Abstract
Activating transcription factor (ATF)3 is known to have an anti-inflammatory function, yet the role of hepatic ATF3 in lipoprotein metabolism or atherosclerosis remains unknown. Here we show that overexpression of human ATF3 in hepatocytes reduces the development of atherosclerosis in Western-diet-fed Ldlr−/− or Apoe−/− mice, whereas hepatocyte-specific ablation of Atf3 has the opposite effect. We further show that hepatic ATF3 expression is inhibited by hydrocortisone. Mechanistically, hepatocyte ATF3 enhances high-density lipoprotein (HDL) uptake, inhibits intestinal fat and cholesterol absorption and promotes macrophage reverse cholesterol transport by inducing scavenger receptor group B type 1 (SR-BI) and repressing cholesterol 12α-hydroxylase (CYP8B1) in the liver through its interaction with p53 and hepatocyte nuclear factor 4α, respectively. Our data demonstrate that hepatocyte ATF3 is a key regulator of HDL and bile acid metabolism and atherosclerosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. RNA-seq data were deposited in the GEO repository (accession no. GSE148301). The genes regulated by ATF3 are presented in Supplementary Data 1. Source data are provided with this paper.
References
Linton, M. F., Tao, H., Linton, E. F. & Yancey, P. G. SR-BI: a multifunctional receptor in cholesterol homeostasis and atherosclerosis. Trends Endocrinol. Metab. 28, 461–472 (2017).
Cuchel, M. & Rader, D. J. Macrophage reverse cholesterol transport: key to the regression of atherosclerosis? Circulation 113, 2548–2555 (2006).
Rosenson, R. S. et al. Cholesterol efflux and atheroprotection: advancing the concept of reverse cholesterol transport. Circulation 125, 1905–1919 (2012).
Zhang, Y. et al. Hepatic expression of scavenger receptor class B type I (SR-BI) is a positive regulator of macrophage reverse cholesterol transport in vivo. J. Clin. Invest. 115, 2870–2874 (2005).
Braun, A. et al. Loss of SR-BI expression leads to the early onset of occlusive atherosclerotic coronary artery disease, spontaneous myocardial infarctions, severe cardiac dysfunction, and premature death in apolipoprotein E-deficient mice. Circ. Res. 90, 270–276 (2002).
Kozarsky, K. F., Donahee, M. H., Glick, J. M., Krieger, M. & Rader, D. J. Gene transfer and hepatic overexpression of the HDL receptor SR-BI reduces atherosclerosis in the cholesterol-fed LDL receptor-deficient mouse. Arterioscler. Thromb. Vasc. Biol. 20, 721–727 (2000).
Zanoni, P. et al. Rare variant in scavenger receptor BI raises HDL cholesterol and increases risk of coronary heart disease. Science 351, 1166–1171 (2016).
Li-Hawkins, J. et al. Cholic acid mediates negative feedback regulation of bile acid synthesis in mice. J. Clin. Invest. 110, 1191–1200 (2002).
Slatis, K. et al. Abolished synthesis of cholic acid reduces atherosclerotic development in apolipoprotein E knockout mice. J. Lipid Res. 51, 3289–3298 (2010).
Miyake, J. H. et al. Transgenic expression of cholesterol-7-α-hydroxylase prevents atherosclerosis in C57BL/6J mice. Arterioscler. Thromb. Vasc. Biol. 22, 121–126 (2002).
Thompson, M. R., Xu, D. & Williams, B. R. ATF3 transcription factor and its emerging roles in immunity and cancer. J. Mol. Med. 87, 1053–1060 (2009).
Hai, T., Wolford, C. C. & Chang, Y. S. ATF3, a hub of the cellular adaptive-response network, in the pathogenesis of diseases: is modulation of inflammation a unifying component? Gene Expr. 15, 1–11 (2010).
Gilchrist, M. et al. Systems biology approaches identify ATF3 as a negative regulator of Toll-like receptor 4. Nature 441, 173–178 (2006).
Whitmore, M. M. et al. Negative regulation of TLR-signaling pathways by activating transcription factor-3. J. Immunol. 179, 3622–3630 (2007).
De Nardo, D. et al. High-density lipoprotein mediates anti-inflammatory reprogramming of macrophages via the transcriptional regulator ATF3. Nat. Immunol. 15, 152–160 (2014).
Yao, B. C. et al. Chronic stress: a critical risk factor for atherosclerosis. J. Int. Med. Res. 47, 1429–1440 (2019).
Black, P. H. & Garbutt, L. D. Stress, inflammation and cardiovascular disease. J. Psychosom. Res. 52, 1–23 (2002).
Vieweg, W. V. et al. Treatment of depression in patients with coronary heart disease. Am. J. Med. 119, 567–573 (2006).
Rosengren, A. et al. Association of psychosocial risk factors with risk of acute myocardial infarction in 11119 cases and 13648 controls from 52 countries (the INTERHEART study): case–control study. Lancet 364, 953–962 (2004).
Brindley, D. N., McCann, B. S., Niaura, R., Stoney, C. M. & Suarez, E. C. Stress and lipoprotein metabolism: modulators and mechanisms. Metabolism 42, 3–15 (1993).
Catalina-Romero, C. et al. The relationship between job stress and dyslipidemia. Scand. J. Public Health 41, 142–149 (2013).
Djindjic, N., Jovanovic, J., Djindjic, B., Jovanovic, M. & Jovanovic, J. J. Associations between the occupational stress index and hypertension, type 2 diabetes mellitus, and lipid disorders in middle-aged men and women. Ann. Occup. Hyg. 56, 1051–1062 (2012).
Assadi, S. N. What are the effects of psychological stress and physical work on blood lipid profiles? Medicine 96, e6816 (2017).
