Abstract
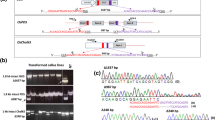
MuDR exhibits the highest transposition activity and insertional mutagenesis frequency in Mutator (Mu) family. If we isolate the MuDR-insertion-specific flanking sequences (MuDRFs), it will be crucial for using Mu element-mediated mutants. The MuDR-TAIL-PCR system was constructed and optimized using a combination of MuDR-TIR-nested specific primers and 12 arbitrary degenerate (AD) primers, modified reaction system and procedure and mutant DNA templates of 87 genotypes from M2 or М2:3 families created by crossing the W22::Mu line (active MuDR donor parent) from the UniformMu population with the Zong31 (Z31) line (recipient parent). Here 129 different MuDRFs were acquired by MuDR-TAIL-PCR, accounting for 86.60% of the total mutant-specific agarose gel bands. In addition, we confirmed the authenticity of the non-redundant flanking sequence amplifications. The amplified non-redundant flanking sequences accounted for 65.12% of the total MuDRFs, and 88.00% of the non-redundant MuDRFs were inserted inside the genes. These results show that the MuDR-TAIL-PCR system that we developed can be used for specifically isolating MuDRFs.
Similar content being viewed by others
References
McCarty, D.R., Settles, A.M., Suzuki, M., Tan, B.C., Latshaw, S., Porch, T., Robin, K., Baier, J., Avigne, W., Lai, J., Messing, J., Koch, K.E., and Hannah, L.C., Steady-state transposon mutagenesis in inbred maize, Plant J., 2005, vol. 44, no. 1, pp. 52–61.
Wang, Y., Yin, G., Yang, Q., Tang, J., Lu, X., Korban, S.S., and Xu, M., Identification and isolation of Mu-flanking fragments from maize, J. Genet. Genom., 2008, vol. 35, no. 4, pp. 207–213.
Muszynski, M.G., Dam, T., Li, B., Shirbroun, D.M., Hou, Z., Bruggemann, E., Archibald, R., Ananiev, E.V., and Danilevskaya, O.N., Delayed flowering1 encodes a basic leucine zipper protein that mediates floral inductive signals at the shoot apex in maize, Plant Physiol., 2006, vol. 142, no. 4, pp. 1523–1536.
Williams-Carrier, R., Stiffler, N., Belcher, S., Kroeger, T., Stern, D.B., Monde, R.A., Coalter, R., and Barkan, A., Use of Illumina sequencing to identify transposon insertions underlying mutant phenotypes in high-copy Mutator lines of maize, Plant J., 2010, vol. 63, no. 1, pp. 167–177.
Liu, S., Dietrich, C.R., and Schnable, P.S., DLA-based strategies for cloning insertion mutants: cloning the gl4 locus of maize using Mu transposon tagged alleles, Genetics, 2009, vol. 183, no. 4, pp. 1215–1225.
Settles, A.M., Latshaw, S., and McCarty, D.R., Molecular analysis of high-copy insertion sites in maize, Nucleic Acids Res., 2004, vol. 32, no. 6, p. e54.
Robbins, M.L., Sekhon, R.S., Meeley, R., and Chopra, S., A mutator transposon insertion is associated with ectopic expression of a tandemly repeated multicopy Myb gene pericarp color1 of maize, Genetics, 2008, vol. 178, no. 4, pp. 1859–1874.
Zhang, F. and Peterson, T., Comparisons of maize pericarp color1 alleles reveal paralogous gene recombination and an organ-specific enhancer region, Plant Cell, 2005, vol. 17, no. 3, pp. 903–914.
Liu, Y.G., Mitsukawa, N., Oosumi, T., and Whittier, R.F., Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR, Plant J., 1995, vol. 8, no. 3, pp. 457–463.
Liu, Y.G. and Huang, N., Efficient amplification of insert end sequences from bacterial artificial chromosome clones by thermal asymmetric interlaced PCR, Plant Mol. Biol. Report., 1998, vol. 16, no. 2, pp. 175–181.
Walbot, V. and Rudenko, G., MuDR/Mu transposable elements in maize, in Mobile DNA, Craig, N., Craigie, R., Gellert, M., and Lambowitz, A., Eds., Washington: ASM Press, 2002, vol. 2, pp. 533–564
Skibbe, D.S., Fernandes, J.F., Medzihradszky, K.F., Burlingame, A.L., and Walbot, V., Mutator transposon activity reprograms the transcriptomes and proteomes of developing maize anthers, Plant J., 2009, vol. 59, no. 4, pp. 622–633.
Su, X., Construction and phenotypic analysis of Mutator-mediated mutants library in maize, M.D. Thesis, Baoding: Agricultural University of Hebei, 2009.
Huang, J., Ge, X., and Sun, M., Modified CTAB protocol using a silica matrix for isolation of plant genomic DNA, Biotechnique, 2000, vol. 28, no. 3, pp. 432–434.
Raizada, M.N., Nan, G.L., and Walbot, V., Somatic and germinal mobility of the RescueMu transposon in transgenic maize, Plant Cell, 2001, vol. 13, no. 7, pp. 1587–1608.
Liu, S., Yeh, C.T., Ji, T., Ying, K., Wu, H., Tang, H.M., Fu, Y., Nettleton, D., and Schnable, P.S., Mu transposon insertion sites and meiotic recombination events co-localize with epigenetic marks for open chromatin across the maize genome, PLoS Genet., 2009, vol. 5, no. 11, p. e1000733.
Lisch, D., Mutator transposons, Trends Plant Sci., 2002, vol. 7, no. 11, pp. 498–504.
Chakrabarti, R. and Schutt, C.E., The enhancement of PCR amplification by low molecular weight amides, Nucleic Acids Res., 2001, vol. 29, no. 11, pp. 2377–2381.
Hayashi, K., Pcr-sscp: a simple and sensitive method for detection of mutations in the genomic DNA, PCR Methods Appl., 1991, vol. 1, no. 1, pp. 34–38.
Zhong, W.J., Zhang, M.D., Yang, L.Q., Wang, M.C., Zheng, Y.L., Yang, W.P., and Gao, Y.J., Isolating the mutator transposable element insertional mutant gene mio16 of maize using double selected amplification of insertion flanking fragments (DSAIFF), J. Integr. Agric., 2012, vol. 11, no. 10, pp. 1592–1600.
Fu, X.Q., Morphological, biochemical and genetic analysis of a brittle stalk mutant of maize inserted by mutator transposon, M.D. Thesis, Wuhan: Huazhong Agricultural University, 2011.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
About this article
Cite this article
Yang, W.F., Tian, Y.H., Wang, T.T. et al. Isolating and confirming the MuDR-inserted flanking sequences of maize. Cytol. Genet. 51, 142–148 (2017). https://doi.org/10.3103/S0095452717020074
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0095452717020074