand concentration range, indicated that the kinetic and binding properties of this enzyme are well described by the set of two kinetic constants, and two binding constants, respectively11,12. The first crystal structure of the enzyme/ligand ternary complex16 revealed the presence of two conformations of the active sites, open and closed, in line with the results obtained for ligand binding and catalysis. However, in this structure only half of the hexamer was present in the asymmetric unit, hence the detailed analysis of the symmetry of the entire hexamer was not possible. The next published crystal structure9 of the E. coli PNP ternary complex, which has a full hexamer in the asymmetric unit, showed three similar dimers, each with one closed and one open active site. Thus, the architecture of the whole enzyme turned out to be a trimer of dimers, which seemed to be consistent with results obtained in solution11,12.
However, further crystallographic studies revealed that the latter structure9 was the only one with such symmetry, namely with all three dimers in the “open-closed” conformation. Subsequent X-ray structures of the WT E. coli PNP complexed with various ligands, when one of the ligands is phosphate13,17,18,19, do not show a trimer of dimers architecture, with the three-fold symmetry axis, but rather a dimer of trimers, having two-fold symmetry, as only two of the dimers are in “open-closed” conformation. In the third dimer both active sites remain open, breaking the three-fold symmetry. However, slight differences in the structure of open sites forming the “open-closed” dimers and open sites in the “open-open” dimer13,19 suggested, that there are three types of active sites, not just two. This discrepancy between the structural studies and the results obtained in solution was a rationale for undertaking experiments presented in this report.
Interaction of E. coli PNP with phosphate, investigated by fluorescence titrations over a broad range of ligand concentration yielded binding curves that are best described by a model with three dissociation constants, consistent with the symmetry of the enzyme molecule observed in the crystal. The engineering of a single tryptophan Y160W mutant, binding studies exploring also physical properties other than the intrinsic fluorescence of the protein and its complexes, as well as crystallisation of the mutant in a complex with substrates analogues were used to further investigate whether this type of symmetry actually exists when the protein is in solution.
Binding of phosphate was followed by fluorescence, circular dichroism and thermophoretic titrations. In the case of two former methods, for both WT PNP and Y160W mutant titration curves are best described by a model of three binding sites, while a thermophoretic titrations are best described by just two-binding-site model (Table 3). Comparing the values of the dissociation constants shows that the constants describing the strongest binding obtained by all three selected techniques, are consistent with each other (under the 3σ test). The same is true for the largest dissociation constant, describing the weakest binding. Two conformations of active sites, open and closed, with elongated and segmented helix H8, respectively, are visible in the crystallographic structures (Fig. 2), and at the same time opposite changes in the thermophoretic mobility are observed upon binding of phosphate to sites in the closed and in the open conformations, as depicted on Fig. 5, left panel and on Figure S2. Thus, the thermophoretic signal for the middle constant may simply be hidden between these two signals of opposite sign, which may account for the fact that the “middle” constant, Kd2, is not detected by the MST. However, there is also another explanation. MST is a technique that relays on changes in protein structure, significant enough to influence the shape and hydration shell of the molecule, and thus the thermophoretic mobility. It is possible that the processes occurring in the protein molecule due to binding described by this constant just do not significantly affect these parameters. Fortunately, phosphate binding is associated with the significant arrest of the molecular motion, observed in the protein backbone fragments that are flexible in the ligand-free state, what was detected in the proton/deuter exchange experiments24. Such changes in the protein chain mobility are reflected in the intrinsic protein fluorescence and in the aromatic amino acids surrounding phosphate, giving rise to the CD signal for all three types of binding sites.
The next thing was to analyze binding of the nucleoside substrate. Calorimetric studies of binding by both WT protein and the Y160W mutant showed that three-binding-site model fits to the experimental titration curves the best. In the case of thermophoretic titrations, the two-binding-site model and the three-binding-site model better describe the data for the WT PNP and Y160W mutant, respectively. This difference for the two enzyme variants should rather be attributed to the better quality of titration curves obtained for the mutant, rather than to the actual difference in the binding model, since the total change in thermophoresis signal upon ligand binding is much larger and therefore the signal to noise ratio is better yielding the more “featured” curves (Fig. 6). Hence, three-binding-site model describing complex formation with the nucleoside seems valid for both protein variants.