Heidt, T. et al. Chronic variable stress activates hematopoietic stem cells. Nat. Med. 20, 754–758 (2014).
Russell, G. & Lightman, S. The human stress response. Nat. Rev. Endocrinol. 15, 525–534 (2019).
Troxler, R. G., Sprague, E. A., Albanese, R. A., Fuchs, R. & Thompson, A. J. The association of elevated plasma cortisol and early atherosclerosis as demonstrated by coronary angiography. Atherosclerosis 26, 151–162 (1977).
Dekker, M. J. et al. Salivary cortisol is related to atherosclerosis of carotid arteries. J. Clin. Endocrinol. Metab. 93, 3741–3747 (2008).
Alevizaki, M., Cimponeriu, A., Lekakis, J., Papamichael, C. & Chrousos, G. P. High anticipatory stress plasma cortisol levels and sensitivity to glucocorticoids predict severity of coronary artery disease in subjects undergoing coronary angiography. Metabolism 56, 222–226 (2007).
Neary, N. M. et al. Hypercortisolism is associated with increased coronary arterial atherosclerosis: analysis of noninvasive coronary angiography using multidetector computerized tomography. J. Clin. Endocrinol. Metab. 98, 2045–2052 (2013).
Hermanowski-Vosatka, A. et al. 11β-HSD1 inhibition ameliorates metabolic syndrome and prevents progression of atherosclerosis in mice. J. Exp. Med. 202, 517–527 (2005).
Cholongitas, E. et al. Relative adrenal insufficiency is associated with the clinical outcome in patients with stable decompensated cirrhosis. Ann. Hepatol. 16, 584–590 (2017).
Spadaro, L. et al. Apolipoprotein AI and HDL are reduced in stable cirrhotic patients with adrenal insufficiency: a possible role in glucocorticoid deficiency. Scand. J. Gastroenterol. 50, 347–354 (2015).
Hayashi, R. Glucocorticoid replacement affects serum adiponectin levels and HDL-C in patients with secondary adrenal insufficiency. J. Clin. Endocrinol. Metab 104, 5814–5822 (2019).
Werumeus Buning, J. et al. Downregulation of cholesteryl ester transfer protein by glucocorticoids: a randomised study on HDL. Eur. J. Clin. Invest. 47, 494–503 (2017).
Nilsson, A. G. et al. Long-term safety of once-daily, dual-release hydrocortisone in patients with adrenal insufficiency: a phase 3b, open-label, extension study. Eur. J. Endocrinol. 176, 715–725 (2017).
Guarnotta, V., Ciresi, A., Pillitteri, G. & Giordano, C. Improved insulin sensitivity and secretion in prediabetic patients with adrenal insufficiency on dual-release hydrocortisone treatment: a 36-month retrospective analysis. Clin. Endocrinol. 88, 665–672 (2018).
Yan, C., Lu, D., Hai, T. & Boyd, D. D. Activating transcription factor 3, a stress sensor, activates p53 by blocking its ubiquitination. EMBO J. 24, 2425–2435 (2005).
Reynier, M. O. et al. Comparative effects of cholic, chenodeoxycholic, and ursodeoxycholic acids on micellar solubilization and intestinal absorption of cholesterol. J. Lipid Res. 22, 467–473 (1981).
Tint, G. S., Salen, G. & Shefer, S. Effect of ursodeoxycholic acid and chenodeoxycholic acid on cholesterol and bile acid metabolism. Gastroenterology 91, 1007–1018 (1986).
Lanzini, A. & Northfield, T. C. Effect of ursodeoxycholic acid on biliary lipid coupling and on cholesterol absorption during fasting and eating in subjects with cholesterol gallstones. Gastroenterology 95, 408–416 (1988).
Hardison, W. G. & Grundy, S. M. Effect of ursodeoxycholate and its taurine conjugate on bile acid synthesis and cholesterol absorption. Gastroenterology 87, 130–135 (1984).
Salvioli, G., Lugli, R. & Pradelli, J. M. Cholesterol absorption and sterol balance in normal subjects receiving dietary fiber or ursodeoxycholic acid. Dig. Dis. Sci. 30, 301–307 (1985).
Chevre, R. et al. Therapeutic modulation of the bile acid pool by Cyp8b1 knockdown protects against nonalcoholic fatty liver disease in mice. FASEB J. 32, 3792–3802 (2018).
Bertaggia, E. et al. Cyp8b1 ablation prevents Western diet-induced weight gain and hepatic steatosis because of impaired fat absorption. Am. J. Physiol. Endocrinol. Metab. 313, E121–E133 (2017).
Watanabe, M. et al. Bile acids lower triglyceride levels via a pathway involving FXR, SHP, and SREBP-1c. J. Clin. Invest. 113, 1408–1418 (2004).
Inoue, Y. et al. Regulation of bile acid biosynthesis by hepatocyte nuclear factor 4α. J. Lipid Res. 47, 215–227 (2006).
Jahan, A. & Chiang, J. Y. Cytokine regulation of human sterol 12α-hydroxylase (CYP8B1) gene. Am. J. Physiol. Gastrointest. Liver Physiol. 288, G685–G695 (2005).
Kliewer, S. A. & Mangelsdorf, D. J. Bile acids as hormones: the FXR–FGF15/19 pathway. Dig. Dis. 33, 327–331 (2015).
Davis, R. A., Miyake, J. H., Hui, T. Y. & Spann, N. J. Regulation of cholesterol-7α-hydroxylase: BAREly missing a SHP. J. Lipid Res. 43, 533–543 (2002).
Wang, J. et al. Studies on LXR- and FXR-mediated effects on cholesterol homeostasis in normal and cholic acid-depleted mice. J. Lipid Res. 47, 421–430 (2006).
