The nuclear envelope (NE) not only protects the genome from being directly accessed by detrimental agents but also regulates genome organization. Breaches in NE integrity threaten genome stability and impede cellular function. Nonetheless, the NE constantly remodels, and NE integrity is endangered in dividing or differentiating cells. Specifically, in unicellular eukaryotes undergoing closed mitosis, the NE expands instead of breaking down during chromosome segregation. The newly assembling nuclear pore complexes (NPCs) penetrate the existing NE in interphase. A peculiar example of NE remodelling during nuclear differentiation in Tetrahymena involves formation of the redundant NE and clustered NPCs. Even under these conditions, the NE remains intact. Many recent studies on unicellular organisms have revealed that nuclear membrane proteins, such as LEM-domain proteins, play a role in maintaining NE integrity. This review summarizes and discusses how nuclear membrane proteins participate in NE integrity.
Eukaryotic cells organize their genome within the nucleus, which is outlined by the nuclear envelope (NE). The nucleus is a seemingly robust but yet dynamic structure that plays an active role in genome organization and gene expression (1–3). Rupture of the NE threatens cell viability and drives genome instability (4). Nonetheless, the NE continuously undergoes dramatic deformation and reformation to facilitate chromosome segregation during cell cycle progression. Furthermore, migratory cancer cells rupture and repair their NE when passing through constricted channels (5). It is therefore important to understand how cells maintain NE integrity during the challenges presented by NE remodelling.
The NE is a double membrane structure. The double membrane of the NE is composed of outer and inner nuclear membranes (ONM and INM), each of which is a lipid bilayer. The ONM is a contiguous extension of the endoplasmic reticulum (ER) in the cytoplasm, whereas the INM faces toward the nucleoplasm. Therefore, the NE presents a physical barrier to hinder direct contact between the cytoplasm and nucleoplasm. Nucleocytoplasmic transport mainly occurs through nuclear pore complexes (NPCs) imbedded at the nuclear pores. NPCs are constructed in modules of multiple copies of about 30 different nucleoporins (6). These modules include the complex of scaffold nucleoporins Nup107-160 that supports the entire NPC, transmembrane nucleoporins that tether the NPC at the nuclear membrane, FG-repeat-containing nucleoporins that interact with cargo passing through the NPC, and cytoplasmic filament and nuclear basket nucleoporins that interact with the cytosol or nuclear factors (7). The nuclear lamina is a mesh-like structure composed of intermediate filament protein lamins that associates with the INM and assures the physicochemical integrity of the nucleus in metazoan cells. Mutations in lamins cause various inheritable diseases, generally called laminopathy, including Emery-Dreifuss muscular disease (mutations in lamin A or emerin) and Hutchison-Gilford Progeria (lamin A); thus, lamins are required for normal nuclear function in metazoans (8, 9). However, lamins are generally absent from most unicellular eukaryotes (10).
Most cells in multicellular organisms break down the NE during chromosome segregation (open mitosis) so that the chromosomes are accessible to the microtubules. After chromosome segregation, the NE immediately reforms around the chromosomes. In living human cells, this process of NE reassembly is regulated by a process involving barrier-to-autointegration factor and the INM protein emerin (11). Using an in vitro system of Xenopus oocyte extracts, the small GTPase Ran and Importin-β were shown to be the major regulators that drive NE reassembly, and the NPC components are one of their effectors (12–14). The Nup107-160 complex and transmembrane nucleoporins are first recruited to the chromatin to initiate post-mitotic NPC assembly during NE reformation (15, 16).
NPC biogenesis also occurs in interphase, during which the new NPCs are assembled onto the existing NE. Interphase NPC assembly, which threatens NPC integrity, requires fusion of the INM and ONM and involves formation of the highly curved membrane (
Fig. 1). In mammalian cells, it is reported that dorm-shaped evagination of the INM is an initial step of interphase NPC assembly (
17). Transmembrane nucleoporin Pom121 and membrane curvature sensor protein Nup133 also have roles in the early steps of interphase NPC assembly (
16,
18). However, molecular mechanisms for forming such curved membranes in interphase remain unknown. In yeast, it has been reported that the ER membrane-bending proteins Reticulons and Dp1/Yop1 facilitate membrane deformation at the onset of NPC biogenesis (
19). Another ER protein, Lunapark 1 (Lnp1), possibly stabilizes the resulting high membrane curvature during NPC assembly (
20,
21). Components of the NPC itself also sense and modulate the membrane curvature (
22,
23). It is thought that a completed NPC architecture acts as a membrane coat on the nuclear pore membrane and consolidates the curved membrane structure (
24,
25).
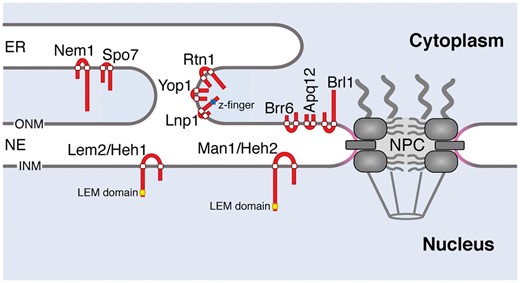
Fig. 1
Nuclear membrane proteins in unicellular organisms. The schematic diagram shows the NE and ER membranes, and the nuclear membrane proteins mentioned in this article. Predicted molecular structure and localization of the NE/ER proteins are indicated in red; domains of the protein shown in yellow, blue and white represent the LEM domain, zinc-finger domain and transmembrane domain, respectively. The highly curved membrane at the NPC is indicated in pink.
