Abstract
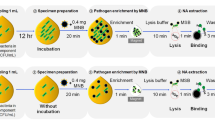
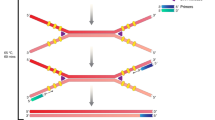
In the present study, we aimed to develop a nucleic acid lateral-flow method for the rapid and sensitive detection of multiple bacteria that contaminate platelet concentrations (PCs). Polymerase chain reaction (PCR) amplicons were produced by a set of board-range primers that recognize the conserved region of bacteria 16S rDNA, followed by hybridization with both an FITC (fluorescein isothiocyanate)-labelled probe and biotin-labelled probe, and then a nucleic acid lateral-flow dipstick (LFD) assay. The LFD accurately identified 7 species of bacteria, but had no cross-reactivity with human genomic DNA. The limit of detection (LOD) of the LFD assay was as low as 101 copies/μL of 16S rDNA for plasmid. In the case of spiked PCs without enrichment, the detection limit of LFD for K. pneumonia was 5 CFU/mL, 6.5 × 104 CFU/mL for the S. epidermidis and 35 CFU/mL for P. aeruginosa.
Similar content being viewed by others
References
M. T. Sullivan and E. L. Wallace, Transfusion, 2005, 45, 141.
M. E. Brecher and S. N. Hay, Clin. Microbiol. Rev., 2005, 18, 195.
R. N. Pietersz, C. P. Engelfriet, H. W. Reesink, E. M. Wood, S. Winzar, A. J. Keller, J. T. Wilson, G. Henn, W. R. Mayr, S. Ramirez-Arcos, M. Goldman, J. Georgsen, P. Morel, P. Herve, G. Andeu, A. Assal, E. Seifried, M. Schmidt, M. Foley, C. Doherty, P. Coakley, A. Salami, E. Cadden, W. G. Murphy, M. Satake, D. de Korte, V. Bosnes, J. Kjeldsen-Kragh, C. McDonald, M. E. Brecher, R. Yomtovian, and J. P. AuBuchon, Vox Sang, 2007, 93, 260.
H. W. Reesink, T. Mohammadi, R. N. Pietersz, and P. H. Savelkoul, Clin. Chem. Lab. Med., 2008, 46, 954.
I. G. Rood, M. H. Koppelman, A. Pettersson, and P. H. Savelkoul, J. Microbiol. Methods, 2008, 75, 64.
S. Philipp, H. P. Huemer, E. U. Irschick, and C. Gassner, Transfus. Med. Hemother., 2010, 37, 21.
Q. Qu, Z. Zhu, Y. Wang, Z. Zhong, J. Zhao, F. Qiao, X. Du, Z. Wang, R. Yang, L. Huang, Y. Yu, L. Zhou, and Z. Chen, J. Microbiol. Methods, 2009, 79, 121.
C. Y. Yu, G. Y. Ang, A. L. Chua, E. H. Tan, S. Y. Lee, G. Falero-Diaz, O. Otero, I. Rodriguez, F. Reyes, A. Acosta, M. E. Sarmiento, S. Ghosh, T. Ramamurthy, C. Yean Yean, P. Lalitha and M. Ravichandran, J. Microbiol. Methods, 2011, 86, 277.
D. P. Kalogianni, L. M. Boutsika, P. G. Kouremenou, T. K. Christopoulos, and P. C. Ioannou, Anal. Bioanal. Chem., 2011, 400, 1145.
P. Noguera, G. A. Posthuma-Trumpie, M. Tuil, F. J. Wal, A. Boer, A. P. H. A. Moers, and A. Amerongen, Anal. Bioanal. Chem., 2010, 399, 831.
J. H. Miller, “Experiments in Molecular Genetics”, 1972, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
M. A. Nadkarni, F. E. Martin, N. A. Jacques, and N. Hunter, Microbiology, 2002, 148, 257.
C. C. Mastronardi and S. Ramirez-Arcos, Can. J. Microbiol., 2007, 53, 1222.
Y. Yanling, T. Sudan, H. Yanmin, X. Xianguo, Z. Faming, L. Hangjun, and Y. Lixing, Chin. J. Nosocomiol., 2011, 21, 205.
W. Yongzhi, X. Lijun, R. Hao, Z. Ping, Z. Shiying, L. Mian, and Q. Zhongtian, Chin. J. Exp. Clin. Infect. Dis., 2007, 1, 204.
I. G. Rood, A. Pettersson, P. H. Savelkoul, and D. de Korte, Transfusion, 2010, 50, 1352.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Wang, J., Wang, X., Li, Y. et al. A Novel, Universal and Sensitive Lateral-Flow Based Method for the Detection of Multiple Bacterial Contamination in Platelet Concentrations. ANAL. SCI. 28, 237–241 (2012). https://doi.org/10.2116/analsci.28.237
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2116/analsci.28.237