Peet, D. J. et al. Cholesterol and bile acid metabolism are impaired in mice lacking the nuclear oxysterol receptor LXRα. Cell 93, 693–704 (1998).
Hong, C. et al. The LXR–Idol axis differentially regulates plasma LDL levels in primates and mice. Cell Metab. 20, 910–918 (2014).
Wang, N., Arai, T., Ji, Y., Rinninger, F. & Tall, A. R. Liver-specific overexpression of scavenger receptor BI decreases levels of very low density lipoprotein ApoB, low density lipoprotein ApoB, and high density lipoprotein in transgenic mice. J. Biol. Chem. 273, 32920–32926 (1998).
Bjorklund, M. M. et al. Induction of atherosclerosis in mice and hamsters without germline genetic engineering. Circ. Res. 114, 1684–1689 (2014).
Joseph, J. J. & Golden, S. H. Cortisol dysregulation: the bidirectional link between stress, depression, and type 2 diabetes mellitus. Ann. N. Y. Acad. Sci. 1391, 20–34 (2017).
Huszar, D. et al. Increased LDL cholesterol and atherosclerosis in LDL receptor-deficient mice with attenuated expression of scavenger receptor B1. Arterioscler. Thromb. Vasc. Biol. 20, 1068–1073 (2000).
Arai, T., Wang, N., Bezouevski, M., Welch, C. & Tall, A. R. Decreased atherosclerosis in heterozygous low density lipoprotein receptor-deficient mice expressing the scavenger receptor BI transgene. J. Biol. Chem. 274, 2366–2371 (1999).
Huby, T. et al. Knockdown expression and hepatic deficiency reveal an atheroprotective role for SR-BI in liver and peripheral tissues. J. Clin. Invest. 116, 2767–2776 (2006).
Ji, Y. et al. Hepatic scavenger receptor BI promotes rapid clearance of high density lipoprotein free cholesterol and its transport into bile. J. Biol. Chem. 274, 33398–33402 (1999).
Houten, S. M., Watanabe, M. & Auwerx, J. Endocrine functions of bile acids. EMBO J. 25, 1419–1425 (2006).
Wang, D. Q., Tazuma, S., Cohen, D. E. & Carey, M. C. Feeding natural hydrophilic bile acids inhibits intestinal cholesterol absorption: studies in the gallstone-susceptible mouse. Am. J. Physiol. Gastrointest. Liver Physiol. 285, G494–G502 (2003).
Wang, J. et al. Relative roles of ABCG5/ABCG8 in liver and intestine. J. Lipid Res. 56, 319–330 (2015).
Huang, Y. et al. Lactobacillus acidophilus ATCC 4356 prevents atherosclerosis via inhibition of intestinal cholesterol absorption in apolipoprotein E-knockout mice. Appl. Environ. Microbiol. 80, 7496–7504 (2014).
Davidson, M. H. et al. Inhibition of intestinal cholesterol absorption with ezetimibe increases components of reverse cholesterol transport in humans. Atherosclerosis 230, 322–329 (2013).
Lim, J. H., Lee, H. J., Pak, Y. K., Kim, W. H. & Song, J. Organelle stress-induced activating transcription factor-3 downregulates low-density lipoprotein receptor expression in Sk-Hep1 human liver cells. Biol. Chem. 392, 377–385 (2011).
Xu, Y. et al. A metabolic stress-inducible miR-34a-HNF4α pathway regulates lipid and lipoprotein metabolism. Nat. Commun. 6, 7466 (2015).
Kozarsky, K. F. et al. Overexpression of the HDL receptor SR-BI alters plasma HDL and bile cholesterol levels. Nature 387, 414–417 (1997).
Xu, Y. et al. Lipocalin-2 protects against diet-induced nonalcoholic fatty liver disease by targeting hepatocytes. Hepatol. Commun. 3, 763–775 (2019).
Zhang, Y., Castellani, L. W., Sinal, C. J., Gonzalez, F. J. & Edwards, P. A. Peroxisome proliferator-activated receptor-γ coactivator 1α (PGC-1α) regulates triglyceride metabolism by activation of the nuclear receptor FXR. Genes Dev. 18, 157–169 (2004).
Ge, X. et al. Aldo-keto reductase 1B7 is a target gene of FXR and regulates lipid and glucose homeostasis. J. Lipid Res. 52, 1561–1568 (2011).
Li, Y. et al. Hepatic forkhead box protein A3 regulates ApoA-I (apolipoprotein A-I) expression, cholesterol efflux, and atherogenesis. Arterioscler. Thromb. Vasc. Biol. 39, 1574–1587 (2019).
Voshol, P. J. et al. Postprandial chylomicron formation and fat absorption in multidrug resistance gene 2 P-glycoprotein-deficient mice. Gastroenterology 118, 173–182 (2000).
Goudriaan, J. R. et al. CD36 deficiency in mice impairs lipoprotein lipase-mediated triglyceride clearance. J. Lipid Res. 46, 2175–2181 (2005).
Khalifeh-Soltani, A. et al. Mfge8 promotes obesity by mediating the uptake of dietary fats and serum fatty acids. Nat. Med. 20, 175–183 (2014).
Turley, S. D., Herndon, M. W. & Dietschy, J. M. Reevaluation and application of the dual-isotope plasma ratio method for the measurement of intestinal cholesterol absorption in the hamster. J. Lipid Res. 35, 328–339 (1994).
Xu, Y. Macrophage miR-34a is a key regulator of cholesterol efflux and atherosclerosis. Mol. Ther. 202–216 (2019).
Rossi, S. S., Converse, J. L. & Hofmann, A. F. High pressure liquid chromatographic analysis of conjugated bile acids in human bile: simultaneous resolution of sulfated and unsulfated lithocholyl amidates and the common conjugated bile acids. J. Lipid Res. 28, 589–595 (1987).