Many unicellular eukaryotes, such as Saccharomyces cerevisiae and Schizosaccharomyces pombe, rely on the interphase pathway for NPC biogenesis because there is no NE breakdown and reformation during their mitosis (closed mitosis) (26). In closed mitosis, the mitotic spindle is formed in the intact nucleus by insertion of the microtubule-organizing centres across the NE (27), allowing the spindle microtubules to interact with the segregating chromosomes while maintaining NE integrity. Moreover, to accommodate the elongated spindle, the NE undergoes expansion during closed mitosis (28, 29). Hence, the unicellular eukaryotes must have devised mechanisms to maintain NE integrity throughout the cell cycle. Here, we summarize recent findings regarding NE integrity maintenance by the nuclear membrane proteins in unicellular eukaryotes (Fig. 1).
The LEM Proteins Maintain NE Integrity and Genome Stability
The role of NE integrity in preventing genome instability has been investigated (
30,
31). The LEM (Lap2 (lamina-associated polypeptide 2), emerin, and Man1)-domain proteins are a group of integral INM proteins that share a helix-extension-helix motif that potentially binds to chromatin (
32,
33), and they have been implicated in both NE maintenance and genome stability (
31,
34–36). In the fission yeast
S.pombe, cells lacking the Lem2-encoding gene display defects in genome organization and stability, such as increased rates of mini-chromosome loss, and reduced levels of the repressive histone marker methylated lysine 9 of histone H3 (H3K9me) leading to defects in heterochromatin formation and mislocalization of peripheral heterochromatin (
35–37). Moreover, Lem2 augments heterochromatin formation in response to nutritional conditions, suggesting that Lem2 have some roles in sensing nutritional signals (
34) (
Fig. 2). Interestingly, an increased amount of the ER protein Lnp1 suppresses the phenotypes of cells lacking Lem2 (
36) (
Fig. 2). In the budding yeast
S.cerevisiae, Lnp1 is required for NPC integrity (
20), and the LEM protein Heh1 participates in NPC quality surveillance (
31). Thus, it is inferred that the LEM protein and Lnp1 act synergistically to maintain NE structural integrity in both of these yeasts. In agreement with this, our electron microscopy observations showed that deletion of the Lem2-encoding gene in the absence of Ima1, another conserved INM protein in
S.pombe, causes an abnormal branched NE inside the nucleus and abnormal vacuole-like membrane compartment within the nucleus (
38). In addition, overexpression of Lem2 causes NE membrane proliferation, whereas deletion of the Lem2-encoding gene leads to abnormal NE bulges and gaps, as observed with electron microscopy (
35). These results suggest an important role of Lem2 in regulating NE membrane homeostasis in
S.pombe.
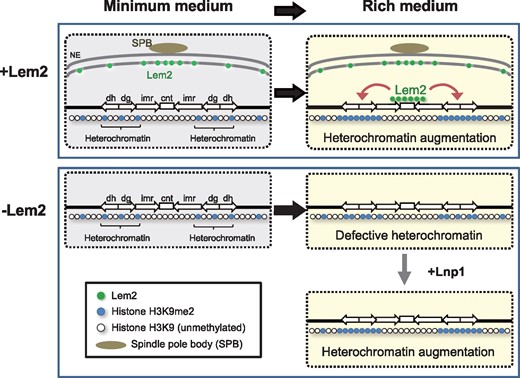
Fig. 2
Lem2 functions on heterochromatin formation. Upon transfer to a nutritionally rich condition, Lem2 associates with the inner centromere region and augments heterochromatin formation at the outer centromeric regions in wild-type cells (upper panel). In the absence of Lem2, augmentation of the centromeric heterochromatin formation is diminished (lower panel). This phenotype is rescued by the additional expression of Lnp1. Green circles, Lem2; blue circles, chromatin containing methylated histone H3K9; and white circles, chromatin containing unmethylated histone H3K9. Reproduced from (36).
The Nem1-Spo7 Complex Negatively Regulates Nuclear Membrane Growth
Nuclear membrane homeostasis is controlled through the metabolism of phosphatidic acid (PA). PA can be converted into the phospholipids such as phosphatidylcholine (PC) and phosphatidylinositol (PI), which are used for membrane lipid synthesis. PA can also be converted into diacylglycerol (DAG); some of DAG is turned into triacylglycerol (TAG) for lipid storage in lipid droplets (LDs) (
39). The PA phosphatase lipin catalyzes PA into DAG (
40,
41). In
S.cerevisiae, the ER/NE-localized phosphatase complex Nem1-Spo7 dephosphorylates and activates the yeast lipin (Pah1 in
S.cerevisiae) at the ER/NE membrane when the cells are under glucose starvation (
40,
42). Moreover, the NE recruitment of the yeast lipin is accompanied by an increase of LD biogenesis (
40). In the absence of Nem1, Spo7, or lipin,
S.cerevisia cells display striking ER/NE membrane expansion (
43,
44), indicating that most PA is used for membrane lipid synthesis in the mutants. Therefore, the activity of Nem1-Spo7 drives the use of PA for neutral lipid storage and restricts membrane lipid synthesis (
Fig. 3).