The structure of the ternary complex of the Y160W mutant with sulfate and formycin A described in this study has a nucleoside bound only in two active sites per hexamer, located in the neighbouring subunits, but not belonging to the same dimer. Since phosphate was not present during the crystallisation, all active sites are in the open conformation. However, the distribution of occupied active sites makes this structure resemble the structure of the WT enzyme obtained at not saturating concentration of formycin A, captured as the first step in binding of this ligand by crystallographic “snapshots”. In the latter structure there is an analogous distribution of the occupied (and closed) active sites19. The distribution of the nucleoside molecules in the Y160W enzyme subunits indicates that although all active sites are in the same open conformation, they do not share the same affinity to the ligand and are therefore not equivalent. Three binding constants necessary to describe the thermophoretic titration curves and similarity of the curve describing the change in thermophoretic signal for both protein variants indicate that in the saturating phosphate concentration in crystallographic structures of the Y160W mutant, some active site pockets will most probably be closed, similarly as in the case of the WT enzyme.
The Y160W mutant turned out to be an excellent tool supporting the investigation of the interactions of hexameric E. coli PNP with ligands. The mutant not only yielded the expected more informative intrinsic protein fluorescence curves but also, unexpectedly, the calorimetric and thermophoretic titrations curves with a greater overall change in the measured parameter. The better signal-to-noise ratio and better reproducibility resulted in higher reliability of the obtained data. Using this tool, we have shown that breaking the three-fold symmetry, which is characteristic of most of the crystal structures of the hexameric E. coli PNP complexes with various ligands, is not a crystallographic artefact. Consistent with the two-fold rather than the three-fold symmetry of the overall enzyme architecture, this study indicates that the active sites in the open conformation of the “open-open” dimer most likely differ from the sites in the open conformation of the “open-closed” dimer, because three binding constants, not just two, are necessary to properly describe the interaction of E. coli PNP with both its substrates, the phosphate and the nucleoside (Fig. 7). Thus, also in solution, the three dimers making up the hexameric architecture of this enzyme are most likely not functionally equivalent. Two of them, with open and closed conformations of active sites, are catalytically active, while the third, “open-open” dimer is able to bind ligands but cannot undergo conformational change, consisting in closing of one active site, which is a necessary step in catalysis.
The schematic view of the E. coli PNP hexameric molecule, in the apo form, with the three-fold and the two-fold symmetry axes (left, based on PDB 1ECP8), and in the complexes with ligands, phosphate and nucleoside (right). In just one ternary complex of the E. coli PNP with its ligands (PDB 1K9S)9 the three-fold symmetry is retained (upper right), and all three dimers are with one monomer in the open (blue) and one in the closed (green) conformation of the active sites. The binary enzyme-phosphate complex, and the majority of various ternary enzyme-phosphate-nucleoside complexes show only two-fold symmetry (lower right, see also Fig. 1), as one of three dimers is symmetric (rotten green), hence different than two other unsymmetrical dimers (blue-green). The present study shows that this is more likely not a crystallographic artefact as also in solution the three-binding-site model is necessary to properly describe interaction of this enzyme with ligands, hence dimers are probably also functionally not equivalent.
In this context it is worth mentioning once again that there is only one structure of the ternary complex of E. coli PNP with its ligands, in which the three-fold symmetry is maintained, and all three dimers occur with one monomer in the open and one in the closed conformation of the active sites9. The results described here, together with the presence of the aforementioned structure9, as well as our previous studies showing that dimers tend to form a hexamer because they are not stable as single molecules15, indicate that the molecular mechanism of E. coli PNP catalysis should take into account not only the cooperation between the two dimer-forming subunits, as indicated by the kinetic properties of this enzyme11 and initial ligand binding studies9,12,13, but also a cross-talk of all six hexamer subunits. The question thus arises whether under optimal conditions this cross-talk can lead to the “open-closed” conformation of all three dimers hence to a fully active enzyme, or the third dimer is doomed to be in the catalytically inactive “open-open” conformation. We tackle this problem in detail in our next study on this fascinating enzyme.
Methods
Chemicals
Formycin A was from Berry & Associates, Inc. (USA), adenosine, inosine, guanosine and xanthine oxidase from bovine milk (1 U/mg) were obtained from Sigma-Aldrich (USA). 7-methylguanosine (m7Guo) was synthesised from guanosine according to Jones’ and Robins' method involving methyl iodide25. This yields the preparation free from sulfate, which as an ion resembling phosphate could bias the results. All other chemicals were from Roth (Germany) or Sigma-Aldrich (USA), and were of the highest available purity. Background contaminant phosphate in buffers and other chemicals used in this study was measured spectrophotometrically using the method involving phosphomolybdate complex reduced by ascorbic acid26.
PNP Y160W mutant
The E. coli strain with the pSE380 plasmid carrying E. coli PNP DeoD gene was a kind gift from Dr. Joanne L. Turnbull. The plasmid was a template in a site-directed mutagenesis performed with the use of Invitrogen GeneTailor Site-Directed Mutagenesis System (Invitrogen, USA) according to manufacturer protocol. Forward primer used for introducing the Y160W point mutation was as follows (the altered codon is underlined): 5′-TCTCCGCTGACCTGTTCTGGTCTCCGGACG-3′. The reverse primer was 5′-AGAACAGGTCAGCGGAGAACAGGTTACCCACC-3′. The synthesis of oligonucleotide primers and the sequencing of the mutated PNP gene were done by Genomed S.A. (Poland).