Wang, D. Q., Lammert, F., Paigen, B. & Carey, M. C. Phenotypic characterization of Lith genes that determine susceptibility to cholesterol cholelithiasis in inbred mice: pathophysiology of biliary lipid secretion. J. Lipid Res. 40, 2066–2079 (1999).
Naik, S. U. et al. Pharmacological activation of liver X receptors promotes reverse cholesterol transport in vivo. Circulation 113, 90–97 (2006).
Xu, Y. et al. FXR activation increases reverse cholesterol transport by modulating bile acid composition and cholesterol absorption. Hepatology 64, 1072–1085 (2016).
Roberts, D. C. et al. An alternative procedure for incorporating radiolabelled cholesteryl ester into human plasma lipoproteins in vitro. Biochem. J. 226, 319–322 (1985).
Acknowledgements
We thank L. Yang’s group for providing human data. The non-clinical work was supported in part by the National Institutes of Health, grants R01DK102619 (Y. Zhang), R01HL103227 (Y. Zhang and L. Yin), R01HL142086 (Y. Zhang), R01DK118941 (Y. Zhang) and R01DK118805 (Y. Zhang).
Author information
Authors and Affiliations
Contributions
Y.X., Y.L., K.J. and Y. Zhang conceived and designed the study and guided the interpretation of the results. Y.X. performed 80% of the studies and data analysis. K.J. generated floxed Atf3 mice. Y.L. and K.J. performed a number of in vivo studies. Y. Zhang supervised the project. Y. Zhang and Y.X. prepared the manuscript. X.P., Y. Zhu, S.H., S.C. and L. Yin performed various studies. The human data were provided by L.C., Y.T. and L. Yang; H.H.W. and D.Q.-H.W. performed the BA composition assays. All authors discussed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks Alexander Bartelt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Christoph Schmitt.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Hepatic ATF3 is repressed by stress signaling.
a. Mouse or human primary hepatocytes were treated with vehicle (Veh), 500 nM angiotensin II (A-II), 500 nM dexamethasone (DEX), 250 μM forskolin (FSK), 100 nM glucagon (GCG) or 250 nM hydrocortisone (HC) (n = 4). After 24 h, protein levels were determined. **P < 1E-6 for Veh versus A-II, DEX, FSK, GCG or HC, respectively for mouse or human primary hepatocytes. b. HepG2 cells were treated with vehicle, angiotensin II (A-II), dexamethasone (DEX), forskolin (FSK), glucagon (GCG) or hydrocortisone (HC) as described in (a) (n = 4). mRNA levels were determined after 6 h (top panel) and protein levels were determined after 24 h (bottom panel). **P = 0.000019, 3E-6, 1E-6, 0.0031 or 2E-6 for Veh versus A-II, DEX, FSK, GSK or HC, respectively. c–f. Mouse primary hepatocytes (c; n = 4), human primary hepatocytes (d; n = 3 for Veh group and n = 4 for cAMP group) or HepG2 cells (e and f; n = 4) were treated with vehicle (Veh) or 500 μM db-cAMP (cAMP). mRNA levels were analyzed after 6 h (c–e) and protein levels in HepG2 cells were analyzed after 24 h (f). In (c), **P = 0.0034 versus Veh. In (d), **P = 0.000051 versus Veh. In (e), *P = 0.01 versus Veh. g and h. Mouse (g) or human (h) primary hepatocytes were pretreated with 10 μM H89 or 1 μM PKI 14–22 (PKI) for 2 h, followed by treatment with hydrocortisone (HC) for 6 h. mRNA levels were determined (n = 4). In (g), **P < 1E-6 for HC versus Veh, HC + H89 or HC + PKI, respectively. In (h), *P = 0.04 for HC versus Veh, and **P = 0.0061 or 0.0016 for HC versus HC + H89 or HC + PKI, respectively. i. C57BL/6J mice were i.p. injected with either vehicle or hydrocortisone (HC; 2 mg/kg) once a day for 7 days (n = 8). Plasma cholesterol lipoprotein profile was analyzed by FPLC. j–l. Atf3fl/fl mice and hepatocyte-specific Atf3−/− (L-Atf3−/−) mice were i.p. injected with either vehicle or hydrocortisone (HC; 2 mg/kg) once a day for 7 days. Hepatic mRNA levels were determined (j; n = 8 for the L-Atf3−/−+HC group, and n = 7 for 3 other groups). Plasma cholesterol lipoprotein profile was analyzed (k). Plasma ALT (left panel) and ASL (right panel) (l) levels were determined (n = 6 for the L-Atf3−/−+Vehicle group, and n = 7 for 3 other groups). In (j), for Atf3 expression, **P = 0.002 or 0.000072 for Atf3fl/fl + HC versus Atf3fl/fl + Veh or L-Atf3−/−+HC, respectively. For Scarb1 expression, **P = 2E-6 or <1E-6 for Atf3fl/fl + HC versus Atf3fl/fl + Veh or L-Atf3−/−+HC, respectively. For Cyp7a1 expression, **P = 1E-6 or <1E-6 for Atf3fl/fl + HC versus Atf3fl/fl + Veh or L-Atf3−/−+HC, respectively. For Cyp8b1 expression, **P = 0.0014 or 2E-6 for Atf3fl/fl + HC versus Atf3fl/fl + Veh or L-Atf3−/−+HC, respectively. In (l), for ALT, P = 0.94 or 0.89 for vehicle versus HC for Atf3fl/fl or L-Atf3−/− mice, respectively. For AST, P = 0.98 or 0.99 for vehicle versus HC for Atf3fl/fl or L-Atf3−/− mice, respectively. m. C57BL/6J mice were i.p. injected with either vehicle or hydrocortisone (HC; 2 mg/kg) once a day for 7 days (n = 8). The correlation between hepatic Atf3 mRNA and plasma HDL-C levels was determined. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (c-e), one-way (a, b, g, h) or two-way (j, l) ANOVA with Turkey’s post hoc test for multiple comparisons, or a two-tailed Pearson correlation analysis (m) was used for statistical analysis. NS, not significant.