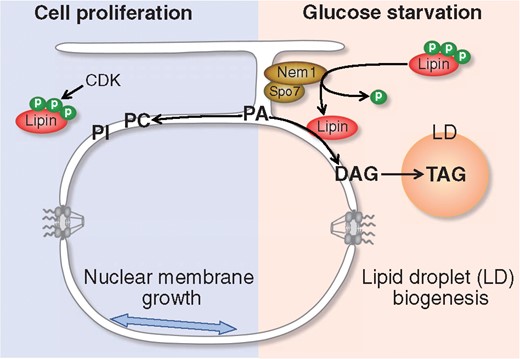
Fig. 3
Nem1-Spo7 complex negatively controls nuclear membrane expansion. Nem1-Spo7 is a serine/threonine phosphatase complex that localizes to ER/NE. Upon glucose starvation (right panel), the Nem1-Spo7 complex dephosphorylates cytosolic lipin. Dephosphorylated lipin localizes to ER/NE and catalyzes conversion of PA into DAG. DAG is then turned into TAG that goes into LD for LD biogenesis. On the other hand, CDK phosphorylates and sequesters lipin in the cytosol during cell proliferation (left panel). Without the action of lipin, PA is used for membrane lipid synthesis (such as PC and PI), which leads to nuclear membrane expansion.
In order to accommodate elongation of the mitotic spindle, membrane lipid synthesis is crucial for NE expansion during closed mitosis (29). In S.pombe, cyclin-dependent kinase (CDK) counteracts the action of Nem1-Spo7 complex by phosphorylating lipin (S.pombe Ned1) and admits nuclear membrane growth during closed mitosis (45). Since S.cerevisiae lipin is also phosphorylated by CDK (43), the two yeasts possibly uses the same regulatory mechanism to ensure NE integrity during cell proliferation (Fig. 3). On the other hand, in Schizosaccharomyces japonicus, which is closely related to S.pombe, lipin is not subjected to CDK regulation, and there is no addition of the nuclear membrane during mitosis (45, 46), possibly reflecting the finding that S.japonicus partially ruptures the NE in anaphase (semi-open mitosis) (46, 47).
The Brr6 Protein Complex Monitors Nuclear Membrane Homeostasis
Yeasts that undergo closed or semi-open mitosis assemble new NPCs and spindle pole bodies (SPBs; the yeast microtubule-organizing centre) on the existing NE, which remodels the NE membranes and challenges NE integrity. The brr6 mutants were found to be defective in nuclear transport, NPC assembly, and SPB insertion in yeast (48, 49), indicating that Brr6 is important for NE integrity maintenance. Brr6 is an ER/NE integral membrane protein (48). In S. cerevisiae, Brr6, Apq12 and Brl1 (Brr6-like protein 1) form a complex adjacent to the NPC at the NE (50). Moreover, brr6 or apq12 mutants exhibit ER/NE membrane protrusions and a characteristic phenotype in which the NPCs are covered with an additional lipid bilayer (51, 52), similar to what has been observed in cells deficient in very-long-chain fatty acids synthesis (53). In accordance with this, analyses of the membrane lipid composition have revealed that short-chain and unsaturated fatty acids accumulate in the brr6, brl1 and apq12 mutants (50). It is believed that the Brr6 complex monitors the membrane lipid content in response to NE biophysical changes (such as membrane bending during NPC assembly and SPB insertion) through yet-unknown mechanisms (50). BLAST searches revealed that Brr6 homologs were found not only in fungi but also in unicellular eukaryotes that maintain intact NEs throughout the cell cycle, namely the amoeba Dictyostelium discoideum (54), the alga Cyanidioschyzon merolae (55), and the protozoan parasite Plasmodium falciparum (56), suggesting that Brr6 or Brr6-like proteins coevolved with the system of closed mitosis to monitor nuclear membrane homeostasis.
Large-Scale NE Remodelling and Nuclear Differentiation in the Ciliate Tetrahymena
The ciliate protozoa Tetrahymena are unicellular organisms, and they contain two functionally and structurally distinct nuclei in the same cytoplasm: a macronucleus (MAC) with somatic functions and a micronucleus (MIC) with germline functions (57). The MAC is transcriptionally active during all life cycle stages, and is used to produce proteins required for all life activities. In contrast, the MIC is transcriptionally inert and is required for the inheritance of the genome during sexual reproduction. The NEs of both the MAC and MIC stay intact throughout all life cycle stages, including the nuclear division stage where the nuclei are extensively elongated. In Tetrahymena thermophila, the structural differences between the MAC and MIC NEs are characterized by the MAC-type and MIC-type NPCs with distinct sets of nucleoporins (58, 59).