The E. coli DH5α-T1 cells containing the recombinant pSE380/PNPY160W plasmid were grown in Superbroth medium (2% peptone, 0.2% sodium phosphate (dibasic), 0.1% potassium phosphate (monobasic), 0.8% sodium chloride, 1.5% yeast extract, 0.2% glucose and 50 μg/ml ampicillin) at 37 °C with constant shaking (250 rpm) to OD600 = 0.42 and the culture was induced with 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG). The cells were grown for additional 5 h in the same conditions and harvested by centrifugation for 20 min at 4000×g at 4 °C.
The purification procedure was based on the previously published method27, but the additional chromatography step was added. The cell pellet obtained from 2.5 l of the bacterial culture was re-suspended in 50 ml of homogenizing buffer H (100 mM Tris–HCl, pH 7.6, 2 mM EDTA, 10% glycerol (v/v), 1 mM DTT, 50 mM NaCl, 0.4 mg/ml lysozyme and 0.015 mg/ml PMSF), incubated on ice for 30 min, sonicated and centrifuged for 20 min at 43,000×g at 4 °C. The supernatant was filtered through 4 layers of gauze, supplemented with an equal volume of 100 mM Tris–HCl buffer (pH 7.6) containing 2 mM EDTA, 10% glycerol (v/v) and 1 mM DTT (buffer A) and applied onto Q-Sepharose Fast Flow column (120 ml, GE Healthcare). The column was washed with 0.5 l of buffer A and subsequently with 360 ml of buffer A containing 0.1 M KCl. The enzyme was eluted with a 1.24 l linear gradient of 0.1–0.3 M KCl in buffer A. Fractions were assessed for PNP activity (using the spectrophotometric assay) and for the purity (using the SDS-PAGE). The purest fractions were pooled (210 ml), concentrated in Amicon stirred cell (Millipore) to 24 ml volume and supplemented with an equal volume of 2 M ammonium sulfate. The enzyme was applied onto Phenyl-Sepharose CL-4B (30 ml, GE Healthcare) equilibrated with 50 mM HEPES pH 7.6 containing 1 M ammonium sulfate. The column was washed with 200 ml of the same buffer and subsequently with the same buffer containing 0.8 M and 0.6 M ammonium sulfate (200 ml each). The enzyme was eluted with decreasing step gradient of ammonium sulfate in 50 mM HEPES pH 7.6, from 0.5 M to 0.1 M (100 ml each). The fractions containing PNP protein (named fraction 3 and fraction 4) were pooled separately, concentrated in the Amicon stirred cell, and dialysed against 10 mM HEPES pH 7.0 with two changes of dialysis buffer. The fraction 4, containing 60 mg of protein had slightly higher purity and therefore was used in all experiments described in this study. The protein was divided into small aliquots and stored frozen at − 80 °C until further use.
The recombinant WT enzyme was purified in the same way as described in our earlier publications13,18,19.
Enzyme activity and kinetic studies
All activity measurements and kinetic studies were done in 50 mM Hepes/HCl buffer pH 7.0 at 25 °C as previously described for the WT E. coli PNP and its active-site mutants, using direct spectrophotometric assay for adenosine, guanosine and 7-methylguanosine as substrates, and a coupled assay with xanthine oxidase in the case of inosine13,28,29. Activity measurements were performed for each protein sample, before and after each experiment as a control of the state of the protein.
The classical Michaelis–Menten equation was fitted to the kinetic curves (i.e. initial rate of the reaction, v0, vs. initial substrate concentration, c0):
If the analysis of residual plots suggested that a more complex model was necessary to describe the data properly, a model taking into account allosteric interaction between active sites was applied:
where KM2 = aKM1 and Vmax2 = bVmax1 characterizes a free active site when one binding site is already occupied by the ligands30. Coefficients a and b express the degree of cooperation between the subunits in a dimer, while constants KM1 and Vmax1 characterize the active sites of the enzyme when the neighbouring site is free.
A more complex model was used if there was a statistically significant decrease in the residual sum of squares as judged by the F test at 95% confidence level.
To determine inhibition model and inhibition constant for formycin A, rate of phosphorolysis with m7Guo as a variable substrate, at constant, 50 mM, phosphate concentration, was registered in the absence of the inhibitor and in the presence of two different concentrations of formycin. A global fitting to the whole data set was performed, using various models (competitive, uncompetitive, noncompetitive, mixed). As expected for the substrate analogue, competitive model described the data sufficiently well:
where ci is inhibitor concentration. Kinetic data were analysed using the GraphPad Prism program (Intuitive Software for Science, San Diego, CA, USA).