Extended Data Fig. 2 Over-expression of hepatic ATF3 reduces plasma HDL-C levels and bile acid hydrophobicity indices in C57BL/6J mice.
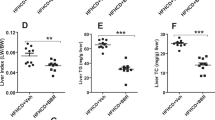
C57BL/6J mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (n = 8). After 2 months, mice were euthanized. a–d. Plasma levels of triglyceride (TG), total cholesterol (TC) (a), HDL-C, non-HDL-C (b), ALT and AST (c) were quantified. Plasma cholesterol lipoprotein profile was analyzed by FPLC (d). Chol, cholesterol. In (a), **P = 0.00024 versus AAV-Null for TC. In (b), **P = 0.00009 versus AAV-Null for HDL. In (c), **P = 0.0024 versus AAV-Null for ALT, and P = 0.079 for AAV-Null versus AAV-ATF3 for AST. e. RNA sequencing was performed using liver samples (n = 4 per group). The volcano plot shows many hepatic genes were differentially regulated by ATF3. f–i. Hepatic mRNA levels were quantified by qRT-PCR (f; n = 8). Hepatic proteins were analyzed by Western blot assays (g, h) and protein levels were quantified (i; n = 6). In (f), **P = 0.000013, 0.00014, 0.00053, 0.00015 or 0.0021 versus AAV-Null for Cyp7a1, Cyp8b1, Scarb1, Ldlr and Apoe expression, respectively. In (i), *P = 0.022 versus AAV-Null for LDLR, and **P = 0.00026, 0.0012, 0.0046, 0.004 or 0.0032 versus AAV-Null for CYP7A1, CYP8B1, ATF3 and ApoE expression, respectively. j,k. Plasma total bile acid (BA) levels (j; n = 8) and biliary BA composition (k; n = 8 for the Null group, and n = 7 for the ATF3 group) were determined. TUDC, tauroursodeoxycholic acid. TCA, taurocholic acid. TCDCA, taurochenodeoxycholic acid. In (j), *P = 0.025 versus Null. In (k), *P = 0.012 and **P = 0.0027 or 0.0006 versus AAV-Null for TCDCA, TUDC or TCA, respectively. l. HepG2 cells were transfected with the pGL3-Cyp7a1 or pGL3-Cyp8b1 luciferase promoters together with pCMV-ATF3 (n = 8) or pCMV-Empty (n = 7). After 36 h, relative luciferase units (RLU) were determined. **P = 0.0063 or 0.00018 versus CMV-Empty for pGL3-Cyp7a1 or pGL3-Cyp8b1, respectively. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (a, c, f, i–l) or two-way ANOVA with Turkey’s post hoc test for multiple comparisons (b) was used for statistical analysis.
Extended Data Fig. 3 Over-expression of hepatic ATF3 reduces plasma HDL-C and LDL-C levels and increases plasma bile acid levels in db/db mice.
db/db mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (n = 8). After 2 months, mice were euthanized. a–d. Plasma total cholesterol (Chol) (a), HDL-C, non-HDL-C (b), ALT and AST (c) levels were determined. Plasma cholesterol lipoprotein profile was analyzed by FPLC (d). In (a), *P = 0.01 versus Null. In (b), **P = 0.0012 or 0.000021 versus AAV-Null for HDL or non-HDL, respectively. In (c), *P = 0.02 or 0.027 versus AAV-Null for ALT or AST, respectively. e. Plasma bile acid (BA) levels. *P = 0.042 versus Null. f and g. Hepatic proteins were analyzed by Western blot assays (f) and protein levels were quantified (g). **P = 3E-6, 8E-6, 0.00074, 7E-6, 0.0039 or 0.0065 versus AAV-Null for CYP7A1, CYP8B1, ATF3, SR-BI, LDLR or ApoE expression, respectively. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (a, c, e, g) or two-way ANOVA with Turkey’s post hoc test for multiple comparisons (b) was used for statistical analysis.
Extended Data Fig. 4 Loss of hepatocyte ATF3 increases plasma HDL-C levels and reduces plasma bile acid levels.
Atf3fl/fl mice and hepatocyte-specific Atf3−/− (L-Atf3−/−) mice were fed a chow diet (n = 6). a–d. Plasma total cholesterol (Chol) (a), HDL, non-HDL-C (b), ALT and AST (c) levels were determined. Plasma cholesterol lipoprotein profile was analyzed by FPLC (d). In (a), **P = 0.0036 versus Atf3fl/fl. In (b), **P = 0.00075 for Atf3fl/fl versus L-Atf3−/− for HDL. In (c), P = 0.49 or 0.28 for Atf3fl/fl versus L-Atf3−/− for ALT or AST, respectively. e. Plasma bile acid (BA) levels. *P = 0.019 versus Atf3fl/fl. f and g. Hepatic proteins were analyzed by Western blot assays (f) and protein levels were quantified (g). **P = 2E-6, 0.000099, 0.000035 or 0.000376 versus Atf3fl/fl for ATF3, CYP7A1, CYP8B1 or SR-BI expression, respectively. h. Hepatic mRNA levels were determined by qRT-PCR. P = 0.12, 0.25 or 0.1 for Atf3fl/fl versus L-Atf3−/− for Srebp2, Hmgcr, or Hmgcs expression, respectively. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (a, c, e, g, h) or two-way ANOVA with Turkey’s post hoc test for multiple comparisons (b) was used for statistical analysis.
