Abstract
Electrochemical aptamer-based (EAB) sensors encompass the only biosensor approach yet reported that is simultaneously: (1) independent of the chemical or enzymatic reactivity of its target, rendering it general; (2) continuous and lag-free; and (3) selective enough to deploy in situ in the living body. Consistent with this, in vivo EAB sensors supporting the seconds-resolved, real-time measurement of multiple drugs and metabolites have been reported, suggesting the approach may prove of value in biomedical research and the diagnosis, treatment, and monitoring of disease. However, to apply these devices in long-duration animal models, much less in human patients, requires that they be free of any significant pathogen load. Thus motivated, here we have characterized the compatibility of EAB sensors with standard sterilization and high-level disinfection techniques. Doing so, we find that, while many lead to significant sensor degradation, treatment with CIDEX OPA (0.55% ortho-phthalaldehyde) leads to effective disinfection without causing any detectable loss in sensor performance.

Export citation and abstract BibTeX RIS

This is an open access article distributed under the terms of the Creative Commons Attribution 4.0 License (CC BY, http://creativecommons.org/licenses/by/4.0/), which permits unrestricted reuse of the work in any medium, provided the original work is properly cited.
Electrochemical aptamer-based (EAB) sensors are a platform technology supporting continuous, real-time measurement of the local concentration of specific molecular targets. 1 Consisting of a redox reporter-modified DNA or RNA aptamer attached to an interrogating electrode via a self-assembled monolayer, EAB sensors employ the binding-induced folding of their aptamer to generate a change in electron transfer rate that is easily monitored using, for example, square wave voltammetry. This signaling mechanism renders EAB sensors; (1) independent of the chemical or enzymatic reactivity of their targets, and thus general; (2) single-step and rapidly reversible, thus enabling high-frequency (typically seconds-resolved) measurements; and (3) highly selective, thus suitable for deployment in situ in the living body. Consistent with this, EAB sensors have been shown to support real-time, seconds-resolved measurements the plasma concentrations of multiple drugs and metabolites when placed intravenously in live rats. 2,3 The value such real-time molecular measurements could bring not only to the biomedical research community, but also to clinical practice, suggests a compelling case for the use of EAB sensors in human health.
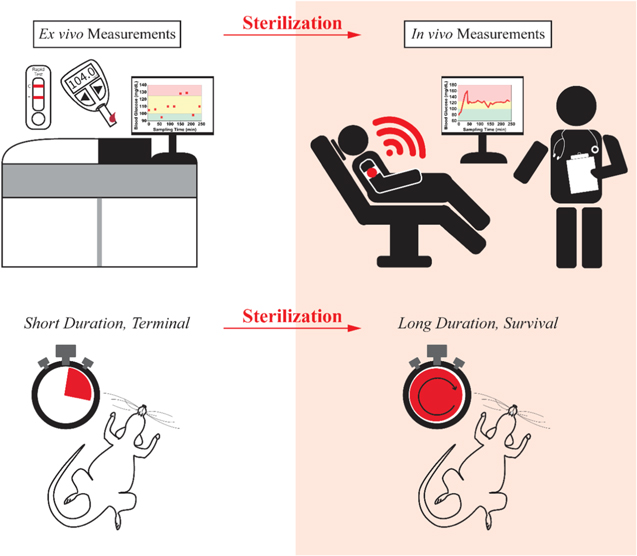
Clinical applications require technologies that are both effective and safe (Fig. 1). In the case of implantable devices, the efficient removal of pathogens is a critical element of safety. Likewise, animal welfare concerns require that devices placed in vivo in support of long-duration or "survival" studies (i.e., in which the animal is not sacrificed within hours of insertion of the device) be disinfected prior to their use. 4,5 Unfortunately, however, sterilization and disinfection are often achieved via destruction of the genetic material of any pathogens present, which suggests that many treatment approaches may irreversibly damage the DNA aptamers that EAB sensors rely on. This said, viral 6 and bacterial 7 genomes are orders of magnitude longer than the few tens of bases length typical of aptamers, leading us to hypothesize that it might prove possible to achieve high level reduction of pathogen loads without unduly damaging EAB sensors. Given this, here we set out to identify protocols that effectively eliminate pathogens while leaving EAB sensor performance largely or entirely unaffected.