Analytical ultracentrifugation
Sedimentation velocity experiments were conducted with an Optima XL-I (Beckman-Coulter Inc., Indianapolis, IN, USA) using the An-50Ti and An-60Ti analytical rotors, and double-sector 1.2-cm cells with Epon-charcoal centerpieces and sapphire windows. Partial specific volume of the Y160W mutant (from its amino acid composition), densities and viscosities of buffers were calculated using the Sednterp program31. Protein sample in 50 mM Tris/HCl buffer pH 7.6 and 0.2 M NaCl was equilibrated at 20 °C or at 4 °C, accelerated to 42,000 rpm (142,250 g) at the cell bottom and radial scans of the protein-absorption profile in the cell were measured at 5- or 7-min intervals. For the analysis, the continuous c(s) distribution model incorporated in the Sedfit program32 was used. The sedimentation coefficient s and the standard sedimentation coefficient s020,w were calculated, from which the molecular weight of the species present in the solution was derived, assuming that they have the same friction ratio (obtained as a parameter in the Sedfit program).
Crystallisation, data collection and structure solution
The Y160W protein was crystallised in the 50 mM citrate buffer pH 5.2 and 14% ammonium sulfate (w/v) in the presence of 50 mM phosphate and/or 5 mM formycin A, PNP concentration was 90 mg/ml. Crystals were obtained at 18 °C by the hanging-drop vapour diffusion method. Prior to flash-freezing in liquid nitrogen crystals were soaked in a cryoprotectant solution containing 30% glycerol. Data were collected at the beamline P14 operated by EMBL Hamburg at the PETRA III storage ring (DESY, Hamburg, Germany), images were processed with the XDS33. The structure was solved by the molecular replacement with the Phaser, being a part of CCP4 suite34, and as a model PNP-WT (PDB 1ECP8) was used. The structure was refined with the Phenix.refine35 and Coot36. The X-ray data collection and refinement statistics are shown in Table 4. The structure was deposited in the Protein Data Bank (PDB 6XZ2).
Fluorimetric titrations
All experiments were carried out on the Perkin-Elmer LS55 spectrofluorimeter. Fluorescence spectra were registered using in each case (protein or protein–ligand complex) several excitation wavelengths. Differential spectra were calculated by subtracting spectrum of a protein from the spectrum of the appropriate complex. The negative differential fluorescence intensity means that a decrease of fluorescence upon complex formation is observed. Titrations of the enzyme with phosphate were registered using several excitation and observation wavelengths combinations. When selecting specific wavelengths for such a combination, two criteria were used: (1) the overall change in the fluorescence signal and (2) the complexity of the shape of the titration as the analysis of the multi-phasic curves is more explicit than a uni-phasic curve analysis.
Path length for the excitation was 2 mm and for the emission was 5 mm (half of the cuvette dimensions). A continuous stirring was used during titrations. All measurements were performed at 25 °C. Enzyme concentration (as a monomer) was in the range of 0.5–2 μM. Molecular weight of the Y160W monomer used in all calculations was 25.970 kDa, and the molar extinction coefficient at absorption maximum ε278nm = 8 940 M−1 cm−1, corresponding to ε1%278 nm = 0.34537. Protein and ligand were dissolved in 50 mM Tris/HCl buffer pH 7.6, ligands stocks contained protein at the same concentration as in the reaction cuvette, as proposed by Breer et al.38, therefore, there was no protein dilution during titration and no correction is needed.
Fluorescence intensity of the WT protein is stable over time, but in the case of the Y160W mutant a slight quenching of fluorescence is observed during titration, which typically lasted about 30 min as the time between each point in the titration was 1 min. Furthermore, adding phosphate during titration stabilises the intrinsic fluorescence of the protein. Therefore, for each phosphate concentration, [Pi], used in titrations (each titration point), fluorescence intensity vs. time was measured, and for each of them the linear fit was found to be sufficient to describe the data. Slopes of these lines, a([Pi]), plotted vs. phosphate concentration, give a typical decay curve, showing signal stabilisation upon increasing concentration of phosphate. Thus corrected binding curves, Fcor([Pi]), are given by the following equation, in which Fn([Pi]) is the fluorescence intensity measured for the phosphate concentration [Pi], and n is the number of a given titration point in the particular titration:
At pH 7.6 phosphate is mostly in a form of a double-negative ion (pKa 7.2)39, thus ionic strength, I, of the solution raises significantly during titration with this ligand. To check its influence on the intrinsic protein fluorescence, titrations with NaCl were performed, giving dependence F(I). For the WT protein F(I) was found to be constant up to the ionic strength of about 100 mM, and then a steep decrease is observed. Titration curves of the WT protein with phosphate were therefore measured up to 100 mM ionic strength of solution, which is enough to observe a plateau. For the Y160W mutant a more complex dependence F(I) was observed, influenced also by the titration conditions (excitation and observation wavelengths). The dependence of the fluorescence on the ionic strength observed for the WT protein suggests that there is no specific influence of the increasing ionic strength on the protein intrinsic fluorescence. Therefore, to correct for the increase of the ionic strength caused by phosphate, the fluorescence intensity measured during titration with this ligand, Fobs([Pi]) was divided by F(I), both obtained for the same ionic strength, to get fluorescence intensity curve, Fbind([Pi]) in which changes of the fluorescence observed are caused only by the phosphate biding to the enzyme. Both Fobs([Pi]) and F(I) were corrected for the time dependent fluorescence quenching, according to Eq. (4), prior to the Fbind([Pi]) calculation.