Extended Data Fig. 5 ATF3 regulates HDL-C and LDL-C uptake by hepatocytes.
a,b. Scarb1fl/fl (Srb1fl/fl) or hepatocyte-specific Scarb1−/− (L-Srb1−/−) mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (a). In a separate study, Atf3+/+ or Atf3−/− mice were i.v. injected with 0.5 × 109 pfu Ad-Empty or Ad-SR-BI (b). After 10 days, primary hepatocytes were isolated from these mice. HDL uptake was carried out after hepatocytes were treated for 4 h with HDL labeled with [14C]cholesteryl Oleate (14C-HDL) (n = 5) (a, b). In (a), *P = 4E-6 or <1E-6 for Srb1fl/fl + AAV-Null versus Srb1fl/fl + AAV-ATF3 or L-Srb1−/− +AAV-Null, respectively. In (b), **P < 1E-6 for Atf3−/−+Ad-Empty versus Atf3+/++Ad-Empty, Atf3+/++Ad-SR-BI or Atf3−/−+Ad-SR-BI. c,d. Ldlr+/+ mice and Ldlr−/− mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3. After 10 days, primary hepatocytes were isolated. LDL uptake was carried out after hepatocytes were treated for 2 h (c, left panel) or 4 h (c, right panel) with LDL labeled with [14C]cholesteryl Oleate (14C-LDL) (n = 5). Western blot assays were performed (d, left panel) and LDLR protein levels were quantified (d, right panel). In (c, left panel), **P = 0.00002 or <1E-6 for Ldlr+/++AAV-Null versus Ldlr+/++AAV-ATF3 or Ldlr−/− +AAV-Null, respectively. In (c, right panel), *P = 0.000011 or <1E-6 for Ldlr+/++AAV-Null versus Ldlr+/++AAV-ATF3 or Ldlr−/− +AAV-Null, respectively. In (d), *P = 0.0038 versus Null. e. Mouse primary hepatocytes were isolated from Atf3fl/fl or L-Atf3−/− mice. LDL uptake was carried out after hepatocytes were treated for 2 or 4 h with LDL labeled with [14C]cholesteryl Oleate (14C-LDL) (n = 5). P = 0.36 or 0.58 for Atf3+/+versus Atf3−/− at 2 h or 4 h, respectively. f,g. HepG2 cells were infected with Ad-Empty or Ad-ATF3 for 24 h. Western blot assays (f) and LDL uptake (g) were performed as described above (n = 5). In (f), *P = 0.04 versus Empty. In (g), **P = 0.0016 or 0.000013 for Ad-Empty versus Ad-ATF3 at 2 h or 4 h, respectively. h,i. HepG2 cells were infected with Ad-shLacZ or Ad-shATF3 for 24 h. Protein levels were determined by Western blot assays (h, left and right panels). LDL uptake was performed as described above (n = 5) (i). In (h), **P = 0.00003 or 0.0015 versus shLacZ for ATF3 or LDLR expression, respectively. In (i), *P = 0.011 and **P = 0.00003 for shLacZ versus shAtf3 at 2 h or 4 h, respectively. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (d, f, h) or two-way ANOVA with Turkey’s post hoc test for multiple comparisons (a–c, e, g, i) was used for statistical analysis. NS, not significant.
Extended Data Fig. 6 Regulation of SR-BI expression by ATF3 or p53.
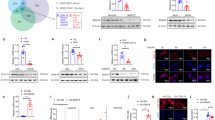
a. C57BL/6J mice were i.v. injected with Ad-Empty or Ad-hP53 (n = 4). After 7 days, hepatic proteins were analyzed by Western blot assays (top panel) and protein levels were quantified (bottom panel). **P = 0.0015 versus Empty. b–d. p53fl/fl mice and hepatocyte-specific p53−/− (L-p53−/−) mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (n = 7 per group). After 30 days, plasma cholesterol lipoprotein profile was analyzed (b). Plasma non-HDL-C (c), ALT and AST (d) levels were quantified. In (c), P = 0.232 or 0.687 for AAV-Null versus AAV-ATF3 for p53fl/fl or L-p53−/− mice, respectively. In (d), for ALT, *P = 0.048 and **P = 0.00025 for AAV-Null versus AAV-ATF3 for p53fl/fl or L-p53−/− mice, respectively; for AST, P = 0.547 or 0.97 for AAV-Null versus AAV-ATF3 for p53fl/fl or L-p53−/− mice, respectively. e,f. p53fl/fl mice and L-p53−/− mice were i.p. injected with either vehicle or hydrocortisone (HC; 2 mg/kg) once a day for 7 days (n = 7 per group). Plasma cholesterol lipoprotein profile (e) and plasma ALT and AST levels (f) were determined. For ALT, P = 0.825 or 0.998 for Vehicle versus HC for p53fl/fl or L-p53−/− mice, respectively. For AST, P = 0.936 or 0.941 for Vehicle versus HC for p53fl/fl or L-p53−/− mice, respectively. g. HepG2 cells were transfected with pGL3-SR-BI-Luc constructs together pCMV-ATF3 or pCMV-Empty (n = 8). After 36 h, relative luciferase units (RLU) were determined. **P = 2E-6, 0.000017, 1.5E-7, 6.9E-8, 4E-7 or 1E-7 for CMV-Empty versus CMV-ATF3 for −0.29, −1.0, −0.8, −0.6, −0.4 or −0.2 kb SR-BI-Luc, respectively. h. EMSA was performed to determine the binding site for ATF3 in the proximal SR-BI promoter. Lane 1: free probe. Lane 2: binding assay for testing if ATF3 protein bound to the SR-BI DNA oligonucleotides. Lanes 3-10: competition assays using SR-BI DNA oligonucleotides containing a wild-type or mutant p53 binding site. Lanes 11 and 12: supershift assays in the presence of IgG or an ATF3 antibody. This experiment was repeated once with similar results. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (a, g) or two-way ANOVA with Turkey’s post hoc test for multiple comparisons (c, d, f) was used for statistical analysis.
