Abstract
Background
Machine learning has been increasingly used for surgical outcome prediction, yet applications in head and neck reconstruction are not well-described. In this study, we developed and evaluated the performance of ML algorithms in predicting postoperative complications in head and neck free-flap reconstruction.
Methods
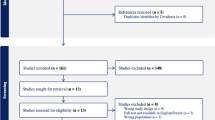
We conducted a comprehensive review of patients who underwent microvascular head and neck reconstruction between January 2005 and December 2018. Data were used to develop and evaluate nine supervised ML algorithms in predicting overall complications, major recipient-site complication, and total flap loss.
Results
We identified 4000 patients who met inclusion criteria. Overall, 33.7% of patients experienced a complication, 26.5% experienced a major recipient-site complication, and 1.7% suffered total flap loss. The k-nearest neighbors algorithm demonstrated the best overall performance for predicting any complication (AUROC = 0.61, sensitivity = 0.60). Regularized regression had the best performance for predicting major recipient-site complications (AUROC = 0.68, sensitivity = 0.66), and decision trees were the best predictors of total flap loss (AUROC = 0.66, sensitivity = 0.50).
Conclusions
ML accurately identified patients at risk of experiencing postsurgical complications, including total flap loss. Predictions from ML models may provide insight in the perioperative setting and facilitate shared decision making.





Similar content being viewed by others
References
Rajkomar A, Dean J, Kohane I. Machine learning in medicine. N Engl J Med. 2019;380(14):1347–58.
Bur AM, Shew M, New J. Artificial intelligence for the otolaryngologist: a state of the art review. Otolaryngol Head Neck Surg. 2019;160(4):603–11.
Sidey-Gibbons JAM, Sidey-Gibbons CJ. Machine learning in medicine: a practical introduction. BMC Med Res Methodol. 2019;19(1):64.
AI: Healthcare's new nervous system. Accenture. Jul 30, 2020. Available at: https://www.accenture.com/au-en/insights/health/artificial-intelligence-healthcare Accessed on Dec 27, 2020.
Ascent of machine learning in medicine. Nat Mater. 2019;18(5):407.
Chen CL, Mahjoubfar A, Tai L-C, et al. Deep learning in label-free cell classification. Sci Rep. 2016;6(1):21471.
Nagendran M, Chen Y, Lovejoy CA, et al. Artificial intelligence versus clinicians: systematic review of design, reporting standards, and claims of deep learning studies. BMJ. 2020;368:m689.
Liu X, Faes L, Kale AU, et al. A comparison of deep learning performance against health-care professionals in detecting diseases from medical imaging: a systematic review and meta-analysis. Lancet Digit Health. 2019;1(6):e271–97.
Thomsen K, Iversen L, Titlestad TL, Winther O. Systematic review of machine learning for diagnosis and prognosis in dermatology. J Dermatolog Treat. 2020;31(5):496–510.
Senders JT, Arnaout O, Karhade AV, et al. Natural and artificial intelligence in neurosurgery: a systematic review. Neurosurgery. 2018;83(2):181–92.
Senders JT, Staples PC, Karhade AV, et al. Machine learning and neurosurgical outcome prediction: a systematic review. World Neurosurg. 2018;109:476-86.e471.
Johnson KW, Torres Soto J, Glicksberg BS, et al. Artificial intelligence in cardiology. J Am Coll Cardiol. 2018;71(23):2668–79.
Chauhan J, Goyal P. BPBSAM: body part-specific burn severity assessment model. Burns. 2020;46(6):1407–23.
Cirillo MD, Mirdell R, Sjöberg F, Pham TD. Time-independent prediction of burn depth using deep convolutional neural networks. J Burn Care Res. 2019;40(6):857–63.
Yadav DP, Sharma A, Singh M, Goyal A. Feature extraction based machine learning for human burn diagnosis from burn images. IEEE J Transl Eng Health Med. 2019;7:1800507.
Jiao C, Su K, Xie W, Ye Z. Burn image segmentation based on Mask Regions with Convolutional Neural Network deep learning framework: more accurate and more convenient. Burns Trauma. 2019;7:6.
Cobb AN, Daungjaiboon W, Brownlee SA, et al. Seeing the forest beyond the trees: predicting survival in burn patients with machine learning. Am J Surg. 2018;215(3):411–6.
Huang Y, Zhang L, Lian G, et al. A novel mathematical model to predict prognosis of burnt patients based on logistic regression and support vector machine. Burns. 2016;42(2):291–9.
Yeong EK, Hsiao TC, Chiang HK, Lin CW. Prediction of burn healing time using artificial neural networks and reflectance spectrometer. Burns. 2005;31(4):415–20.
Bhalodia R, Dvoracek LA, Ayyash AM, Kavan L, Whitaker R, Goldstein JA. Quantifying the severity of metopic craniosynostosis: a pilot study application of machine learning in craniofacial surgery. J Craniofac Surg. 2020;31(3):697–701.
Angullia F, Fright WR, Richards R, Schievano S, Linney AD, Dunaway DJ. A novel RBF-based predictive tool for facial distraction surgery in growing children with syndromic craniosynostosis. Int J Comput Assist Radiol Surg. 2020;15(2):351–67.
Mendoza CS, Safdar N, Okada K, Myers E, Rogers GF, Linguraru MG. Personalized assessment of craniosynostosis via statistical shape modeling. Med Image Anal. 2014;18(4):635–46.
Formeister EJ, Baum R, Knott PD, et al. Machine learning for predicting complications in head and neck microvascular free tissue transfer. Laryngoscope. 2020;130(12):E843–9.