Titrations were performed in series (consisting of five to eight curves), and analysed globally using the DynaFit4 program40 (see below).
Calorimetric titrations
The isothermal titration experiments were performed on the MicroCal iTC200 (Malvern) microcalorimeter. Before each experiment, protein solution was dialysed against 50 mM Tris/HCl buffer pH 7.6, and ligands were dissolved in a dialysate buffer, subsequently all samples were filtered using 0.22 μm pore diameter filters, and degassed. All experiments were carried out at 25°C. For the WT protein series of three calorimetric titrations, with protein concentration in the range of 70 to 800 μM, for the Y160W mutant series of four titration with protein concentration in the range of 300 μM to 1.3 mM were performed. Samples of each protein used for these series came from a single preparation. Ligand samples were 8–12 times more concentrated than the protein sample, in order to achieve a diverse ligand/protein molar ratio during titration and thus better explore binding curves41. In some ITC experiments, in order to sample the lower plateau region more accurately, several initial points were measured with less ligand volume added at each step than used in the subsequent titration points measurements. An example of the ITC data is shown in the Supplementary Information (Figure S6). Raw thermograms were analysed with NITPIC43. Global analysis of titration curves was performed with Sedphat20,21.
MST experiments
Measurements were performed with Monolith NT115 (NanoTemper). Enzyme was labelled with NT647 fluorescent dye (NanoTemper) according to the protocol provided by the manufacturer. Labelled proteins show fluorescence excitation and emission maxima of approximately 650 and 670 nm, respectively. The series of titrations were performed at a constant protein concentration. Both proteins (WT and mutant), and ligands were dissolved in 50 mM Tris/HCl pH 7.6 buffer, measurements were performed at 25 °C. We have proved previously, that PNP-WT is stable over a broad range of temperatures13, thus temperature rise in the focal volume during MST experiment, typically 2–6 °C42 should have no significant influence on the protein activity and its biding capability. In the pre-experiments, it was carefully checked for both proteins, WT and Y160W mutant whether any sign of aggregation or inactivation occurs, and if mutant behaviour is similar to that of the WT enzyme. An example of the raw thermophoretic curves obtained with the relative heating laser power at 60% and 80% are shown in the Supplementary Information (Figure S7).
For experiments of the binary complex formation with phosphate, curves were measured at four different powers of the heating laser. Two to five titrations at each heating power were measured, and curves were averaged for a particular heating power. Global fitting was performed for a series of all four titration curves obtained at four heating laser powers. Protein concentrations in capillaries were 0.14 μM for the WT and 0.21 μM for the Y160W mutant. Ligand concentration was in the range of 3.5 nM—750 mM. The standard sixteen point curves were insufficient to test necessary ligand concentration range, thus measurements for two ligand concentration ranges were performed. Concentrations were selected so that three to four points from each range overlap, allowing to stitch obtained curves into one titration curve, covering entire required ligand concentration range. Measurements have been made for high concentrations of phosphate in order to investigate if for the concentration above about 50 mM, any additional binding constants can be observed. This is an area inaccessible in fluorescence titration due to the quenching of the protein fluorescence at such a high ionic strength of the solution. It was verified that so high ionic strength does not affect the fluorescence of the dye used to label the protein.
Experiments of the nucleoside analogue binding were performed in the presence of saturating concentration of phosphate, i.e. 50 mM. For both, WT PNP and Y160W mutant seven repetitions of each curve were measured, for a single power of the heating laser. The power of the heating laser at which the strongest change of the thermophoretic signal upon ligand binding occurs was selected, namely 80% for both enzyme variants. The raw MST traces, for two highest tested laser powers, 60% and 80%, are presented in the Supplementary information (Figure S7) Protein concentration in the capillary was constant and equal to 0.215 μM and 0.0751 μM for the WT protein and the Y160W mutant, respectively. Series of titrations were analysed globally using the DynaFit4 program40 (see below).