Extended Data Fig. 7 Hepatic ATF3 regulates fat absorption and VLDL secretion.
a. C57BL/6J mice were i.v. injected with AAV8-ALB-Null (n = 8) or AAV8-ALB-hATF3 (n = 7 per group). Fat absorption was performed by gavaging mice with [3H]triolein. Plasma radioactivity at indicated time points was measured. **P = 0.0019 for AAV-Null versus AAV-ATF3. b. Fat absorption was performed in Atf3fl/fl mice and L-Atf3−/− mice as described in (a) (n = 8). **P = 0.0011 for Atf3fl/fl versus L-Atf3−/− mice. c. C57BL/6J mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (n = 7). VLDL secretion was performed. **P = 0.0062 for AAV-Null versus AAV-ATF3. d. VLDL secretion was performed in Atf3fl/fl mice and L-Atf3−/− mice (n = 8). **P = 0.000056 for Atf3fl/fl versus L-Atf3−/− mice. All the data are expressed as mean ± SD. All the data points are biological replicates. Two-way ANOVA with Turkey’s post hoc test for multiple comparisons was used for statistical analysis (a-d).
Extended Data Fig. 8 ATF3 induces hepatic LXRα expression and regulates cholesterol or fat absorption independent of FXR signaling.
a. C57BL/6J mice were i.v. injected with AAV8-ALB-Null, AAV8-ALB-ATF3, and/or AAV8-ALB-CYP8B1 (n = 8). After two months, plasma ALT and AST levels were determined. For ALT, *P = 0.042 and **P = 0.001 for AAV-Null versus AAV-ATF3 for CYP8B1 and Null groups, respectively. For AST, P = 0.35 or 0.39 AAV-Null versus AAV-ATF3 for Null or CYP8B1 groups, respectively. b,c. Hnf4αfl/fl mice and L-Hnf4αfl/fl mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (n = 6). After two months, plasma ALT and AST levels (b) and plasma cholesterol lipoprotein profile (c) were determined. In (b), for ALT, *P = 0.011 and **P = 0.0097 for AAV-Null versus AAV-ATF3 for L-Hnf4α−/− and Hnf4αfl/fl mice, respectively. For AST, P = 0.93 or 0.16 for AAV-Null versus AAV-ATF3 for Hnf4αfl/fl or L-Hnf4α−/− mice, respectively. d. HepG2 cells were transfected with pGL3-Cyp8b1-Luc constructs together pCMV-ATF3 or pCMV-Empty (n = 6 per group). After 36 h, relative luciferase units (RLU) were determined. **P < 1E-6 for CMV-Empty versus CMV-ATF3 for −1.1, −0.9, −0.85, −0.75 or −0.66 kb Cyp8b1-Luc, respectively. e. C57BL/6J mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (n = 6 per group). After 2 months, hepatic or intestinal mRNA levels were determined. **P = 0.0096 or 0.0019 for AAV-Null versus AAV-ATF3 for Cyp7a1 or Fgf15, respectively. f. C57BL/6J mice were i.v. injected with AAV8-ALB-Null, AAV8-ALB-hATF3 and/or AAV8-ALB-CYP8B1 (n = 7 for the Null or ATF3 groups, n = 6 for the CYP8B1 group, and n = 8 for the ATF3 + CYP8B1 group). After two months, hepatic mRNA levels were determined. For Lxrα, **P = 1E-6 or 0.00011 for AAV-ATF3 versus AAV-Null or AAV-ATF3 + AAV-CYP8B1, respectively. g,h. Fxr+/+ mice and Fxr−/− mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3 (n = 6). After 2 months, intestinal cholesterol (g) or fat (h) absorption was determined. In (g), *P = 0.033 for Fxr+/++AAV-Null versus Fxr−/−+AAV-Null, and **P = 0.00007 or <1E-6 for AAV-Null versus AAV-ATF3 for Fxr+/+ or Fxr−/− mice, respectively. In (h), **P < 1E-6 for AAV-Null versus AAV-ATF3 for Fxr+/+ or Fxr−/− mice, respectively. The data are expressed as mean ± SEM (a, b, d-g) or mean ± SD (h). All the data points are biological replicates. A two-tailed Student’s t-test (e) and one-way (f) or two-way (a, b, d, g, h) ANOVA with Turkey’s post hoc test for multiple comparisons were used for statistical analysis. NS, not significant.
Extended Data Fig. 9 Over-expression of hepatocyte ATF3 lowers plasma lipid levels in Apoe−/−, Ldlr−/− or Apoe−/−Ldlr−/− mice.