Tetrahymena undergo sexual conjugation under starvation conditions, during which the MICs of each cell of the conjugating pair generate gamete nuclei through meiosis, subsequently producing the zygotic nucleus after reciprocal fertilization. The zygotic nucleus, which originates from the MIC, subsequently differentiates into two MACs and two MICs after two rounds of post-zygotic divisions (
Fig. 4A). During this stage of nuclear differentiation, the NE is extensively remodelled. First, redundant NEs are formed in the differentiating post-zygotic nuclei. Second, for the MAC-destined nucleus, the MIC-type NPC in the post-zygotic nucleus is replaced with the MAC-type NPC. Third, in the MIC-destined nucleus, the NPCs are clustered in the inner set of the NE at the region of redundant NEs (
Fig. 4B and C) (
60). This curious example of clustered NPCs is also found in the early developmental stages of
Drosophila blastoderm embryos and appears to be regulated by nuclear membrane proteins (
61), suggesting that the ciliate may also use the NE proteins to regulate NPC organization during nuclear differentiation, although the existence of such NE proteins has not been demonstrated.
Fig. 4
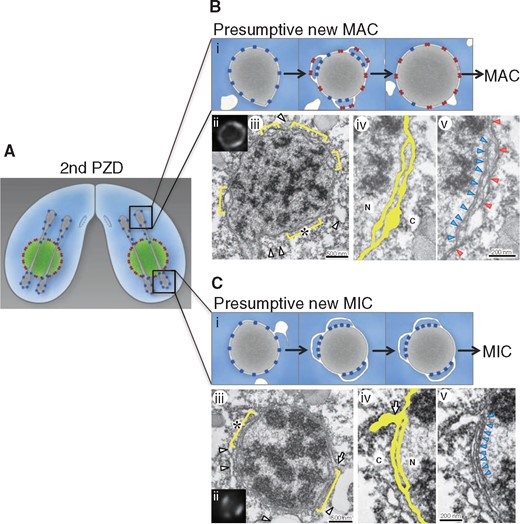
Redundant NE and NPC clusters formed in differentiating nuclei of Tetrahymena. (A) Schematic diagram of conjugating T.thermophila cells at the stage of second post-zygotic division (second PZD). Green circles with red dots and dumbbell-shaped structures with blue dots represent the MAC nuclei and the dividing zygotic nuclei originated from the MIC, respectively; red and blue dots represent MAC-type and MIC-type NPCs, respectively. Upper and lower parts of the cells represent anterior and posterior regions, respectively. MIC-type NPCs (blue dots) are distributed equally to both daughter nuclei during the second PZD. (B, C) Images of an anterior nucleus (presumptive new MAC) (B) and a posterior nucleus (presumptive new MIC) (C). Panels (i–v) in (B) and (C) are as follows. (i) Schematic diagram of differentiating presumptive new MAC (B) and MIC (C) during early developmental stages. At or immediately after the end of the second PZD, the preexisting MIC-type NPCs make several clusters on each post-zygotic nucleus. Various membranes, including ER/vacuolar membranes, approach both nuclei and fuse to the NE to generate regions with redundant NE; the MIC-type NPCs are packed in clusters in the inner set of these redundant NEs. Newly synthesized MAC-type NPCs (red dots) assemble on the NE. (ii–v) Correlative light-electron microscopic images. (ii) Fluorescence image of the nucleus stained with GFP-Nup93. (iii) Electron micrographs of the same nucleus shown in (ii). Yellow brackets indicate the regions where NE redundancy (two sets of NE) is observed. Arrowheads indicate membrane structures near the nucleus. (iv, v) Enlarged images of the electron micrograph indicated by asterisks in (iii). Yellow paint represents the structure of the redundant NE. ‘N’ and ‘C’ represent nuclear and cytoplasmic sides, respectively. Blue and red arrowheads in (v) represent the positions of the NPCs clustered in the inner (nucleoplasmic side) set of the NE and the NPCs facing the outer (cytoplasm side) set of the NE, respectively. Modified from (60). For details, see Supplementary Movie S2 in the web site: http://jcs.biologists.org/content/joces/suppl/2015/04/29/128.9.1812.DC1/JCS167353.pdf.
Concluding Remarks
The yeasts S.cerevisiae and S.pombe maintain NE integrity throughout their cell cycles, during which NPC or SPB assembly/disassembly and nuclear growth periodically occur. It is becoming clearer that many of the nuclear membrane proteins play an active role in NE integrity maintenance. In S.cerevisiae, the LEM proteins recognize the defective NPCs and recruit repair machinery (31). Whether this role in NPC quality control is conserved in other organisms remains an open question. Nonetheless, NE membrane homeostasis is affected by the misexpression of the LEM proteins in S.pombe (35). NE membrane homeostasis is regulated by lipid biogenesis. The Nem1-Spo7 complex and the Brr6 protein complex monitor membrane lipid homeostasis in response to environmental changes (40, 50). Brr6, which is likely unique to unicellular eukaryotic cells, has no homologue to any known lipid enzyme, thus raising the question of how it regulates membrane lipid homeostasis.
On the other hand, the LEM proteins and the lipin activation pathway by the Nem1-Spo7 complex are found in metazoans (34, 39, 62). Mutations in human Lipin-1 and LEM-domain protein emerin are associated with recurrent rhabdomyolysis and Emery-Dreifuss muscular dystrophy, respectively (63, 64), emphasizing the pathological importance of maintaining NE integrity by these proteins.