CD titrations
Experiments were performed with Chirascan spectrophotometer (Applied Photophysics), in the near UV region (240–290 nm), thus in the region where aromatic amino acids absorb. Full spectra of the protein/complex were recorded, with 0.2 nm resolution and 2 s averaging per point. Enzyme concentration was in the range of 20–120 μM, the 50 mM Tris/HCl buffer pH 7.6 was used, ligands were dissolved in the same buffer. All measurements were performed at 25 °C. Upon addition of each volume of phosphate solution spectrum was recorded in the range 230–300 nm, in the amino acids absorption region, with 0.25 nm step and 2 s per point signal averaging. Spectra were smoothed with Savitzky-Golay algorithm44 using a fourth degree polynomial and eleven points in the averaging window. The wavelengths of the greatest signal change were determined by calculating differential spectra. These wavelengths were used to plot the titration curves. Global analysis of five to seven curves was performed with DynaFit440.
Global analysis of binding curves
Three models were taken into account in analysis of binding curves: model with one (first equation below), two (first two equations) and three binding sites.
The model with one binding site is based on the assumption that all monomers are equivalent, and there is no cooperation between them. In the model with two binding sites the basic functional unit of the protein is a dimer, possible non-equivalence of binding sites or cooperativity between active sites in the dimer are taken into account (in the case of no cooperativity dissociation constants should be related statistically: Kd2 = 4Kd1), the dimers within hexamer are equivalent. The model with three binding sites is based on the crystallographic structures of binary and ternary complexes of PNP with ligands, where two of the dimers are in the “open-closed” conformation, but the third dimer is symmetrical, in the “open-open” conformation. Thus, active sites in the open conformation belonging to “open-closed” and “open-open” dimer may not be equivalent. This model takes into account possibility of cooperation between all monomers within a hexameric molecule of the protein (in the case of no cooperativity dissociation constants should be related statistically: Kd3 = 9Kd1, Kd2 = 3Kd1).
Titration series were analysed globally, always starting from the simplest, one-binding-site model. If the residual plots, judged visually and by Runs of Sign Test/Wald-Wolfowitz Runs Test (WWtest)22, suggested that one-binding-site model does not describe data well, the two-binding-site model was tested. Only if the analysis of the residual plots for the two-binding-site model, judged by the WWtest, suggested that still more complex model should be tested, the three-binding-site model was fitted, and the residual plots analysis was performed for the results. For all tested models the Akaike information criterion23 was also calculated. Generally both criteria used in the discrimination analysis were pointing the same model as best describing the data. The only exception was thermophoretic titration of the WT protein with formycin A, and it is described in details in the Results. Analysed models were fitted to the raw data, whereas graphs presented in this study were prepared from the normalised data and the fitted curves. The parameters of the fit are dissociation constants and responses of the system. Responses are defined as changes in a measured physical quantity related to the formation of subsequent protein–ligand complexes: intrinsic protein fluorescence intensity, fluorescence intensity of the dye, heat and ellipticity for fluorimetric, thermophoretic, calorimetric and CD titrations, respectively. Responses are normalised for the protein concentration present in the titration, thus giving molar coefficients.
Many titration curves obtained in this study are uniphasic. In the Supplementary Information (Figure S8) an example of such a uniphasic titration curve is shown (WT PNP titration with Pi, followed by the near-UV CD detection), with one-, two- and three-binding site models fitted. Respective residual plots are also included. Although the signal changes monotonically it is clear even on the basis of the visual inspection that the one-binding site model should be rejected. In turn, the residual plots indicate that the two-binding site model should be rejected as well.
Accession number
Structural data are available in PDB database under accession number PDB 6XZ2.
Abbreviations
- PNP:
-
Purine nucleoside phosphorylase
- Y160W:
-
PNP with Trp at position 160
- WT:
-
Wild type
- Pi:
-
Orthophosphate
- Ino:
-
Inosine
- Guo:
-
Guanosine
- m7Guo:
-
7-Methylguanosine
- Ado:
-
Adenosine
- FA:
-
Formycin A
- Ki :
-
Inhibition constant
- Km :
-
Michaelis constant
- Vmax :
-
Maximal velocity
References
Bzowska, A., Kulikowska, E. & Shugar, D. Purine nucleoside phosphorylases: properties, functions, and clinical aspects. Pharmacol. Ther. 88, 349–452 (2000).
Pugmire, M. J. & Ealick, S. E. Structural analyses reveal two distinct families of nucleoside phosphorylases. Biochem J. 361, 1–25 (2002).
Sattui, S. E. & Gaffo, A. L. Treatment of hyperuricemia in gout: current therapeutic options, latest developments and clinical implications. Ther. Adv. Musculoskelet. Dis. 8, 149–149 (2016).