a–h. Three-months-old Apoe−/− mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3, and then fed a Western diet for 3 months. Food intake (n = 15) and body weight (n = 10) were measured (a). Plasma total cholesterol (n = 9 for the Null group, n = 13 for the ATF3 group), LDL-C (n = 8) (b), ApoA-I, ApoB (c; n = 8), cholesterol lipoprotein profile (d), triglyceride (TG) lipoprotein profile (e), and plasma ALT or AST levels (f; n = 7) were determined. Aortic roots were stained using an MOMA-2 antibody (g) and MOMA-2-positive areas were quantified (h; n = 6). In (d), the inset shows HDL fraction. In (a), P = 0.62 or 0.41versus Null for food intake or body weight, respectively. In (b), **P = 0.0002 or 0.0008 versus Null for plasma cholesterol or LDL-C, respectively. In (c), *P = 0.023 or 0.048 versus Null for plasma ApoA-I or ApoB, respectively. In (f), *P = 0.023 versus Null for AST, and **P = 0.0057 versus Null for ALT. In (h), **P = 0.0011 versus Null. i–n. Ldlr−/− mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3, and then fed a Western diet for 3 months (n = 8). Food intake and body weight were measured. Plasma total cholesterol, LDL-C (j), ApoA-I, ApoB (k), cholesterol lipoprotein profile (l), triglyceride lipoprotein profile (m), and ALT or AST levels (n) were determined. In (i), P = 0.59 or 0.13 versus Null for food intake or body weight, respectively. In (j), *P = 0.035 versus Null for cholesterol, and **P = 0.0035 versus Null for LDL-C. In (k), *P = 0.04 or 0.046 versus Null for plasma ApoA-I or ApoB, respectively. In (n), *P = 0.011 versus Null for AST, and **P = 0.0045 versus Null for ALT. o–t. Nine-weeks-old Apoe−/− mice or Apoe−/−Ldlr−/− mice were i.v. injected with AAV8-ALB-Null or AAV8-ALB-hATF3, and then fed a Western diet for 3 months. Food intake (n = 10 for Apoe−/−+AAV-Null or ATF3/Apoe−/−+AAV-ATF3, n = 15 for Apoe−/−Ldlr−/−+AAV-Null or Apoe−/−Ldlr−/−+AAV-ATF3) and body weight (n = 8 for Apoe−/−+AAV-Null or Apoe−/−+AAV-ATF3, n = 11 for Apoe−/−Ldlr−/−+AAV-Null, and n = 12 for Apoe−/−Ldlr−/−+AAV-ATF3) were measured (o). Plasma total cholesterol (p, left panel; n = 8 for Apoe−/−+AAV-Null or Apoe−/−+AAV-ATF3, n = 11 for Apoe−/−Ldlr−/−+AAV-Null, and n = 12 for Apoe−/−Ldlr−/−+AAV-ATF3), LDL-C (p, right panel; n = 10), ApoA-I (q, left panel; n = 10), ApoB (q, right panel; n = 8), cholesterol lipoprotein profile (r), triglyceride lipoprotein profile (s), and ALT or AST levels (t; n = 8 for Apoe−/−+AAV-Null or Apoe−/−+AAV-ATF3, n = 11 for Apoe−/−Ldlr−/−+AAV-Null, and n = 12 for Apoe−/−Ldlr−/−+AAV-ATF3) were determined. In (o), P = 0.288 or 0.93 for AAV-Null versus AAV-ATF3 for food intake or body weight, respectively. In (p), for plasma cholesterol, *P = 0.013 or **P = 0.000066 for AAV-Null versus AAV-ATF3 for Apoe−/− or Apoe−/−Ldlr−/− mice, respectively; for plasma LDL-C, **P = 0.0011 versus Null. In (q), *P = 0.031 or 0.022 versus Null for plasma ApoA-I or ApoB, respectively. In (t), for ALT, *P = 0.025 and **P = 0.0021 for AAV-Null versus AAV-ATF3 for Apoe−/− and Apoe−/−Ldlr−/− mice, respectively. For AST, *P = 0.019 or 0.015 for AAV-Null versus AAV-ATF3 for Apoe−/− or Apoe−/−Ldlr−/− mice, respectively. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (a–c, f, h–k, n, p (LDL-C), q) or two-way ANOVA with Turkey’s post hoc test for multiple comparisons (o, p (total cholesterol), t) was used for statistical analysis.
Extended Data Fig. 10 Loss of hepatic ATF3 increases plasma lipid levels.
a. C57BL/6J (WT) mice, Ldlr−/− mice or Apoe−/− mice (on a C57BL/6J background) were fed a standard chow diet (CD) or Western diet (WD) for 3 months (n = 8). Hepatic proteins were analyzed by Western blot assays (left panel) and protein levels were quantified (right panel). **P = 0.0048, 0.000089 or 0.000062 for WT + CD versus WT + WD, Ldlr−/−+WD or Apoe−/−+WD, respectively. b–e. Six-weeks-old Atf3fl/flApoe−/− mice and L-Atf3−/−Apoe−/− mice were fed a Western diet for 3 months (n = 8). Plasma cholesterol levels were analyzed (b). Plasma cholesterol (c) or TG (d) lipoprotein profiles were determined. Plasma ALT or AST levels were quantified (e). In (b), **P = 0.0016 versus Atf3fl/flApoe−/−. In (e), *P = 0.032 and **P = 0.0001 for Atf3fl/flApoe−/− versus L-Atf3−/−Apoe−/− for AST and ALT, respectively. All the data are expressed as mean ± SEM. All the data points are biological replicates. A two-tailed Student’s t-test (b, e) or one-way ANOVA with Turkey’s post hoc test for multiple comparisons (a) was used for statistical analysis.
Supplementary information
Supplementary Information
Supplementary Data 1
Supplementary Table 1
Characteristics of participants. Plasma parameters in 24 participants are presented. A two-tailed Student’s t-test was used for statistical analysis.
Supplementary Table 2
Oligonucleotide sequences for RT–qPCR.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Xu, Y., Li, Y., Jadhav, K. et al. Hepatocyte ATF3 protects against atherosclerosis by regulating HDL and bile acid metabolism. Nat Metab 3, 59–74 (2021). https://doi.org/10.1038/s42255-020-00331-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-020-00331-1
This article is cited by
-
Identification of novel SCD1 inhibitor alleviates nonalcoholic fatty liver disease: critical role of liver-adipose axis
Cell Communication and Signaling (2023)
-
Brown adipose tissue-derived Nrg4 alleviates endothelial inflammation and atherosclerosis in male mice
Nature Metabolism (2022)
-
ATF3 in atherosclerosis: a controversial transcription factor
Journal of Molecular Medicine (2022)