Figure 1. Both in vivo clinical applications and long-duration (i.e., survival) animal model studies will require the high-level disinfection of EAB sensors prior to use. Here we explore the extent to which EAB function is compatible with many commonly employed disinfection and sterilization approaches.
Download figure:
Standard image High-resolution imageResults and Discussion
As our initial testbed for determining the extent to which various sterilization and disinfection approaches are compatible with EAB sensor operation, we employed a previously reported sensor against the antibiotic vancomycin. 3,8 We selected this sensor because, at 45 bases, the DNA aptamer it employs is longer than those generally seen in EAB sensors, suggesting this sensor may be particularly susceptible to DNA damage. To evaluate the compatibility of sterilization and disinfection techniques with the function of this sensor, we employed three performance metrics. First, we compared the "signal gain" of treated (i.e., sterilized or disinfected) and control (untreated) sensors at the square-wave frequency where gain is largest. Gain, which is defined as the relative change in current between the absence of target and saturating target, provides a good general measure of sensor performance. Next, we measured any difference in the square wave peak current per unit area (referred to as "packing density") of treated and matched control sensors, as this measure is sensitive to cleavage of the aptamer, loss of the thiol-on-gold self-assembled monolayer (SAM) that attaches the aptamer to the electrode, or loss or irreversible modification of the redox reporter. Of note, while they are weakly related (i.e., gain is a function of packing density 9 ), these first two metrics can also change independently of one another. Finally, we compared the binding curves (e.g., the binding midpoints) of sterilized and unsterilized sensors, a measure that, when taken with gain, indicates whether or not the treatment has changed the aptamer's target binding properties and thus altered the sensor's calibration. For reference, we have previously observed 3 that, when interrogated with a square-wave frequency of 60 Hz, untreated vancomycin sensors achieve gain of 120(±3)% and exhibit binding mid-points (KD) of 27(±3) μM (here as elsewhere in-text of this paper, we report 95% confidence intervals derived from replicate devices).
We began our study with high-level chemical disinfectants, rationalizing that, due to the mild treatment conditions they require, this class of treatments would be the least damaging to EAB sensors. Specifically, such treatments often utilize temperatures below 60 °C and ambient pressure, rendering them suitable for use with temperature- and moisture—sensitive medical devices, such as endoscopes. 10,11 Conveniently, these chemical techniques are also compatible with manual bench top processing in the laboratory (which is not true for, for example, gamma irradiation or ethylene oxide treatment, two approaches we did not explore). For each disinfectant we followed the manual processing protocols recommended by the Centers for Disease Control 10,12 to treat the gold working electrode (see Methods).
EAB sensors are compatible with disinfection using ortho-phthalaldehyde (OPA), a commercially available aromatic dialdehyde disinfectant (known as CIDEX OPA) that is commonly employed for the high-level disinfection of heat-sensitive medical devices in hospital settings. OPA, like other aldehydes, is thought to kill pathogens via the crosslinking of cellular amines, 13 and while OPA does not crosslink proteins as efficiently as glutaraldehyde, the aromatic component of OPA confers the ability to permeabilize pathogenic plasma membranes, promoting uptake of the compoud. 13,14 This uptake is also thought to block spore germination, 15 further contributing to the efficiency of OPA disinfection. Indeed, as a growing number of bacterial strains have become resistant to other reactive aldehydes, the use of CIDEX OPA as a high-level disinfectant has become commonplace. 16 Consistent with OPA's relatively mild chemical action, we observed negligible change in the sensor's gain (Fig. 2A), in its packing density (Fig. 2B), or in its target affinity (Fig. 2C) after we treat aptamer modified gold EAB working electrodes with this sterilant. This said, the working electrode is only one component of EAB sensors, which also include reference and working electrodes comprised of silver/silver chloride and platinum wires, respectively. When we performed the same CIDEX OPA disinfection on complete EAB sensors (i.e., a bundle including all three electrodes), we once again observe no significant change in sensor performance (Fig. 2D).