Bennett, E. M. et al. Designer gene therapy using an Escherichia coli purine nucleoside phosphorylase/prodrug system. Chem. Biol. 10, 1173–1181 (2003).
Zhou, J. H., Tang, B., Liu, X. L., He, D. W. & Yang, D. T. hTERT-targeted E. coli purine nucleoside phosphorylase gene/6-methylpurine deoxyribose therapy for pancreatic cancer. Chin. Med. J. 120, 1348–1352 (2007).
Hassan, A. E., Abou-Elkhair, R. A., Parker, W. B. & Allan, P. W. 6-Methylpurine derived sugar modified nucleosides: Synthesis and in vivo antitumor activity in D54 tumor expressing M64V-Escherichia coli purine nucleoside phosphorylase. Eur. J. Med. Chem. 108, 616–622 (2016).
Cacciapuoti, G. et al. Efficient fludarabine-activating PNP from archaea as a guidance for redesign the active site of E. Coli PNP. J. Cell. Biochem. 117, 1126–1135 (2016).
Mao, C. et al. The crystal structure of Escherichia coli purine nucleoside phosphorylase: A comparison with the human enzyme reveals a conserved topology. Structure 5, 1373–1383 (1997).
Koellner, G. et al. Open and closed conformation of the E. coli purine nucleoside phosphorylase active center and implications for the catalytic mechanism. J. Mol. Biol. 315, 351–371 (2002).
Štefanić, Z. et al. Still a long way to fully understanding the molecular mechanism of Escherichia coli purine nucleoside phosphorylase. Croat. Chem. Acta 86, 117–127 (2013).
Jensen, K. F. Purine nucleoside phosphorylase from Salmonella typhimurium and Escherichia coli Initial velocity kinetics, ligand binding, reaction mechanism. Eur. J. Biochem. 61, 377–386 (1976).
Kierdaszuk, B., Modrak-Wójcik, A. & Shugar, D. Binding of phosphate and sulfate anions by purine nucleoside phosphorylase from E. Coli: ligand-dependent quenching of enzyme intrisic fluorescence. Biophys. Chem. 63, 107–118 (1997).
Mikleušević, G. et al. Validation of the catalytic mechanism of Escherichia coli purine nucleoside phosphorylase by structural and kinetic studies. Biochemie 93, 1610–1622 (2011).
Wielgus-Kutrowska, B., Grycuk, T. & Bzowska, A. Part-of-the-sites binding and reactivity in the homooligomeric enzymes—Facts and artifacts. Arch. Biochem. Biophys. 642, 31–45 (2018).
Bertosa, B. et al. Homooligomerization is needed for stability: a molecular modelling and solution study of Escherichia coli purine nucleoside phosphorylase. FEBS J. 281, 1860–1871 (2014).
Koellner, G., Luić, M., Shugar, D., Saenger, W. & Bzowska, A. Crystal structure of the ternary complex of E. coli purine nucleoside phosphorylase with formycin B, a structural analogue of the substrate inosine, and phosphate (Sulphate) at 2.1 A resolution. J. Mol. Biol. 280, 153–166 (1998).
Bennet, E. M., Li, C., Allan, P. W., Parker, W. B. & Ealick, S. E. Structural basis for substrate specificity of Escherichia coli purine nucleoside phosphorylase. J. Biol. Chem. 278, 47110–47118 (2003).
Štefanić, Z. et al. New phosphate binding sites in the crystal structure of Escherichia coli purine nucleoside phosphorylase complexed with phosphate and formycin A. FEBS Lett. 586, 967–971 (2012).
Štefanić, Z. et al. Crystallographic snapshots of ligand binding to hexameric purine nucleoside phosphorylase and kinetic studies give insight into the mechanism of catalysis. Sci. Rep. 8, 15427 (2018).
Zhao, H., Piszczek, G. & Schuck, P. SEDPHAT—A platform for global ITC analysis and global multi-method analysis of molecular interactions. Methods 76, 137–148 (2015).
Houtman, J. C. D. et al. Studying multi-site binary and ternary protein interactions by global analysis of isothermal titration calorimetry data in SEDPHAT: Application to adaptor protein complexes in cell signaling. Prot. Sci. 16, 30–42 (2007).
Wald, A. & Wolfowitz, J. On a test whether two samples are from the same population. Ann. Math. Statist. 11, 147–162 (1940).
Burnham, K. P. & Anderson, D. R. Model Selection and Multimodel Inference (Springer, 2002).
Kazazić, S. et al. New insights into active site conformation dynamics of E. coli PNP revealed by combined H/D exchange approach and molecular dynamic simulations. J. Am. Soc. Mass. Spectrom. 27, 73–82 (2015).
Jones, J. W. & Robins, R. K. Purine nucleosides. III. Methylation studies of certain naturally ocurring purine nucleosides. J. Am. Chem. Soc. 85, 193–221 (1963).