Furthermore, studies on Tetrahymena suggest that the formation of the NE structure of the redundant NEs and clustered NPCs may be important in a wide range of eukaryotes for controlling nuclear differentiation. Highly conserved NE proteins such as LEM-domain proteins or other NE proteins found in yeasts are probably involved in the formation of this peculiar NE structure while maintaining NE integrity. Elucidating the molecular basis of NE remodelling in Tetrahymena will provide insights into the addition of the NE to the preexisting NE and into the maintenance of NE integrity when switching nuclear functions.
Acknowledgements
We thank Ms Yasuha Kinugasa (Osaka University) and Dr Yasuhiro Hirano (Osaka University) for valuable discussions.
Funding
This study was supported by JSPS Kakenhi, Grant Numbers JP13F03384 to H.-J.Y., JP15K07066 to M.I., JP16H01309 to Y.H. and JP26291007, JP25116006 to T.H.
Conflict of Interest
None declared.
References
1Casolari
J. M.
, Brown
C. R.
, Komili
S.
, West
J.
, Hieronymus
H.
, Silver
P. A.
(
2004
)
Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization
.
Cell
117
,
427
–
39
2Finlan
L.E.
, Sproul
D.
, Thomson
I.
, Boyle
S.
, Kerr
E.
, Perry
P.
, Ylstra
B.
, Chubb
J.R.
, Bickmore
W.A.
(
2008
)
Recruitment to the nuclear periphery can alter expression of genes in human cells
.
PLoS Genet
.
4
,
e1000039
3Mekhail
K.
, Moazed
D.
(
2010
)
The nuclear envelope in genome organization, expression and stability
.
Nat. Rev. Mol. Cell Biol
.
11
,
317
–
28
4Lim
S.
, Quinton
R. J.
, Ganem
N. J.
(
2016
)
Nuclear envelope rupture drives genome instability in cancer
.
Mol. Biol. Cell
27
,
3210
–
3
5Denais
C. M.
, Gilbert
R. M.
, Isermann
P.
, McGregor
A. L.
, te Lindert
M.
, Weigelin
B.
, Davidson
P.M.
, Friedl
P.
, Wolf
K.
, Lammerding
J.
(
2016
)
Nuclear envelope rupture and repair during cancer cell migration
.
Science
352
,
353
–
8
6Kabachinski
G.
, Schwartz
T. U.
(
2015
)
The nuclear pore complex–structure and function at a glance
.
J. Cell Sci
.
128
,
423
–
9
7Beck
M.
, Hurt
E.
(
2017
)
The nuclear pore complex: understanding its function through structural insight
.
Nat. Rev. Mol. Cell Biol
.
18
,
73
–
89
8Worman
H.J.
, Schirmer
E.C.
(
2015
)
Nuclear membrane diversity: underlying tissue-specific pathologies in disease?
.
Curr. Opin. Cell Biol
.
34
,
101
–
12
9Burke
B.
, Stewart
C.L.
(
2002
)
Life at the edge: the nuclear envelope and human disease
.
Nat. Rev. Mol. Cell Biol
.
3
,
575
–
85
10Iwamoto
M.
, Hiraoka
Y.
, Haraguchi
T.
(
2016
)
Uniquely designed nuclear structures of lower eukaryotes
.
Curr. Opin. Cell Biol
.
40
,
66
–
73
11Haraguchi
T.
, Koujin
T.
, Segura-Totten
M.
, Lee
K.K.
, Matsuoka
Y.
, Yoneda
Y.
, Wilson
K.L.
, Hiraoka
Y.
(
2001
)
BAF is required for emerin assembly into the reforming nuclear envelope
.
J. Cell Sci
.
114
,
4575
–
85
12Askjaer
P.
, Galy
V.
, Hannak
E.
, Mattaj
I. W.
(
2002
)
Ran GTPase cycle and importins alpha and beta are essential for spindle formation and nuclear envelope assembly in living Caenorhabditis elegans embryos
.
Mol. Biol. Cell
13
,
4355
–
70
13Harel
A.
, Chan
R. C.
, Lachish-Zalait
A.
, Zimmerman
E.
, Elbaum
M.
, Forbes
D. J.
(
2003
)
Importin beta negatively regulates nuclear membrane fusion and nuclear pore complex assembly
.
Mol. Biol. Cell
14
,
4387
–
96
14Walther
T. C.
, Askjaer
P.
, Gentzel
M.
, Habermann
A.
, Griffiths
G.
, Wilm
M.
, Mattaj
I.W.
, Hetzer
M.
(
2003
)
RanGTP mediates nuclear pore complex assembly
.
Nature
424
,
689
–
94
15Dultz
E.
, Zanin
E.
, Wurzenberger
C.
, Braun
M.
, Rabut
G.
, Sironi
L.
, Ellenberg
J.
(
2008
)
Systematic kinetic analysis of mitotic dis- and reassembly of the nuclear pore in living cells
.
J. Cell Biol
.
180
,
857
–
65
16Doucet
C. M.
, Talamas
J. A.
, Hetzer
M. W.
(
2010
)
Cell cycle-dependent differences in nuclear pore complex assembly in metazoa
.