Figure 2. EAB sensors survive treatment with the disinfectant CIDEX OPA without any detectable degradation of their performance. (A) As shown, for example, both CIDEX OPA-treated and untreated control sensors exhibit similar signal gain (i.e., respond similarly to the addition of 20 μM vancomycin). Note, to remove fabrication variability and ease comparison we normalized all peaks to the peak heights seen in the absence of target in this and the following figures. (B) Treatment with CIDEX OPA does not affect the density with which methylene-blue-modified aptamers are packed on the surface of the sensor, indicating that treatment does not damage the aptamer, the self-assembled monolayer, or the redox reporter. (The difference in packing density— the number of aptamers per unit area— between the control and CIDEX OPA-treated sensors arises due to fabrication variability in these hand-made devices.) (C) Titration with the sensor's target, vancomycin, likewise shows that treatment with CIDEX OPA does not alter sensor performance. (D) Complete in vivo sensors, which are comprised of a gold-wire EAB sensor, an Ag/AgCl quasi-reference, and a platinum wire counter electrode, are similarly unaffected by CIDEX OPA treatment. The "error bars" shown in (C), (D) and in the following figures reflect standard deviations derived from multiple (three to six), independently fabricated sensors. In contrast, the confidence intervals noted for the dissociation constants in this and the following figures reflect 95% confidence intervals derived from the same, independently fabricated sensors. (E) CIDEX OPA treatment likewise does not harm sensor stability; shown are the responses of untreated and CIDEX-treated vancomycin sensors when interrogated 10 times per hour. The minor fluctuations seen reflect changes in room temperature over the course of this 15 h experiment.
Download figure:
Standard image High-resolution imageCIDEX OPA treatment likewise does not harm EAB sensor longevity. To see this, we challenged vancomycin-detecting sensors by interrogating them in target-free PBS 10 times per hour over the course of 15 h. We observe no significant degradation in either CIDEX-treated or control sensors over the course of these experiments (Fig. 2E).
The compatibility of EAB sensors with CIDEX OPA disinfection appears to be general. To show this, we subjected EAB sensor working electrodes employing two other aptamers to such treatment, one employing a 38-base DNA aptamer for the detection of phenylalanine, 17–19 and a second employing a 26-base DNA aptamer for the detection of the aminoglycoside antibiotics. 2,20,21 Once again, both sensors survive CIDEX OPA treatment without suffering any statistically significant degradation (Fig. 3).
Figure 3. The compatibility of CIDEX OPA disinfection with EAB sensor function extends to (A) phenylalanine-detecting and (B) aminoglycoside antibiotic-detecting EAB sensors.
Download figure:
Standard image High-resolution imageThe disinfection efficiency of the CIDEX OPA treatment we employed surpass Centers for Disease Control (CDC) requirements, which call for a six orders of magnitude reduction in bacterial load. 22,23 To see this, we incubated EAB sensors in 0.1 ml of media inoculated with 10,000 E. coli 24 at 37 °C for 24 h. Following this, we immersed these "contaminated" sensors in either CIDEX OPA or sterile phosphate-buffered saline (PBS) for 12 min at room temperature before moving the treated sensors into fresh, sterile media overnight at 37 °C to regrow any remaining bacteria. Plating serial dilutions of this wash media and monitoring colony growth, we found that treatment by CIDEX OPA results in a reduction of bacteria by more than seven orders of magnitude relative to the PBS control (Fig. 4), indicating high-level disinfection that exceeds the minimum requirements set by the CDC.
Figure 4. CIDEX OPA treatment achieves CDC-mandated levels of disinfection, which require a reduction in pathogenic load of at least six orders of magnitude. 22,23 (A) Specifically, while E. coli growth is present in plated cultures originating from control-treated wash media at dilutions from 103 to 109, we observe no detectable bacterial growth in the cultures from CIDEX OPA-treated wash media (for the complete dilution series, see Fig. S1 (available online at stacks.iop.org/ECSSP/1/011604/mmedia)). (B) E. coli growth challenge exhibits a decrease in the number of colony-forming units (CFU) per milliliter from 4 × 108 to fewer than 40 after CIDEX OPA treatment, demonstrating a reduction in bacterial load of at least seven orders of magnitude.
Download figure:
Standard image High-resolution imageTreatment with ethanol, a commonly used contact antiseptic, is also compatible with EAB sensor function. Specifically, we find that treatment by 70% ethanol does not significantly affect the vancomycin sensor's performance (Fig. 5). The CDC has not, however, cleared ethanol for clinical disinfection use due to its lack of sporicidal activity. 25 Consistent with this, cultures of E. coli that have been plated from the wash media of EAB sensors treated by 70% ethanol grew into dense, punctiform cultures (Fig. S2).