Ames, B. N. Assay for inorganic phosphate, total phosphate and phosphonates. Meth. Enzymol. 8, 115–118 (1966).
Lee, J. et al. Expression, purification, and characterization of recombinant purine nucleoside phosphorylase from Escherichia coli. Protein. Expr. Purif. 22, 180–188 (2001).
Kalckar, H. M. Differential spectrophotometry of purine compounds by means of specific enzymes. J. Biol. Chem. 167, 429–443 (1947).
Kulikowska, E., Bzowska, A., Wierzchowski, J. & Shugar, D. Properties of two unusual, and fluorescent, substrates of purine-nucleoside phosphorylase: 7-methylguanosine and 7-methylinosine. Biochim. Biophys. Acta. 874, 355–363 (1986).
Segel, I. H. Enzyme Kinetics: Behavior and Analysis of Rapid Equilibrium and Steady-State Enzyme Systems (Wiley-Interscience, 1993).
Laue, T. M. et al. Computer-aided interpretation of sedimentation data for proteins. In Analytical Ultracentrifugation in Biochemistry and Polymer Science (eds. Harding S. E., Rowe, A. J. & Horton, J. C.) 90–125 (Royal Society of Chemistry, Cambridge 1992).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Kabsch, W. XDS. A. Cryst. D 66, 125–132 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. A. Cryst. D 68, 352–367 (2012).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. A. Cryst. D 60, 2126–2132 (2004).
Gasteiger, E. et al. Protein identification and analysis tools on the ExPASy server. In The Proteomics Protocols Handbook (ed. Walker, J. M.) 571–607 (Humana Press, 2005).
Breer, K. et al. 9-Deazaguanine derivatives connected by a linker to difluoromethylene phosphonic acid are slow-binding picomolar inhibitors of trimeric purine nucleoside phosphorylase. FEBS J. 277, 1747–1760 (2010).
CRC Handbook for Chemistry and Physics. (100th Edition, CRC Press, 2019).
Kuzmic, P. Program DYNAFIT for the analysis of enzyme kinetic data: Application to HIV proteinase. Anal. Biochem. 237, 260–273 (1996).
Brautigan, C. A. Fitting two- and three-site binding models to isothermal titration calorimetric data. Methods 76, 124–136 (2015).
Jerabek-Willemsen, M. et al. MicroScale Thermophoresis: Interaction analysis and beyond. J. Mol. Str. 1077, 101–133 (2014).
Keller, S. et al. High-precision isothermal titration calorimetry with automated peak-shape analysis. Anal. Biochem. 84, 5066–5073 (2012).
Savitzky, A. & Golay, M. J. E. Smoothing and differentiation of data by simplified least squares procedures. Anal. Chem. 36, 1627–1639 (1964).
Acknowledgements
Plasmid pSE380 containing the deoD gene (pSE380deoD) was a gift from Joanne L. Turnbull, Concordia University, Montreal, Canada. We thank Alicja Dyzma for crystallising Y160W mutant, Elżbieta Nowak for collecting X-ray data at the synchrotron, Biserka Žinić and Jacek Jemielity for the synthesis of 7-methylguanosine, Lucyna Magnowska and Natasza Gajda for the excellent technical assistance in kinetic experiments. These studies were supported by the National Science Centre of Poland and Polish Ministry of Science and Higher Education (Grants No. N N301 044939, BST - 163500/BF, BST - 173300/BF task 34, BST-0180100-03/BF task 22, 501-D111-01-1110105 (PP/BF)), and also by the Faculty of Chemistry, Warsaw University of Technology. This work was partially performed in the NanoFun laboratories co-financed by the European Regional Development Fund within the Innovation Economy Operational Program, the Project No. POIG.02.02.00-00-025/09/ and POIG.02.01.00-14-122/09.
Author information
Authors and Affiliations
Contributions
A.B. designed and supervised the study; A.G. and J.C. prepared the Y160W genetic construct and all protein samples; M.N. and A.B. done kinetic and inhibition characterization of the Y160W mutant; BWK did ultracentrifugation experiments; M.N. designed, conducted and analysed majority of titrations, and did crystallographic studies; L.M. designed, conducted and analysed fluorescence titrations of the Y160W mutant; M.W.S. supervised MST titrations and participated in writing the paper; A.O. conducted titrations with the near UV CD detection; M.N. and A.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Narczyk, M., Mioduszewski, Ł., Oksiejuk, A. et al. Single tryptophan Y160W mutant of homooligomeric E. coli purine nucleoside phosphorylase implies that dimers forming the hexamer are functionally not equivalent. Sci Rep 11, 11144 (2021). https://doi.org/10.1038/s41598-021-90472-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-90472-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.