Cell
141
,
1030
–
41
17Otsuka
S.
, Bui
K. H.
, Schorb
M.
, Hossain
M. J.
, Politi
A.Z.
, Koch
B.
, Eltsov
M.
, Beck
M.
, Ellenberg
J.
, (
2016
)
Nuclear pore assembly proceeds by an inside-out extrusion of the nuclear envelope
.
Elife
5
,
e19071
18Funakoshi
T.
, Clever
M.
, Watanabe
A.
, Imamoto
N.
(
2011
)
Localization of Pom121 to the inner nuclear membrane is required for an early step of interphase nuclear pore complex assembly
.
Mol. Biol. Cell
22
,
1058
–
69
19Dawson
T. R.
, Lazarus
M. D.
, Hetzer
M. W.
, Wente
S. R.
(
2009
)
ER membrane-bending proteins are necessary for de novo nuclear pore formation
.
J. Cell Biol
.
184
,
659
–
75
20Casey
A. K.
, Chen
S.
, Novick
P.
, Ferro-Novick
S.
, Wente
S. R.
(
2015
)
Nuclear pore complex integrity requires Lnp1, a regulator of cortical endoplasmic reticulum
.
Mol. Biol. Cell
26
,
2833
–
44
21Chen
S.
, Desai
T.
, McNew
J.A.
, Gerard
P.
, Novick
P.J.
, Ferro-Novick
S.
(
2015
)
Lunapark stabilizes nascent three-way junctions in the endoplasmic reticulum
.
Proc. Natl. Acad. Sci. U. S. A
.
112
,
418
–
23
22Drin
G.
, Casella
J.-F.
, Gautier
R.
, Boehmer
T.
, Schwartz
T.U.
, Antonny
B.
(
2007
)
A general amphipathic alpha-helical motif for sensing membrane curvature
.
Nat. Struct. Mol. Biol
.
14
,
138
–
46
23Mészáros
N.
, Cibulka
J.
, Mendiburo
M.J.
, Romanauska
A.
, Schneider
M.
, Köhler
A.
(
2015
)
Nuclear pore basket proteins are tethered to the nuclear envelope and can regulate membrane curvature
.
Dev. Cell
33
,
285
–
98
24Devos
D.
, Dokudovskaya
S.
, Williams
R.
, Alber
F.
, Eswar
N.
, Chait
B.T.
, Rout
M.P.
, Sali
A.
(
2006
)
Simple fold composition and modular architecture of the nuclear pore complex
.
Proc. Natl. Acad. Sci. U. S. A
.
103
,
2172
–
7
25Rexach
M.
(
2009
)
Piecing together nuclear pore complex assembly during interphase
.
J. Cell Biol
.
185
,
377
–
9
26Asakawa
H.
, Yang
H.-J.
, Hiraoka
Y.
, Haraguchi
T.
(
2016
)
Virtual nuclear envelope breakdown and its regulators in fission yeast meiosis
.
Front. Cell Dev. Biol
.
4
,
5
–
12
27Ding
R.
, West
R.R.
, Morphew
D.M.
, Oakley
B.R.
, McIntosh
J.R.
(
1997
)
The spindle pole body of Schizosaccharomyces pombe enters and leaves the nuclear envelope as the cell cycle proceeds
.
Mol. Biol. Cell
.
8
,
1461
–
79
28Gu
Y.
, Yam
C.
, Oliferenko
S.
(
2012
)
Divergence of mitotic strategies in fission yeasts
.
Nucleus
3
,
220
–
5
29Takemoto
A.
, Kawashima
S.A.
, Li
J.-J.
, Jeffery
L.
, Yamatsugu
K.
, Elemento
O.
, Nure
P.
(
2016
)
Nuclear envelope expansion is crucial for proper chromosomal segregation during a closed mitosis
.
J. Cell Sci
.
129
,
1250
–
9
30Webster
B.M.
, Lusk
C.P.
(
2016
)
Border safety: quality control at the nuclear envelope
.
Trends Cell Biol
.
26
,
29
–
39
31Webster
B.M.
, Thaller
D.J.
, Jäger
J.
, Ochmann
S.E.
, Borah
S.
, Lusk
C.P.
(
2016
)
Chm7 and Heh1 collaborate to link nuclear pore complex quality control with nuclear envelope sealing
.
embo J
.
35
,
2447
–
67
32Cai
M.
, Huang
Y.
, Ghirlando
R.
, Wilson
K.L.
, Craigie
R.
, Clore
G.M.
(
2001
)
Solution structure of the constant region of nuclear envelope protein LAP2 reveals two Lem-domain structures: one binds BAF and the other binds DNA
.
embo J
.
20
,
4399
–
407
33Lin
F.
, Blake
D.L.
, Callebaut
I.
, Skerjanc
I.S.
, Holmer
L.
, McBurney
M.W.
, Paulin-Levasseur
M.
, Worman
H.J.
(
2000
)
Man1, an inner nuclear membrane protein that shares the Lem domain with lamina-associated polypeptide 2 and emerin
.
J. Biol. Chem
.
275
,
4840
–
7
34Brachner
A.