Figure 5. (A) EAB sensor function is compatible with sterilization by ethanol; we observe no significant difference in the gain of sensors treated in 70% ethanol for 10 min relative to the performance of untreated controls. (B) Likewise, ethanol treatment does not significantly affect aptamer packing density or (C) target affinity. Unfortunately, however, ethanol is not an effective disinfectant (Fig. S2).
Download figure:
Standard image High-resolution imageIn contrast to CIDEX OPA or ethanol disinfection, EAB sensors are incompatible with treatment by the reactive bis-aldehyde glutaraldehyde. Specifically, although this compound, which works to reduce pathogenic loads through aminic crosslinking, 26 is not expected to react with nucleic acids at the 21 °C temperature employed here, 27 treatment with 2% glutaraldehyde nevertheless significantly degrades sensor performance. As a result of such treatment, for example, the gain of our vancomycin-detecting sensor (the relative change in signal upon binding 20 μM of target) is decreased by two thirds (Fig. 6A) despite leaving the number of methylene blues on the sensor surface unchanged (Fig. 6B). Glutaraldehyde treatment likewise shifts the sensor's binding curve to significantly higher target concentrations (Fig. 6C). Given these observations, we speculate that glutaraldehyde treatment leads to DNA modification, such that many of the aptamers do not bind their target at all (reducing gain) and those that do bind do so with poorer affinity.
Figure 6. Treatment with 2% glutaraldehyde significantly degrades EAB sensor gain. (B) It does not, however, significantly affect aptamer packing density. (C) Along with reducing gain, glutaraldehyde treatment also reduces the sensor's affinity for its target.
Download figure:
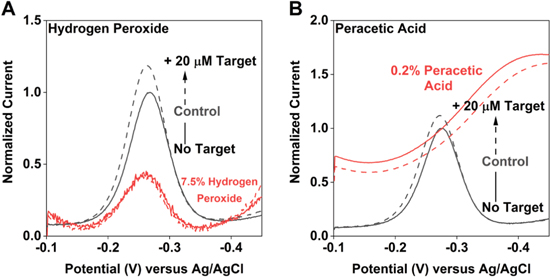
Standard image High-resolution imageTreatment with disinfecting levels of hydrogen peroxide or sterilizing levels of peracetic acid (treatment conditions specified 12 by the CDC) profoundly damage EAB sensors. Treatment with 7.5% hydrogen peroxide, for example, reduces the vancomycin sensor's peak current by 94% and entirely eliminates the response to its target (Fig. 7A). Treatment with 0.2% peracetic acid likewise leads to complete loss of the methylene blue peak and the appearance of an oxygen reduction peak at approximately −0.4 V. The latter indicates that oxygen is reaching the surface of the electrode, suggesting that the loss of methylene blue is due to loss of the self-assembled monolayer. This is perhaps not unexpected, given that peracetic acid is an even stronger oxidant than peroxide. 28,29
Figure 7. Treatment with either (A) 7.5% hydrogen peroxide or (B) 0.2% peracetic acid, two commonly employed disinfectants, gravely damages EAB sensors.
Download figure:
Standard image High-resolution imageDue to the ubiquity with which it is used for in-lab processing, we also explored the treatment of EAB sensors with steam sterilization. Perhaps unsurprisingly, this technique, which requires high temperatures, pressures, and long exposure times, significantly reduces EAB signaling currents (Figs. 8, S3). However, under "wet" conditions, wherein we submerge the working electrode in PBS, steam sterilization does not alter other key aspects of their performance. Specifically, while the peak currents associated with sensors autoclaved at 121 °C for 5 min while immersed in PBS are reduced significantly (Fig. 8B), their gain and target affinity remained largely unaffected (Fig. 8C). This suggests that such sterilization could prove useful in applications for which reduced signaling current (and, with that, reduced signal-to-noise ratios) is acceptable. In contrast, sensors treated under dry autoclaving conditions (i.e., exposed directly to steam) no longer respond measurably to their target (Fig. S3).
