Abstract
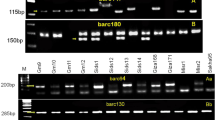
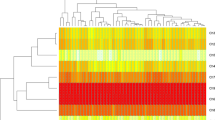
Bread wheat (Triticum aestivum L.) germplasm consisting of 45 genotypes were clustered phenotypically using ten morphological traits and Area Under Disease Progress Curve (AUDPC) as measure of stripe rust resistance. The clustering was ratified by using twenty three molecular markers (SSR, EST and STS) linked to stripe rust (Puccinia striiformis f. sp. tritici) resistant QTLs. The aim was to asses the extent of genetic variability among the genotypes in order to select the parents for crossing between the resistant and susceptible genotypes with respect to stripe rust. The Euclidian dissimilarity values resulted from phenotypic data regarding morphological traits and AUDPC were used to construct a dendrogram for clustering the accessions. Using un-weighted pair group method with arithmetic means, another dendrogram resulted from the similarity coefficient values was used to distinguish the genotypes with respect to stripe rust. Clustering based on phenotypic data produced two major groups and five clusters (with Euclidian dissimilarity ranging from 2.44 to 16.16) whereas genotypic data yielded two major groups and four clusters (with percent similarity coefficient values ranging from 0.1 to 46.0) to separate the gene pool into highly resistant, resistant, moderately resistant, moderately susceptible and susceptible genotypes. With few exceptions, the outcome of both type of clustering was almost similar and resistant as well as susceptible genotypes came in the same clusters of molecular genotyping as yielded by phenotypic clustering. As a result seven genotypes (Bakhtawar-92, Frontana, Saleem 2000, Tatara, Inqilab-91, Fakhre Sarhad and Karwan) of diverse genetic background were selected for pyramiding stripe rust lesistant genes as well as some other agronomic traits after hybridization.
Similar content being viewed by others
References
Fahima, T., Röder, M.S., Grama, A., Nevo, E., Microsatellite DNA Polymorphism Divergence in Triticum dicoccoides Accessions Highly Resistant to Yellow Rust, Theor. Appl. Genet., 1998, vol. 96, pp. 187–195.
Imtiaz, M., Cromey, M.G., Hampton, J.G., and Hill, M.J., Inheritance of Seedling Resistance to Stripe Rust (Puccinia striifomis f. sp. tritici) in ‘Otan’ and ‘Tritea’ Wheat (Triticum aestivum), New Zealand J. Crop Hort. Sci., 2003, vol. 31, pp. 15–22.
Wang, X., Zheng, W., Buchenauer, H., Zhao, J., Han, Q., Huang, L., and Kang, Z., The Development of a PCR-Based Method for Detecting Puccinia striiformis Latent Infections in Wheat Leaves, Eur. J. Plant Pathol., 2008, vol. 120, pp. 241–247.
Irfaq, M., Mir, A.K., Khattak, G.S.S., Tila, M., and Shah, S.J.A., Genetic Behavior of Controlling Area Under Disease Progress Curve for Stripe Rust (Puccinia striiformis f. sp. tritici) in Two Wheat (Triticum aestivum) Crosses, Phytopathology, 2009, vol. 99, pp. 1265–1272.
Smith, S.E., Doss, A.A., and Warburton, M., Morphological and Agronomic Variation in North Africa and Arabian Alfalfas, Crop Sci., 1991, vol. 31, pp. 1159–1163.
Smith, J.S.C. and Smith, O.S., The Description and Assessment of Distances Between Inbreed Lines of Maize. The Utility of Morphological, Biochemical and Genetic Descriptions and a Scheme of Testing of Distinctiveness Between Inbreed Lines, Maydica, 1989, vol. 34, pp. 151–161.
Ashraf, M., Cheema, A.A., Qamar, Z., and Rashid, M., Detection of Polymorphism in Rice Germplasm using RAPD Markers, Pak. J. Bot., 2007, vol. 39, pp. 2483–2493.
Peter, M., Darcatos, L.J., Dumsday, S., Rhiannon, N., Olle, O.I., Noel, O., Cogan, P.M., Dobrowobki, M., Fujimori, H., Roderick, V.S., Alan, W.S., Smith, K.F., and John, W.F., Development and Characterization of EST-SSR Markers for the Crown Rust Pathogen of Ryegrass (Puccinia coronata f. sp. lollii), Genome, 2006, vol. 49, pp. 572–583.