Kuo PJ, Wu SC, Chien PC, et al. Artificial neural network approach to predict surgical site infection after free-flap reconstruction in patients receiving surgery for head and neck cancer. Oncotarget. 2018;9(17):13768–82.
Zhao EH, Nishimori K, Brady J, et al. Analysis of risk factors for unplanned reoperation following free flap surgery of the head and neck. Laryngoscope. 2018;128(12):2790–5.
Zhou W, Zhang WB, Yu Y, et al. Risk factors for free flap failure: a retrospective analysis of 881 free flaps for head and neck defect reconstruction. Int J Oral Maxillofac Surg. 2017;46(8):941–5.
Carniol ET, Marchiano E, Brady JS, et al. Head and neck microvascular free flap reconstruction: an analysis of unplanned readmissions. Laryngoscope. 2017;127(2):325–30.
Maricevich M, Lin LO, Liu J, Chang EI, Hanasono MM. Interposition vein grafting in head and neck free flap reconstruction. Plast Reconstr Surg. 2018;142(4):1025–34.
Tang Z-H, Liu J, Zeng F, Li Z, Yu X, Zhou L. Comparison of prediction model for cardiovascular autonomic dysfunction using artificial neural network and logistic regression analysis. PLoS One. 2013;8(8):e70571.
Jaimes F, Farbiarz J, Alvarez D, Martínez C. Comparison between logistic regression and neural networks to predict death in patients with suspected sepsis in the emergency room. Crit Care. 2005;9(2):R150–6.
Tu JV. Advantages and disadvantages of using artificial neural networks versus logistic regression for predicting medical outcomes. J Clin Epidemiol. 1996;49(11):1225–31.
Chen JH, Asch SM. Machine learning and prediction in medicine: beyond the peak of inflated expectations. N Engl J Med. 2017;376(26):2507–9.
Collins GS, Reitsma JB, Altman DG, Moons KGM. Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): the TRIPOD Statement. BMC Medicine. 2015;13(1):1.
Yap B, Rani KA, Rahman HA, Fong S, Khairudin Z, Abdullah NN. An application of oversampling, undersampling, bagging and boosting in handling imbalanced datasets. Paper presented at: DaEng2013.
Pfob A, Sidey-Gibbons C, Tasoulis MK, et al. Artificial intelligence to accurately identify breast cancer patients with a pathologic complete response for omission of surgery after neoadjuvant systemic therapy: An international multicenter analysis. J Clin Oncol. 2020;38(15_suppl):565.
Parikh RB, Manz C, Chivers C, et al. Machine learning approaches to predict 6-month mortality among patients with cancer. JAMA Netw Open. 2019;2(10):e1915997.
Li C, Zhang S, Zhang H, et al. Using the K-nearest neighbor algorithm for the classification of lymph node metastasis in gastric cancer. Comput Math Methods Med. 2012;2012:876545.
Menon R, Bhat G, Saade GR, Spratt H. Multivariate adaptive regression splines analysis to predict biomarkers of spontaneous preterm birth. Acta Obstet Gynecol Scand. 2014;93(4):382–91.
R Core Team. R: A language and environment for statistical computing. Published online 2020. Available at: https://www.r-project.org/ Accessed Jan 12, 2021.
Bertsimas D, Dunn J, Velmahos GC, Kaafarani HMA. Surgical risk is not linear: Derivation and validation of a novel, user-friendly, and machine-learning-based Predictive OpTimal Trees in Emergency Surgery Risk (POTTER) Calculator. Ann Surg. 2018;268(4):574–83.
Hassan AM, Rajesh A, Asaad M, et al. A surgeon's guide to artificial intelligence-driven predictive models. Am Surg. 2022:31348221103648.
Hassan AM, Rajesh A, Asaad M, et al. Artificial intelligence and machine learning in prediction of surgical complications: Current state, applications, and implications. Am Surg. 2022:31348221101488.
Hassan AM, Lu SC, Asaad M, et al. Novel machine learning approach for the prediction of hernia recurrence, surgical complication, and 30-day readmission after abdominal wall reconstruction. J Am Coll Surg. 2022;234(5):918–27.
Hassan AM, Biaggi AP, Asaad M, et al. Development and assessment of machine learning models for individualized risk assessment of mastectomy skin flap necrosis. Ann Surg. 2022.
O’Neill AC, Yang D, Roy M, Sebastiampillai S, Hofer SOP, Xu W. Development and evaluation of a machine learning prediction model for flap failure in microvascular breast reconstruction. Ann Surg Oncol. 2020;27(9):3466–75.
Bzdok D, Altman N, Krzywinski M. Statistics versus machine learning. Nat Methods. 2018;15(4):233–4.
Breiman L. Statistical modeling: the two cultures. Stat Sci. 2001;16:199–231.
Trevethan R. Sensitivity, specificity, and predictive values: foundations, pliabilities, and pitfalls in research and practice. Front Public Health. 2017;5:307.
Mallett S, Halligan S, Thompson M, Collins GS, Altman DG. Interpreting diagnostic accuracy studies for patient care. BMJ. 2012;345:e3999.
Funding
No direct or indirect financial support was used for this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosures
None of the authors has a financial interest in any of the products, devices, or drugs mentioned in this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Asaad, M., Lu, SC., Hassan, A.M. et al. The Use of Machine Learning for Predicting Complications of Free-Flap Head and Neck Reconstruction. Ann Surg Oncol 30, 2343–2352 (2023). https://doi.org/10.1245/s10434-022-13053-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-022-13053-3