Figure 8. Autoclaving an EAB sensor emersed in PBS (A) does not significantly alter its gain. (B) It does, however, reduce the amount of signal-generating aptamer on the electrode surface by a factor of six. (C) Wet autoclaving likewise does not alter the sensor's affinity for its target.
Download figure:
Standard image High-resolution imageConclusions
EAB sensors are compatible with CIDEX OPA treatment sufficient to reduce bacterial loads by more than seven orders of magnitude, a level of disinfection that exceeds CDC guidelines for in vivo devices. Coupled with the generalizability of this disinfectant to other target-recognizing EAB sensors, this compatibility sets the stage for the use of EAB sensors in both long duration in vivo animal studies and, ultimately, in human patients. Ethanol, while compatible with EAB sensor functionality, does not meet the regulatory guidelines for disinfection of medical devices. In contrast, peroxide and peracetic acid treatments effectively destroy EAB signaling. Finally, while glutaraldehyde treatment and wet steam sterilization reduce functionality, EAB sensors nevertheless respond reproducibly to their targets after such treatments.
Experimental
Materials
As our primary testbed, we fabricated EAB sensors employing a 4-base truncation of a previously reported vancomycin-binding aptamer (5'-SS-(CH2)6- CGA GGG TAC CGC AAT AGT ACT TAT TGT TCG CCT ATT GTG GGT CGG-methylene blue-3', IDT). In our additional studies of the utility of CIDEX OPA treatment, we employed sensors fabricated with previously reported aptamers binding phenylalanine (5'-CGA CCG CGT TTC CCA AGA AAG CAA GTA TTG GTT GGT CG-3', Biosearch Technologies) and the aminoglycosides (5'-HO-(CH2)6-SS-(CH2)6-GGG ACT TGG TTT AGG TAA TGA GTC CC-O-CH2-CHCH2OH-(CH2)4-NH-CO-(CH2)2-methylene blue-3', Biosearch Technologies).
All reagents were obtained commercially and used as received. 1X Phosphate Buffered Saline at pH 7.4 containing 2 mM magnesium chloride was used in sensor fabrication and subsequent characterization assays. We prepared this buffer using a stock of 20X Phosphate Buffered Saline (ChemCruz, SC-362299) and 1 M magnesium chloride (Boston BioProducts, BM-670). CIDEX OPA (Advanced Sterilization Products) was used as received (i.e., without dilution). Other chemical disinfectants were diluted to their working concentrations (Table I) using high purity Milli-Q water (18.2 MΩ) from stocks of 30% hydrogen peroxide (Fisher Chemical, H325), 25% glutaraldehyde (Sigma-Aldrich, G6257), and 32% peracetic acid (Sigma-Aldrich, 269336).
Table I. Sterilizing and disinfecting techniques and conditions.
| Technique | Treatment Parameters |
|---|---|
| OPA, 0.55% (CIDEX OPA) | 12 min at 20 to 25 °C |
| Ethanol, 70% | 30 min at 20 to 25 °C |
| Glutaraldehyde, 2.0% | 90 min at 21 °C |
| Hydrogen peroxide, 7.5% | 30 min at 20 to 25 °C |
| Peracetic acid, 0.2% | 12 min at 55 °C |
| Steam sterilization | 5 min at 121 °C |
Sensor fabrication
We prepared gold wire working electrodes as previously described. 30,31 In brief, we soldered a length of 0.2 mm diameter gold wire to a gold-plated pin connector with 60/40 lead-selenium solder. We encased the electrode by shrink-wrapping poly-tetrafluoroethylene insulation (Zeus, HS Sub-Lite-Wall) around the gold surface, leaving a 6.5 mm length of bare gold surface. The bare gold surface was then trimmed to 6 mm for aptamer deposition. The construct underwent electro-chemical cleaning prior to aptamer modification. For this, we cycled the working electrode in 0.5 M NaOH from −1 V to −1.8 V at a scan rate of 1 V s−1 for 500 cycles to remove any contaminating organics. We next electrochemically roughened the electrodes in 0.5 M H2SO4 to increase the surface area using a previously described procedure 32 by stepping the potential between 0 and 2.2 V. By monitoring the gold oxide reduction peak in 0.05 M H2SO4 using cyclic voltammetry, we found the electroactive electrode area.