, Foisner
R.
(
2011
)
Evolvement of Lem proteins as chromatin tethers at the nuclear periphery
.
Biochem. Soc. Trans
.
39
,
1735
–
41
35Gonzalez
Y.
, Saito
A.
, Sazer
S.
(
2011
)
Fission yeast Lem2 and Man1 perform fundamental functions of the animal cell nuclear lamina
.
Nucleus
3
,
60
–
76
36Tange
Y.
, Chikashige
Y.
, Takahata
S.
, Kawakami
K.
, Higashi
M.
, Mori
C.
, Kojidani
T.
, Hirano
Y.
, Asakawa
H.
, Murakami
Y.
, Haraguchi
T.
, Hiraoka
Y.
(
2016
)
Inner nuclear membrane protein Lem2 augments heterochromatin formation in response to nutritional conditions
.
Genes Cells
21
,
812
–
32
37Barrales
R.R.
, Forn
M.
, Georgescu
P.R.
, Sarkadi
Z.
, Braun
S.
(
2016
)
Control of heterochromatin localization and silencing by the nuclear membrane protein Lem2
.
Genes Dev
.
30
,
133
–
48
38Hiraoka
Y.
, Maekawa
H.
, Asakawa
H.
(
2011
)
Inner nuclear membrane protein Ima1 is dispensable for intranuclear positioning of centromeres
.
Genes Cells
16
,
1000
–
11
39Bahmanyar
S.
, Biggs
R.
, Schuh
A.L.
, Desai
A.
, Müller-Reichert
T.
, Audhya
A.
, Dixon
J.E.
, Oegema
K.
(
2014
)
Spatial control of phospholipid flux restricts endoplasmic reticulum sheet formation to allow nuclear envelope breakdown
.
Genes Dev
.
28
,
121
–
6
40Barbosa
A.D.
, Sembongi
H.
, Su
W.-M.
, Abreu
S.
, Reggiori
F.
, Carman
G.M.
, Siniossoglou
S.
(
2015
)
Lipid partitioning at the nuclear envelope controls membrane biogenesis
.
Mol. Biol. Cell
26
,
3641
–
57
41Han
G.-S.
, Wu
W.-I.
, Carman
G.M.
(
2006
)
The Saccharomyces cerevisiae Lipin homolog is a Mg2+-dependent phosphatidate phosphatase enzyme
.
J. Biol. Chem
.
281
,
9210
–
8
42Karanasios
E.
, Han
G.-S.
, Xu
Z.
, Carman
G.M.
, Siniossoglou
S.
(
2010
)
A phosphorylation-regulated amphipathic helix controls the membrane translocation and function of the yeast phosphatidate phosphatase
.
Proc. Natl. Acad. Sci. U. S. A
.
107
,
17539
–
44
43Santos-Rosa
H.
, Leung
J.
, Grimsey
N.
, Peak-Chew
S.
, Siniossoglou
S.
(
2005
)
The yeast lipin Smp2 couples phospholipid biosynthesis to nuclear membrane growth
.
embo J
.
24
,
1931
–
41
44Siniossoglou
S.
, Santos-Rosa
H.
, Rappsilber
J.
, Mann
M.
, Hurt
E.
(
1998
)
A novel complex of membrane proteins required for formation of a spherical nucleus
.
embo J
.
17
,
6449
–
64
45Makarova
M.
, Gu
Y.
, Chen
J.-S.
, Beckley
J.R.
, Gould
K.L.
, Oliferenko
S.
(
2016
)
Temporal regulation of lipin activity diverged to account for differences in mitotic programs
.
Curr. Biol
.
26
,
237
–
43
46Yam
C.
, He
Y.
, Zhang
D.
, Chiam
K.-H.
, Oliferenko
S.
(
2011
)
Divergent strategies for controlling the nuclear membrane satisfy geometric constraints during nuclear division
.
Curr. Biol
.
21
,
1314
–
9
47Aoki
K.
, Hayashi
H.
, Furuya
K.
, Sato
M.
, Takagi
T.
, Osumi
M.
, Kimura
A.
, Niki
H.
(
2011
)
Breakage of the nuclear envelope by an extending mitotic nucleus occurs during anaphase in Schizosaccharomyces japonicus
.
Genes Cells
16
,
911
–
26
48de Bruyn Kops
A.
, Guthrie
C.
(
2001
)
An essential nuclear envelope integral membrane protein, Brr6p, required for nuclear transport
.
embo J
.
20
,
4183
–
93
49Tamm
T.
, Grallert
A.
, Grossman
E.P.S.
, Alvarez-Tabares
I.
, Stevens
F.E.
, Hagan
I.M.
(
2011
)
Brr6 drives the Schizosaccharomyces pombe spindle pole body nuclear envelope insertion/extrusion cycle
.
J. Cell Biol
.
195
,
467
–
84
50Lone
M.A.
, Atkinson
A.E.
, Hodge
C.A.
, Cottier
S.
, Martínez-Montañés
F.
, Maithel
S.
, Mène-Saffrané
L.
, Cole
C.N.
, Schneiter
R.
(
2015
)
Yeast integral membrane proteins Apq12, Brl1, and Brr6 form a complex important for regulation of membrane homeostasis and nuclear pore complex biogenesis
.
