Abstract
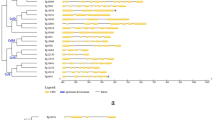
In silico search for the nucleotide sequences encoding cellulose synthases in the flax genome was carried out, together with the comparison of the identified sequences with the orthologous genes in dicotyledonous plants. The analysis resulted in the identification of 32 flax candidate genes, 16 of which encoded cellulose synthases with high possibility rate, while the remaining 16 encoded cellulose synthase-like proteins (Csl). The phylogenetic analysis of the protein products of the cellulose synthase genes allowed dividing them into six groups comprising cellulose synthases of different classes: CesA1/10, CesA3, CesA4, CesA5/6/2/9, CesA7, and CesA8. Paralogous sequences belonging to the classes CesA1/10 and CesA5/6/2/9, associated with the primary cell wall formation, were characterized by the higher intra-class similarity rate, than the orthologous sequences. At the same time, the genes that control cellulose biosynthesis for the secondary cell wall formation constituted distinct clades, CesA4, CesA7, and CesA8. The analysis of the 16 selected flax candidate cellulose synthase genes demonstrated that the flax genome contains at least 12 different variants of cellulose synthase genes, which belong to all six cellulose synthase clades. In such a way, at this stage, the cellulose synthase genes from all ten known CesA classes have been identified in the flax genome; however, their correct attribution to each of these classes requires some additional study.
Similar content being viewed by others
References
Brown, R.M., Cellulose structure and biosynthesis: what is in store for the 21st century?, J. Polym. Sci. A. Polym. Chem., 2004, vol. 42, pp. 487–495.
Kimura, S., Immunogold labeling of rosette terminal cellulose synthesizing complexes in the vascular plant Vigna angularis, Plant Cell, 1999, vol. 11, pp. 2075–2085.
Scheible, W.R., Eshed, R., Richmond, T., et al., Modifications of cellulose synthase confer resistance to isoxaben and thiazolidinone herbicides in Arabidopsis CFPxr1 mutants, Proc. Nat. Acad. Sci. USA, 2001, vol. 98, pp. 10079–10084.
Taylor, N.G., Howells, R.M., Huttly, A.K., et al., Interactions among three distinct cesa proteins essential for cellulose synthesis, Proc. Nat. Acad. Sci. USA, 2003, vol. 100, pp. 1450–1455.
Sethaphong, L., Haigler, C.H., Kubicki, J.D., et al., Tertiary model of a plant cellulose synthase, Proc. Nat. Acad. Sci. USA, 2013, vol. 110, no. 18, pp. 7512–7517.
Ross, P., Mayer, R., and Benziman, M., Cellulose biosynthesis and function in bacteria, Microbiol. Rev., 1991, vol. 55, no. 1, pp. 35–58.
Hamann, T., Osborne, E., Youngs, H.L., et al., Global expression analysis of CESA and CSL genes in Arabidopsis, Cellulose, 2004, vol. 11, pp. 279–286.
Holland, N., Holland, D., Helentjaris, T., et al., A comparative analysis of the plant cellulose synthase (CESA) gene family, Plant Physiol., 2000, vol. 123, pp. 1313–1324.
Ranik, M. and Myburg, A.A., Six new cellulose synthase genes from eucalyptus associated with primary and secondary cell wall biosynthesis, Tree Physiol., 2006, vol. 26, pp. 545–556.
Kumar, M., Thammannagowda, S., Bulone, V., et al., An update on the nomenclature for the cellulose synthase genes in Populus, Trends Plant Sci., 2009, vol. 14, no. 5, pp. 248–254.
Carroll, A. and Specht, C., Understanding plant cellulose synthases through a comprehensive investigation of the cellulose synthase family sequences, Front. Plant Sci., 2011, vol. 2, pp. 1–11.
Wang, Z., Hobson, N., Galindo, L., et al., The genome of flax (Linum usitatissimum) assembled de novo from short shotgun sequence reads, Plant J., 2012, vol. 72, pp. 461–473.
Henikoff, S. and Henikoff, J.G., Amino acid substitution matrices from protein blocks, Proc. Nat. Acad. Sci. USA, 1992, vol. 89, pp. 10915–10919.
Punta, M., Coggill, P.C., Eberhardt, R.Y., et al., The PFAM protein families database, Nucleic Acids Res., 2012, vol. 40, pp. D290–301.
Dereeper, A., Guignon, V., Blanc, G., et al., Phylogeny. fr: robust phylogenetic analysis for the non-specialist, Nucleic Acids Res., 2008, vol. 36 (Web Server issue), pp. W465–W469.
Edgar, R.C., MUSCLE: multiple sequence alignment with high accuracy and high throughput, Nucleic Acids Res., 2004, vol. 32, no. 5, pp. 1792–1797.
Castresana, J., Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis, Mol. Biol. Evol., 2000, vol. 17, pp. 540–552.
Guindon, S., Dufayard, J.F., Lefort, V., et al., New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0, System. Biol., 2010, vol. 59, no. 3, pp. 307–321.
Chevenet, F., Brun, C., Banuls, A.L., et al., TreeDyn: towards dynamic graphics and annotations for analyses of trees, MBC Bioinform., 2006, vol. 7, p. 439.
Larkin, M.A., Blackshields, G., Brown, N.P., et al., ClustalW and ClustalX version 2, Bioinformatics, 2007, vol. 23, no. 21, pp. 2947–2948.
Richmond, T.A. and Somerville, C.R., The cellulose synthase superfamily, Plant Physiol., 2000, vol. 124, pp. 495–498.
Hazen, S., Scott-Craig, J.S., and Walton, J.D., Cellulose synthase-like (CSL) genes of rice, Plant Physiol., 2002, vol. 128, pp. 336–340.
Fincher, G.B., Revolutionary times on our understanding of cell wall biosynthesis and remodeling in the grasses, Plant Physiol., 2009, vol. 149, pp. 27–37.
Sveinsson, S., McDill, J., Wong, G.K., et al., Phylogenetic pinpointing of a paleopolyploidy event within the flax genus (Linum) using transcriptomics, Ann. Bot., 2014, vol. 113, no. 5, pp. 753–761.
Mokshina, N., Gorshkova, T., and Deyholos, M.K., Chitinase-like (CTL) and cellulose synthase (CESA) gene expression in gelatinous-type cellulosic walls of flax (Linum usitatissimum L.) bast fibers, PLoS One, 2014, vol. 9, no. 2, p. e94949.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © N.A. Pydiura, G.Ya. Bayer, D.V. Galinousky, A.I. Yemets, Ya.V. Pirko, T.A. Padvitski, N.V. Anisimova, L.V. Khotyleva, A.V. Kilchevsky, Ya.B. Blume, 2015, published in Tsitologiya i Genetika, 2015, Vol. 49, No. 5, pp. 3–12.
About this article
Cite this article
Pydiura, N.A., Bayer, G.Y., Galinousky, D.V. et al. Bioinformatic search for cellulose synthase genes in flax (Linum usitatissimum) and their phylogenetic analysis. Cytol. Genet. 49, 279–287 (2015). https://doi.org/10.3103/S0095452715050084
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0095452715050084