Following this, we modified the electrode with probe aptamer, as detailed. We thawed a stock solution of the aptamer (0.1 mM) from storage at −20 °C. We reduced any present disulphide bonds using 6 mM tris(2-carboxyethyl) phosphine hydrochloride (Sigma Aldrich) for 1 h under dark conditions, then diluted the aptamer mixture to a concentration of 500 nM in 1X PBS with 2 mM MgCl2. Immediately after roughening, we immersed the cleaned electrodes in the aptamer solution for 1 h for probe immobilization under dark conditions at room temperature. We further passivated the surface after aptamer deposition by immersion in 35 mM 6-mercapto-1-hexanol (Sigma Aldrich) suspended in 1X PBS for 10 to 12 h.
For in vivo sensors, we prepared the gold working electrode as above and sintered silver/silver chloride reference electrodes as follows. A length of 0.2 mm silver wire was soldered to a gold-plated pin connector with 60/40 lead-selenium solder. We encased the electrode by shrink-wrapping insulation around the silver surface, leaving approximately 1 cm of bare silver wire. We then submerged the exposed surface to sodium hypochlorite (commercial laundry bleach) overnight, covered. These reference electrodes were then thoroughly rinsed with water and submerged in 1X PBS buffer for a minimum of 1 h prior to use.
Before subjecting our working electrodes to the treatment conditions under investigation, we first ensured they were working properly by collecting 50 scans of square-wave data at 100 Hz using the parameters described above. Doing this, we confirmed that our methylene blue peak was (i) observable and (ii) not drifting significantly (the latter indicates poor SAM formation).
To determine aptamer packing density, 32 we collected cyclic voltammograms between −0.1 and −0.45 V at a scan rate of 0.1 V s−1 to reduce all methylene blue reporters present on the sensor surface. We integrate the area under the reductive curve and divide by scan rate to obtain the total charge associated with reporter-modified reporters, then convert this charge to moles of reporter present. We obtain packing density by dividing the number of methylene blue moieties by the electroactive surface area.
Electrochemical interrogation
We performed all electrochemical interrogation of our electrodes on a CH Instrument Multipotentiostat CHI1040C (Austin, TX). We used a three-electrode setup to collect measurements in which our sensors served as the working electrode, a silver/silver chloride aqueous electrode in saturated KCl (CH Instruments) served as the reference electrode, and a platinum wire (CH Instruments) completed the cell as the counter electrode. To interrogate our electrochemical cell, we applied a square-wave potential of 25 mV amplitude between −0.1 V and −0.45 V. We collected data at several square-wave frequencies over the range 5 Hz and 1 kHz but show here only the data associated with the frequency at which the sensor's signal gain is highest. For vancomycin, phenylalanine, and aminoglycosides sensors in PBS, these frequencies are 60 Hz, 120 Hz, and 300 Hz, respectively. All measurements were conducted at room temperature.
Sterilization and disinfection treatments
We applied sterilizing and disinfecting treatments to EAB sensors as follows. We manually processed chemical treatments per the conditions indicated (Table I). For steam sterilization, we used a Tomy ES-315 autoclave (San Diego, CA). After treatment, we submerged electrodes in water for 5 min, then in PBS for 5 min, and finally in the measurement cell containing PBS for 30 min. We then electrochemically interrogated the sensors again.
Determining treatment efficiency
We determined the treatment efficiency of the CIDEX OPA and ethanol treatment. To do so, we placed EAB sensors in 0.1 ml of sterile growth media (Difco SOB Media) inoculated with 10,000 bacterial cells (E. coli BL21) for 24 h at 37 °C and with 200 rpm shaking. We then subjected these "contaminated" sensors to either the treatment under investigation or incubated them in sterile PBS under the equivalent conditions. We then moved the treated sensors to fresh, sterile media for an overnight "wash" and regrowth at 37 °C and 250 rpm. We then plated serial dilutions of this overnight liquid culture at a volume of 0.25 ml on agar plates (1% NaCl, 1% tryptone, 0.5% yeast extract, and 1.5% agar). We monitored these plates for colony growth after overnight incubation at 37 °C.
Acknowledgments
This work was funded by NIH grant R01AI145206. We thank Dr. Kaylyn K. Leung and Lisa Fetter for technical insights in the fabrication and testing of in vivo probes.