Eukaryot. Cell
14
,
1217
–
27
51Hodge
C.A.
, Choudhary
V.
, Wolyniak
M.J.
, Scarcelli
J.J.
, Schneiter
R.
, Cole
C.N.
(
2010
)
Integral membrane proteins Brr6 and Apq12 link assembly of the nuclear pore complex to lipid homeostasis in the endoplasmic reticulum
.
J. Cell Sci
.
123
,
141
–
51
52Scarcelli
J.J.
, Hodge
C.A.
, Cole
C.N.
(
2007
)
The yeast integral membrane protein Apq12 potentially links membrane dynamics to assembly of nuclear pore complexes
.
J. Cell Biol
.
178
,
799
–
812
53Schneiter
R.
, Hitomi
M.
, Ivessa
A.S.
, Fasch
E.
, Kohlwein
S.D.
, Tartakoff
A.M.
(
1996
)
A yeast acetyl coenzyme a carboxylase mutant links very-long-chain fatty acid synthesis to the structureand function of the nuclear membrane-pore pomplex
.
Mol. Cell. Biol
.
16
,
7161
–
72
54O’Day
D.H.
, Budniak
A.
(
2014
)
Nucleocytoplasmic protein translocation during mitosis in the social amoebozoan Dictyostelium discoideum
.
Biol. Rev
90
,
126
–
41
55Yagisawa
F.
, Fujiwara
T.
, Kuroiwa
H.
, Nishida
K.
, Imoto
Y.
, Kuroiwa
T.
(
2011
)
Mitotic inheritance of endoplasmic reticulum in the primitive red alga Cyanidioschyzon merolae
.
Protoplasma
249
,
1129
–
35
56Gerald
N.
, Mahajan
B.
, Kumar
S.
(
2011
)
Mitosis in the human malaria parasite Plasmodium falciparum
.
Eukaryot. Cell
10
,
474
–
82
57Orias
E.
, Cervantes
M.D.
, Hamilton
E.P.
(
2011
)
Tetrahymena thermophila, a unicellular eukaryote with separate germline and somatic genomes
.
Res. Microbiol
.
162
,
578
–
86
58Iwamoto
M.
, Hiraoka
Y.
, Haraguchi
T.
(
2015
)
The nuclear pore complex acts as a master switch for nuclear and cell differentiation
.
Commun. Integr. Biol
.
8
,
1
–
3
59Iwamoto
M.
, Mori
C.
, Kojidani
T.
, Bunai
F.
, Hori
T.
, Fukagawa
T.
, Hiraoka
Y.
, Haraguchi
T.
(
2009
)
Two distinct repeat sequences of Nup98 nucleoporins characterize dual nuclei in the binucleated ciliate Tetrahymena
.
Curr. Biol
.
19
,
843
–
7
60Iwamoto
M.
, Koujin
T.
, Osakada
H.
, Mori
C.
, Kojidani
T.
, Matsuda
A.
, Asakawa
H.
, Hiraoka
Y.
, Haraguchi
T.
(
2015
)
Biased assembly of the nuclear pore complex is required for somatic and germline nuclear differentiation in Tetrahymena
.
J. Cell Sci
.
128
,
1812
–
23
61Hampoelz
B.
, Mackmull
M.T.
, Machado
P.
, Ronchi
P.
, Bui
K.H.
, Schieber
N.
, Santarella-Mellwig
R.
, Necakov
A.
, Andrés-Pons
A.
, Philippe
J.M.
, Lecuit
T.
, Schwab
Y.
, Beck
M.
(
2016
)
Pre-assembled nuclear pores insert into the nuclear envelope during early development
.
Cell
166
,
664
–
78
62Han
S.
, Bahmanyar
S.
, Zhang
P.
, Grishin
N.
, Oegema
K.
, Crooke
R.
, Graham
M.
, Reue
K.
, Dixon
J.E.
, Goodman
J.M.
(
2012
)
Nuclear envelope phosphatase 1-regulatory subunit 1 (formerly TMEM188) is the metazoan Spo7p ortholog and functions in the lipin activation pathway
.
J. Biol. Chem
.
287
,
3123
–
37
63Zeharia
A.
, Shaag
A.
, Houtkooper
R.H.
, Hindi
T.
, de Lonlay
P.
, Erez
G.
, Hubert
L.
, Saada
A.
, de Keyzer
Y.
, Eshel
G.
, Vaz
F.M.
, Pines
O.
, Elpeleg
O.
(
2008
)
Mutations in LPIN1 cause recurrent acute myoglobinuria in childhood
.
Am. J. Hum. Genet
83
,
489
–
94
64Berk
J.M.
, Tifft
K.E.
, Wilson
K.L.
(
2014
)
The nuclear envelope Lem-domain protein emerin
.
Nucleus
4
,
298
–
314
Abbreviations
Abbreviations
CDK
DAG
ER
INM
LDs
Lnp1
MAC
MIC
NE
NPCs
ONM
PA
PC
PI
SPBs
TAG
© The Authors 2017. Published by Oxford University Press on behalf of the Japanese Biochemical Society. All rights reserved
PDF