Squirrel, J., Hollingsworth, P.M., Woodhead, M., Russell, J., Lowe, A.J, Gibby, M., and Powel, W., How Much Effort Is Required to Isolate Nuclear Microsatellits from Plants?, Mol. Ecol., 2003, vol. 12, pp. 1339–1348.
Zadoks, J.C., Chang, T.T., and Konzak, C.F., A Decimal Code for the Growth Stages of Cereals, Weed Res., 1974, vol. 14, pp. 415–421.
Rattu, A.R., Akhtar, M.A., Fayyaz, M., and Bashir, M., Report on Screening of Wheats Against Yellow Rusts and Leaf Rusts under NUWYT and NWDSN during 2005–2006, Islamabad, Pakistan: Crop Diseases Research Programme, Institute of Plant and Environmental Protection, National Agriculture Research Centre, Pakistan Agriculture Research Council, 2006, pp. 13–14.
Peterson, R.F., Campbell, A.B., and Hannah, A.E., A Diagnostic Scale for Estimating Rust Intensity on Leaves and Stems of Cereals, Can. J. Res., 1948, vol. 26, pp. 496–500.
Line, R.F., Konzak, C.F., and Alan, R.E., Evaluating Resistance to Puccinia striiformis in Wheat. Induced Mutation for Disease Resistance in Crop Plants, in Proceedings of a Research Co-ordination Meeting, IAEA, Austria, 1974, pp. 125–132.
Loegering, W.Q., Methods for Recording Cereal Rust Data, USDA International Spring Wheat Nursery, 1959.
Doling, D.A., A Method for the Transformation of Field Data for Comparing the Mildew Resistance of Cereal Varieties and the Systemic Deviation of the Values in NIAB Farmer’s Leaflets, J. Nat. Inst. Agric. Bot., 1965, vol. 10, pp. 169–179.
Aslam, M., Uniform Procedure for Development and Release of Improved Wheat Varieties, Mimeograph, PARC, Islamabad, 1982.
Arama, P.F., Parievliet, J.E., and Van Silfhout, C.H., Trangressive Segregation for Resistance in Wheat to Septoria tritici Blotch, Afric. Crop Sci. J., 2000, vol. 8, pp. 213–222.
Steel, R.J.D. and Torrie, J.H., Duncan’s New Multiple Range Test, in Principles and Procedures of Statistics, 2nd ed., Mc Graw Hill Book Comp. Inc., New York, 1980.
Maroof, M.A.S, Soliman, K., Jorgensen, R.A., and Allerd, R.W., Ribosomal DNA Spacer Length Lymorphism in Barley: Mendelian Inheritance, Chromosome Location and Population Dynamics, Proc. Natl. Acad. Sci. U.S.A., 1984, vol. 81, pp. 8014–8018.
NTSYSpc, Exeter Software for Windows, Version 2.02i: Copyright 1986-1998. Applied Biostatistics Inc., Setauket, New York, 1998.
Sixin, L. and Anderson, J.A., Marker Assisted Evaluation of Fusarium Head Blight Resistant Wheat Germplasm, Crop Sci., 2003, vol. 43, pp. 760–766.
McCartney, C.A., Somers, D.J., Fedak, G., and Cao, W., Haplotype Diversity at Fusarium Head Blight Resistance QTLs in Wheat, Theor. Appl. Genet., 2004, vol. 109, pp. 261–271.
Zhuping, Y., Gilbert, J., and Procunier, L.D., Genetic Diversity of Resistance Genes Controlling Fusarium Head Blight with Simple Sequence Repeat Markers in Thirty-Six Wheat Accessions from East Asian Origin, Euphytica, 2006, vol. 148, pp. 345–352.
Singh, R.P., and Rajaram, S., Genetics of Adult-Plant Resistance to Leaf Rust in ‘Frontana’ and Three CIMMYT Wheats, Genome, 1992, vol. 35, pp. 24–31.
McDonald, D.B., McIntosh, R.A., Wellings, C.R., Singh, R.P., and Nelson, J.C., Cytogenetical Studies in Wheat XIX. Location and Linkage Studies on Gene Yr27 for Resistance to Stripe (Yellow) Rust, Euphytica, 2004, vol. 136, pp. 239–248.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
About this article
Cite this article
Khan, M.I., Khan, M.A., Hongxiang, M. et al. Selection of parents for crossing based on genotyping and phenotyping for stripe rust (Puccinia striiformis) resistance and agronomic traits in bread wheat breeding. Cytol. Genet. 45, 379–394 (2011). https://doi.org/10.3103/S0095452711060065
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0095452711060065